3.2
Impact Factor
ISSN: 1837-9664
J Cancer 2017; 8(13):2501-2510. doi:10.7150/jca.17407 This issue Cite
Research Paper
miR-618 Inhibits Prostate Cancer Migration and Invasion by Targeting FOXP2
1. Department of Radiation Oncology, Affiliated Cancer Hospital & Institute of Guangzhou Medical University, Guangzhou 510095, China;
2. Department of Pathology, Integrated Hospital of Traditional Chinese Medicine, Southern Medical University, Guangzhou 510315, China;
3. Guangdong Provincial Key Laboratory of Molecular Oncologic Pathology, Southern Medical University, Guangzhou 510515, China;
4. Department of Pathology, School of Basic Medical Sciences, Southern Medical University, Guangzhou 510515, China;
5. Department of Pathology, Affiliated Chenzhou Hospital, Southern Medical University (The First People's Hospital of Chenzhou), Chenzhou 423000, China;
6. Department of Urology, Nanfang Hospital, Southern Medical University/The First School of Clinical Medicine, Southern Medical University, Guangzhou 510515, China.
* These authors contributed equally to this work.
Received 2016-8-30; Accepted 2017-5-28; Published 2017-8-2
Abstract
miRNAs play critical role in the development and progression of prostate cancer. Here we studied the role of miR-618 in prostate cancer migration and invasion. miR-618 was downregulated in metastatic androgen-independent prostate cancer (AIPC), patients with low miR-618 had poor outcome. Overexpression of miR-618 inhibited migration and invasion and induced mesenchymal to epithelial transition (MET). Conversely, knockdown of miR-618 promoted migration and invasion and induced epithelial to mesenchymal transition (EMT). FOXP2 was the direct target of miR-618, and promoted TGF-β expression, inhibition of TGF-β reversed the effect of miR-618 knockdown. We further analyzed the correlation between miR-618 expression and FOXP2 in human prostate cancer tissues, and found there was a negative correlation between miR-618 expression and FOXP2 levels. In conclusion, we found miR-618 inhibited prostate cancer migration and invasion by targeting FOXP2 and inhibiting TGF-β.
Keywords: prostate cancer, miR-618, FOXP2, migration, TGF-β.
Introduction
Prostate cancer is the second most common cause of male cancer deaths. Some biomarkers are used for its diagnosis and prognosis, for example, prostate-specific antigen (PAS), EZH2, Hepsin and TGF-β1 [1, 2], but the survival of patients is still poor. Androgens promote prostate cancer growth, inhibition its secretion or function using chemical/surgical castration or antiandrogens which block the androgens bind to the androgen receptor (AR) has been used to treat prostate cancer, androgen deprivation therapy (ADT) suppresses the progression of prostate cancer [3], we call prostate cancer which is sensitive to ADT as androgen-dependent prostate cancer (ADPC), but prostate cancer often generates resistance to ADT, we call prostate cancer which is insensitive to ADT as androgen-independent prostate cancer (AIPC) or castration-resistant prostate cancer (CRPC) [4]. CRPC has strong metastatic ability and high recurrent rate, and is the main challenge for prostate cancer therapy now. Many genes and non-coding RNAs have been found to regulate the development and progression of prostate cancer, such as mixed-lineage leukemia protein (MLL) complex [5], BRD4 [6], miR-32 [7] and long non-coding RNA SChLAP1 [8]. However, the regulatory mechanism of prostate cancer hasn't been well understood, new regulators are waiting to be discovered.
The role of microRNA (miRNA) in tumor biology has been widely studied, it suppresses target mRNA through a combination of translational suppression and mRNA destabilization [9], it functions as an oncogene or a tumor suppressor, for example, miR-221 promotes metastasis of breast cancer and self-renewal of breast cancer stem cells by targeting ATXN1 [10], it also promotes the progression of CRPC by targeting HECTD2 and RAB1A [11]. miR-618 can serve as biomarker for lymphomagensis [12], male breast cancer [13], head and neck squamous cell carcinoma [14] and hepatocellular carcinoma [15]. It suppresses proliferation of anaplastic thyroid cancer by targeting XIAP [16]. The function and molecular regulatory mechanism of miR-618 in prostate cancer have not been studied until now. FOXP2 belongs to forkhead box P family, the role of FOXP2 in tumors hasn't studied widely. In hepatocellular carcinoma, patients with low FOXP2 expression have poor survival, FOXP2 downregulation promotes invasion [17]. FOXP2 downregulation promotes breast cancer metastasis and self-renewal of breast cancer stem cells, it inhibits metastasis and cancer stem cell associated factors, such as POSTN, ALDH1 and CD44, patients with low FOXP2 expression have poor survival [18], suggesting it's a tumor suppressor for breast cancer.
Here, we studied the role of miR-618 in prostate cancer progression, we first analyzed miR-618 expression in ADPC samples and AIPC samples, and the relationship between miR-618 expression and clinical outcome, then we investigated the effect of miR-618 on migration and invasion. Finally, we looked for the target gene of miR-618. We found miR-618 inhibited prostate cancer migration and invasion by targeting FOXP2, FOXP2 could inhibit TGF-β expression, TGF-β inhibited the effect of miR-618 knockdown in prostate cancer.
Materials and methods
Cell culture and transfection
Two primary-cultured prostate epithelial cell lines were maintained in 5% Ham's F12 (Life Technologies) with 5% fetal bovine serum (FBS) (GIBCO), 5 ug/ml insulin, 10ng/ml epidermal growth factor, 1ug/ml hydrocortisone (Sigma). Prostate cancer cell lines, including LNCap, 22RV1, Tsu-Pr1, PC3 and DU145 were obtained from the American Type Culture Collection (ATCC). LNCap and 22RV1 were androgen receptor (AR)-positive ADPC cells, Tsu-Pr1, PC3 and DU145 were AR-negative AIPC cells. They were cultured in RPMI-1640 medium supplemented with 10% FBS.
miR-618 mimic, miR-618 inhibitor and their cognate control RNAs (indicated as negative control (NC)) were synthesized by RIBOBIO (Guangzhou, China), Cells were seeded in six-well plates, and 20nm oligonucleotides were transfected into indicated prostate cancer cells using Lipofectamine RNAiMAX Reagent (Life Technoligies) according to the manufacturer's instructions, when the density of cells was up to 60 ~ 70%.
To overexpress FOXP2, we cloned the CDS fragment of FOXP2 into pMSCV-puro vector (Clontech). To determine whether miR-618 directly binds to FOXP2, the 3'UTR of FOXP2 fragment containing wild or mutational binding site of miR-618 was cloned into pEGFP-C1 and pGL3 vectors, the vectors were transfected into indicated cells using Fugene HD (Promega) according to the manufacturer's instructions.
Human prostate cancer samples
To determine miR-618 expression in prostate cancer tissues, 60 clinical samples were collected from the Department of Pathology, TCM-Integrated Hospital, Southern Medical University, Guangzhou, China, they were frozen and stored in liquid nitrogen for further use. A cohort of 126 samples used for survival analysis were also obtained from the same institute. To perform clinical follow-up, we constructed an electronic file for each patient, including name, age, address, telephone number, hospitalization outcome and discharge diagnosis. Patients were followed up from 2-4 weeks after discharge, then every 2 months at least once a visit to understand the treatment effect, survival and other condition changes. The follow-up time was up to 60 months. The clinicopathological characteristics were shown in Supplemental table 1. For the use of these samples for research purposes. All samples were obtained with written informed and approved by the Institutional Research Ethics Committee of Southern Medical University.
RNA isolation and real time RT-PCR
Total miRNA of cells and tissues was extracted using miRNeasy mini Kit (Qiagen) according to the manufacturer's protocol, cDNA was synthesized from 10ng of total miRNA using miR-618 specific primers (TaqMan miRNA assay, Applied Biosystems) and Taqman miRNA Reverse Transcription Kit (Applied Biosystems). miR-618 expression level was quantified using miRNA-specific TaqMan miRNA Assay Kit (Applied Biosystems). Total RNA was extracted using TRIzol (Life Technologies), cDNA was reverse transcribed from 5ug total RNA using HiScript 1st Strand cDNA Synthesis Kit (Vazyme). The gene expression level was quantified using AceQ qPCR SYBR Green Master Mix (Vazyme) following the manufacturer's instructions. Real-time RT-PCR was performed using the Applied Biosystems 7500 Sequence Detection System. RUN24 was used as an endogenous control for miRNA date normalization, GAPDH was used as an endogenous control for gene normalization. Relative expressions were calculated using a comparative Ct method [19].
Western blot assay
Western blot assay was performed as previously described [20]. Antibodies used were described as follow: FOXP2 (Cell signaling, #8198), Vimentin (Cell signaling, #5741), E-Cadherin (Cell signaling, #3195), α-Catenin (Cell signaling, #2163), Fibronectin (Cell signaling, #4706), Snail (Cell signaling, #3879), TGF-β (Cell signaling, #3709) and GFP (Cell signaling, #2956). The membranes were stripped and re-probed with anti-GAPDH (Sigma, G9545) or anti-β-actin (Sigma, SAB5500001) antibodies as loading control.
Wound healing assay
Wound healing assay was performed as previously described [21].
Transwell invasion assay
Transwell invasion assay was performed as previously described [20]. In brief, 24-well Transwell inserts (BD Biosciences) were coated with Matrigel matrix for 2h at 37℃. The same amount of prostate cancer cells were seeded onto 24-well Transwell inserts and allowed to migrate through about 8um pores towards 10% RPMI-1640 in the bottom chamber. After 24h, Membranes were then fixed in 4% paraformaldehyde. The seeding section of the membrane was removed with a cotton swab and the bottom surface stained with 0.2% crystal violet, washed and imaged. Cells were counted from the images. Each experiment was repeated at least in triplicate.
3D spheroid invasion assay
3D spheroid invasion assay was performed according to the standard method described previously [22]. Briefly, prostate cells were seeded on 2% Matrigel coated in 24-well plates, the medium was refreshed every other day, Cells could form spheres, and were photographed at 2-day intervals for 10 days.
Luciferase reporter assay
Cells which were seeded in triplicate in 24-well plates were cotransfected with constructed pGL3 vector and miR-618 mimic using Lipofectamine 2000 (Life Technologies) at 60% confluence, luciferase activities were investigated after 48h using Dual Luciferase Reporter Assay Kit (Promega) according to the manufacturer's instruction. All experiments were carried out in triplicate.
Statistical analysis
SPSS 13.0 (SPSS, Chicago, IL. USA) was used to analyze the data. Results are presented as the mean ± standard deviation (STDEV) for at least three repeated individual experiments for each group, statistical differences were analyzed using student's t-test for independent. For the clinical samples, One-way analysis of variance (ANOVA) with Tukey post-test was used to analyze miR-618 expression in tissue samples. Chi-Square test was applied to analyze the correlation between miR-618 expression and FOXP2 expression in patients. Survival curves were plotted using the Kaplan-Meier method and were compared by the log-rank test. A p-value of less than 0.05 was considered significant.
Results
miR-618 is downregulated in aggressive malignancy prostate cancer tissues and cells
To investigate the role of miR-618 in prostate cancer, we determined miR-618 expression in untreated primary prostate tumor samples and metastatic castration-resistant prostate tumor samples, and found miR-618 was significantly downregulated in castration-resistant prostate tumor samples (p< 0.05) (Figure 1A). We also determined miR-618 expression in prostate cancer tissues which came from nine patients who had treated with ADT and generated resistance to antiandrogen therapy, Real-time RT-PCR analysis showed miR-618 was downregulated in AIPC samples (Figure 1B), suggesting miR-618 might inhibit prostate cancer progression. We also determined the relationship between miR-618 expression and clinical outcome, follow-up data showed the overall survival of prostate cancer patients with high miR-618 levels were significantly longer than those with low miR-618 levels (log-rank test, p<0.001) (Figure 1C). These revealed that miR-618 expression had a negative correlation with advanced prostate cancer.
miR-618 is downregulated in aggressive malignancy prostate cancer tissues and cells. (A). Real-time RT-PCR analyzed miR-618 expression in untreated primary prostate cancer tissues and castration-resistant prostate cancer tissues. (B). Real-time PCR determined miR-618 expression in paired ADPC tissues and AIPC tissues. (C). Kaplan-Meier overall survival curves for the patients with high versus low miR-618 expression. (D). Real-time RT-PCR analyzed miR-618 expression in primary-cultured prostate epithelial cells (indicated as N1 and N2) and prostate cancer cells. Error bars represent mean ± SD for at least three biological replicates. * P<0.05.
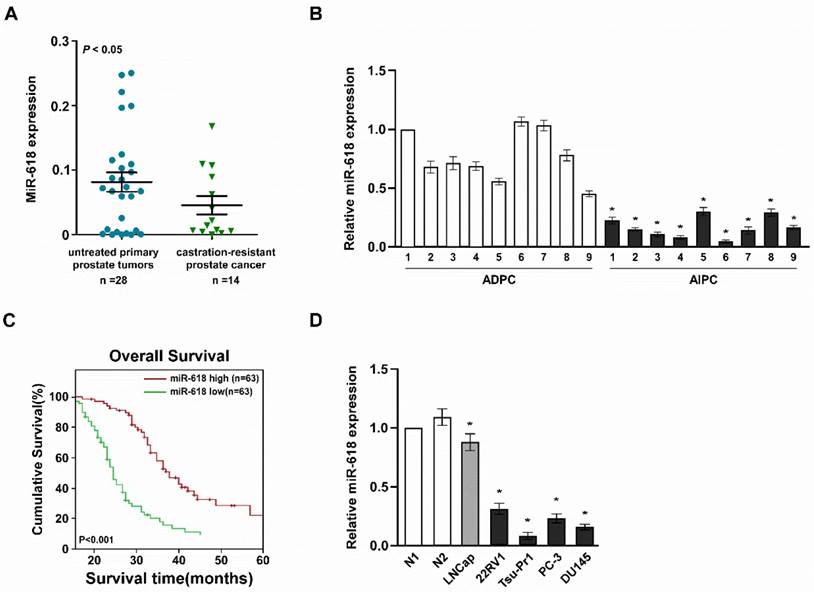
We also determined miR-618 expression in primary-cultured prostate epithelial cells and prostate cancer cells, and found miR-618 was downregulated in prostate cancer cells, its expression was lower in AR-negative AIPC cell lines than AR-positive AIPC cell lines (Figure 1D). These suggested miR-618 is downregulated in prostate cancer cells and aggressive malignancy prostate cancer cells compared to normal prostate epithelial cells, but its expression was lower in aggressive malignancy prostate cancer cells.
Overexpression of miR-618 inhibits migration and invasion of prostate cancer and induces mesenchymal to epithelial transition (MET)
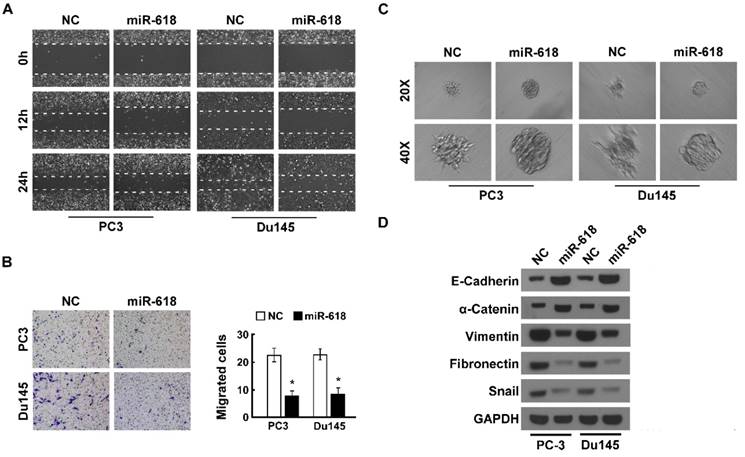
miR-618 was low expressed in castration-resistant prostate cancer with strong metastatic ability, so we determined the effect of miR-618 on migration and invasion of prostate cancer. We overexpressed miR-618 in prostate cancer cells PC3 and Du145. Wound healing assay showed miR-618 overexpression inhibited cell migration in indicated prostate cancer (Figure 2A). Transwell invasion assay found the number of invaded prostate cancer cells overexpressing miR-168 significantly reduced, compared to the negative control (NC), suggesting overexpression of miR-618 significantly inhibited cell invasion (Figure 2B). 3D spheroid invasion assay showed that the invasive projections were disappeared in the cells-overexpressing miR-618, compared to the negative control (NC) (Figure 2C), this result further suggested miR-618 inhibited cell invasion. Cell migration and invasion always accompany the change of cell morphology, we further confirmed this result by determined the expression of epithelial markers and mesenchymal markers [23], western blot assay showed when miR-618 was overexpressed, epithelial markers E-Cadherin and α-Catenin were upregulated, mesenchymal markers Vimentin, Frbronrcyin and Snail were downregulated (Figure 2D), suggesting overexpression of miR-618 induces MET.
Overexpression of miR-681 inhibits migration and invasion of prostate cancer and induces MET. (A). Wound healing assay analyzed the effect of miR-618 overexpression on migration of prostate cancers. Wound closures were photographed at 0, 12 and 24 hours after wounding. (B). Transwell invasion assay analyzed the effect of miR-618 overexpression on invasion. (left) Representative micrographs of transwell invasion assay of the indicated cells, (right) quantification of indicated invading cells. (C). 3D spheroid invasion assay determined the effect of miR-618 overexpression on metastasis. (D). Western blot analyzed the expression of epithelial markers and mesenchymal markers when miR-618 was overexpression, GAPDH was used as the loading control. Error bars represent mean ± SD for at least three biological replicates. *P<0.05.
Knockdown of miR-618 promotes migration and invasion of prostate cancer and induces epithelial to mesenchymal transition (EMT)
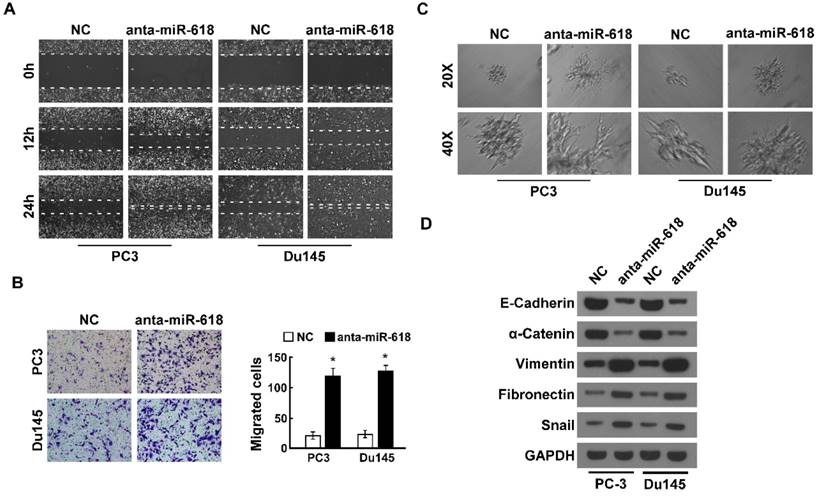
In order to confirm that miR-618 inhibited migration and invasion of prostate cancer, we knocked down miR-618 in the same cells. Wound healing assay showed miR-618 knockdown promoted prostate cell migration (Figure 3A). Transwell invasion assay showed the number of invaded prostate cancer cells with miR-168 knockdown significantly increased. This revealed that knockdown of miR-618 significantly promoted cell invasion (Figure 3B). 3D spheroid invasion assay revealed that indicated cells with miR-618 knockdown generated more invasive projections, compared to the negative control (NC), suggesting miR-618 knockdown promoted cell invasion (Figure 3C). We further determined the expression of epithelial markers and mesenchymal markers, and found E-Cadhein and α-Catenin were downregulated, the expression of Vimentin, Frbronrcyin and Snail was upregulated, suggesting knockdown of miR-618 induces EMT.
miR-618 directly targets to the 3'UTR of FOXP2
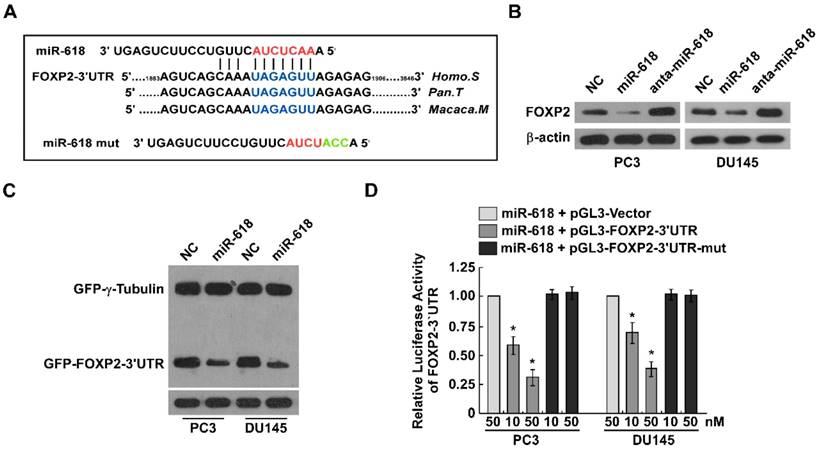
We used miRWALK to predict the target gene of miR-618 [24], and found FOXP2 might be the target gene of miR-618. FOXP2 is a key transcription factor for regulation of the development human speech, language and central nervous system [25]. The seed sites of miR-618 bound to the 3'UTR of FOXP2, this seed sites were evolutionarily conservative, miR-618 also bound to the 3'UTR of FOXP2 in Pan. T and Macaca.M (Figure 4A). We further determined whether FOXP2 is the target of miR-618, western blot assay showed that FOXP2 was downregulated when miR-618 was upregulated, while when miR-618 was downregulated, FOXP2 was upregulated (Figure 4B), suggesting FOXP2 expression was opposite to miR-618 expression. In order to investigate whether miR-618 directly binds to the 3'UTR of FOXP2, we cloned the 3'UTR of FOXP2 fragment containing the binding sites of miR-618 into pEGFP-C1 vector, overexpression of miR-618 inhibited GFP expression, but it hadn't no effect on GFP-γ-Tubulin expression (Figure 4C). We also cloned the 3'UTR of FOXP2 into pGL3 vector, Luciferase activity assay revealed that miR-618 overexpression inhibited the luciferase activity of 3'UTR of FOXP2 in a dose-dependent manner, but mutation of the binding site in 3'UTR of FOXP2 abrogated responsiveness to miR-618 (Figure 4D). These results suggested FOXP2 is the direct target of miR-618.
Knockdown of miR-618 promotes migration and invasion of prostate cancer and induces EMT. (A). Wound healing assay analyzed the effect of miR-618 knockdown on migration of prostate cancers. Wound closures were photographed at 0, 12 and 24 hours after wounding. (B). Transwell invasion assay analyzed the effect of miR-618 knockdown on invasion. (left) Representative micrographs of transwell invasion assay of the indicated cells, (right) quantification of indicated invading cells. (C). 3D spheroid invasion assay determined the effect of miR-618 knockdown on metastasis. (D). Western blot analyzed the expression of epithelial markers and mesenchymal markers when miR-618 was knock-downed, GAPDH was used as the loading control. Error bars represent mean ± SD for at least three biological replicates. *P<0.05.
FOXP2 is the direct target of miR-618. (A). Predicted miR-618 target sequences (blue) in the 3'UTR of FOXP2 (FOXP2-wt-3'UTR) and positions of three mutated nucleotides (cyan) in the 3'UTR of FOXP2 (FOXP2-mut-3'UTR). (B). Western blot analyzed FOXP2 expression in indicated cells transfecting miR-618 mimic or miR-618 inhibitor, β-Actin was used as the loading control. (C). Western blot analyzed GFP expression in indicated cells transfecting miR-618 mimic or miR-618 inhibitor, α-Tubulin was used as the loading control. (D). Luciferase reporter assay determined luciferase activity in indicated cells cotransfected miR-618 mimic and pGL3-FOXP2-wt-3'UTR or pGL3-FOXP2-mut-3'UTR. Error bars represent mean ± SD for at least three biological replicates. *P<0.05.
FOXP2 activates TGF-β to promote migration and invasion
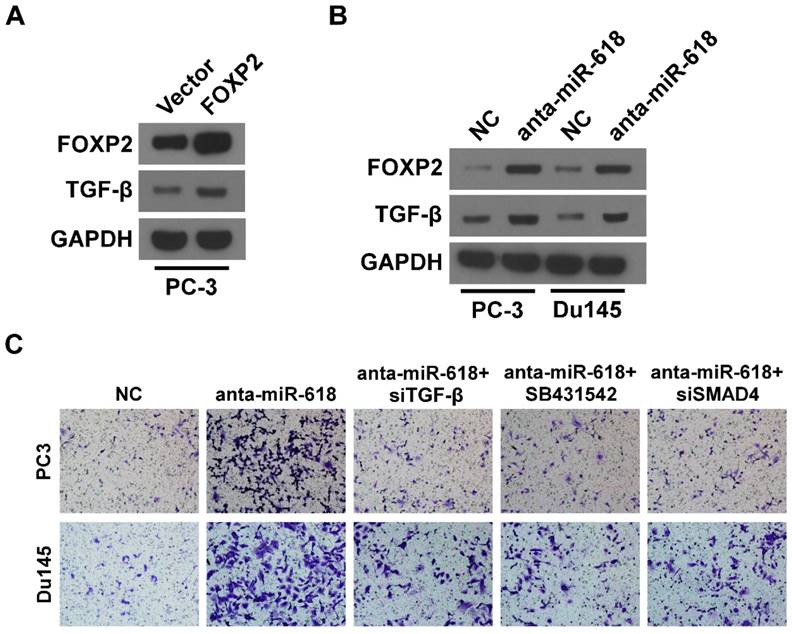
TGF-β signaling is a key player in tumor development, its misregulation often be discovered in diverse tumors. In normal and premalignant cells, TGF-β can function as a tumor suppressor. In malignant tumor cells, TGF-β can function as a tumor promoter, and promotes tumor invasiveness, dissemination and metastatic colonization [26, 27]. We investigated whether FOXP2 regulates TGF-β expression. Western blot assay revealed that FOXP2 overexpression increased TGF-β expression (Figure 5A). Inhibition of miR-618 also increased TGF-β expression (Figure 5B), suggesting the miR-618 suppressed FOXP2 to inhibit TGF-β, miR-618 knockdown promoted FOXP2 expression to activate TGF-β and then induced EMT. In other word, if TGF-β was inhibited, the effect of miR-618 knockdown on cellular migration could be inhibited. We used TGF-β small inference RNA (siTGF-β), small molecule inhibitor for TGF-β, SB431542 [28] and SMAD4 small inference RNA (siSMAD4) to inhibit TGF-β. SMAD4 can form hetero-oligomers with pathway-restricted SMADs to mediate SMADs translocate into the nucleus and activate response elements [29]. Tranwell migration assay showed inhibition TGF-β indeed suppressed cellular invasion (Figure 5C). These suggested FOXP2 promoted migration and invasion by increasing TGF-β.
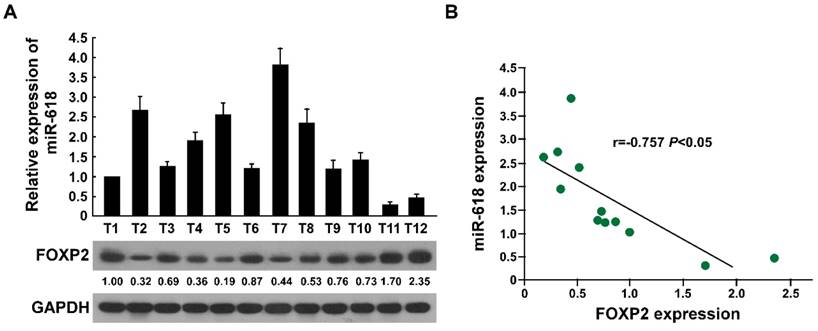
To further confirm FOXP2 was the target of miR-618, we collected 12 prostate cancer tissues. Real-time RT-PCR and western blot assay found patients with high miR-618 had low FOXP2 expression and patients with low miR-618 expression had high FOXP2 expression (Figure 6A). There was a negative correlation between miR-618 expression and FOXP2 expression (r=-0.757, p< 0.05) (Figure 6B). This suggested FOXP2 is the target gene of miR-618.
FOXP2 promotes cell invasion by increasing TGF-β expression. (A). Western blot analyzed TGF-β expression once miR-618 overexpression in PC3 cells. GAPDH was used as the loading control. (B). Western blot analyzed FOXP2 and TGF-β expression when miR-618 was knock-downed in PC3 and DU145 cells. GAPDH was used as the loading control. (C). when miR-618 and TGF-β were inhibited simultaneously, transwell invasion assay was used to assess cell invasion.
The relationship between miR-618 expression and FOXP2 expression. (A). Real-time RT-PCR assessed miR-618 expression in 12 prostate cancer tissues, western blot analyzed FOXP2 expression in the same tissues. GAPDH was used as the loading control. (B). The correlation analysis between miR-618 expression and FOXP2 expression in 12 prostate cancer tissues.
Discussion
In present study, we found miR-618 was downregulated in AIPC tissues, prostate cancer patients with low miR-618 expression had shorter survival than those with high miR-618 expression, suggesting there was a negative correlation between miR-618 expression and clinical outcome. We further studied the effect of miR-618 on migration and invasion, and found overexpression miR-618 inhibited migration and invasion and induced MET, miR-618 knockdown promoted migration and invasion and induced EMT. FOXP2 was the direct target of miR-618, mechanism analysis revealed FOXP2 could increase TGF-β expression to promote migration and invasion, we further confirmed the correlation between miR-618 expression and FOXP2 expression in prostate cancer tissues, and found miR-618 expression was negative correlated with FOXP2 expression.
Laura Stumm and colleagues use immunohistochemistry to analyze FOXP2 expression in 7779 prostate cancer tissues, they found there is a positive correlation between FOXP2 expression and advanced tumor stage, Gleason grade and lymph node metastases, especially in ERG-negative prostate cancer, high FOXP2 expression is correlated with early biochemical recurrence. This suggests FOXP2 is an oncogene in prostate cancer [30]. This is consistent with our results, FOXP2 promoted migration and invasion of prostate cancer. This also suggested FOXP2 has different role in different kinds of tumors. TGF-β promotes migration and invasion and the expansion of cancer stem cells, FOXP2 overexpression promoted TGF-β expression to promote prostate cancer migration and invasion, so we thought miR-618 inhibits prostate cancer migration and invasion by inhibition TGF-β. Some target genes of FOXP2 might activate TGF-β signaling, this inference was still to be confirmed by RNA-seq or chromatin immunoprecipitation (ChIP). Many miRNAs could regulate prostate cancer stem cells, for example, miR-320 suppresses the self-renewal of prostate cancer stem cells by inhibiting Wnt signaling [31], miR-7 suppresses prostate cancer stem cells by repressing KLF4/PI3K/AKT/P21 pathway [32]. TGF-β signaling often promotes cancer stem cells, miR-618 inhibited TGF-β, it might inhibit the self-renewal of prostate cancer stem cells.
Increased AR activity is the major driver of CRCP, for example, amplification of AR gene, constitutive activation mutation and AR signaling activating by other signaling pathway [33]. TGF-β signaling promotes apoptosis in normal and premalignant cells, AR signaling could inhibit TGF-β signaling by down-regulating TGF-β receptors, but TGF-β is upregulated in CRCP, it promotes CRCP metastasis, these suggests AR signaling doesn't suppress TGF-β signaling pathway in CRCP, the cross-talking between AR signaling and TGF-β signaling was changed in CRCP, but the detailed mechanism was unclear [34].
The nanoparticle-based delivery systems (nanovector) can deliver miRNA to cancer cells in vivo, for example, miR-34a-delivering therapeutic nanocomplexes with CC9 peptide can efficiently treat pancreatic cancers [35, 36]. miR-100 is high expression in normal mammary tissues, and inhibits breast cancer growth, metastasis and self-renewal of breast cancer stem cell. Anti-CD44-nanovector can deliver miR-100 to breast cancer cells to inhibit breast cancer development and initiation [37]. We found miR-618 expression was higher in normal prostate epithelial cells than prostate cancer cells, it could be delivered to prostate cells using nanovector. In summary, miR-618 is downregulated in aggressive malignancy prostate cancer tissues and cells, patients with low miR-618 expression have poor survival. miR-186 inhibits the migration, invasion and induces MET of prostate cancer by targeting FOXP2 and inhibiting TGF-β expression. Furthermore, there is a negative correlation between FOXP2 expression and miR-618 levels in prostate cancer patients.
Supplementary Material
Supplementary table 1.
Acknowledgements
This study was supported by two grants from the National Natural Science Foundation of China (No.81372769, No.81328017), three Science and Technology planning Projects of Guangdong Province (No.2013B051000050, No.2014A020212538 and No.2016A020215175), a Natural Science Foundation of Guangdong Province (No.2016A030313583), a Medical Scientific Research Foundation of Guangdong Province (No. A2016555), and a Science and Technology planning Project of Guangzhou (No. 201704020070).
Authors' Contributions
XL Song, SC Zhao and ZQ Wu designed experiment. Y Tang, XL Song and XH Lei performed experiment. Y Tang, XL Song, SC Zhao and ZQ Wu analyzed and interpreted the data. Y Tang, XL Song and ZQ Wu wrote the paper. XH Lei provided administrative, technical, or material support. SC Zhao and ZQ Wu supervised the study.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Bradford TJ, Tomlins SA, Wang X, Chinnaiyan AM. Molecular markers of prostate cancer. Urologic oncology. 2006;24(6):538-551
2. Varambally S, Dhanasekaran SM, Zhou M, Barrette TR, Kumar-Sinha C, Sanda MG, Ghosh D, Pienta KJ, Sewalt RG, Otte AP. et al. The polycomb group protein EZH2 is involved in progression of prostate cancer. Nature. 2002;419(6907):624-629
3. Harris WP, Mostaghel EA, Nelson PS, Montgomery B. Androgen deprivation therapy: progress in understanding mechanisms of resistance and optimizing androgen depletion. Nature clinical practice Urology. 2009;6(2):76-85
4. Li P, Yang R, Gao WQ. Contributions of epithelial-mesenchymal transition and cancer stem cells to the development of castration resistance of prostate cancer. Molecular cancer. 2014;13:55
5. Malik R, Khan AP, Asangani IA, Cieslik M, Prensner JR, Wang X, Iyer MK, Jiang X, Borkin D, Escara-Wilke J. et al. Targeting the MLL complex in castration-resistant prostate cancer. Nature medicine. 2015;21(4):344-352
6. Asangani IA, Dommeti VL, Wang X, Malik R, Cieslik M, Yang R, Escara-Wilke J, Wilder-Romans K, Dhanireddy S, Engelke C. et al. Therapeutic targeting of BET bromodomain proteins in castration-resistant prostate cancer. Nature. 2014;510(7504):278-282
7. Jalava SE, Urbanucci A, Latonen L, Waltering KK, Sahu B, Janne OA, Seppala J, Lahdesmaki H, Tammela TL, Visakorpi T. Androgen-regulated miR-32 targets BTG2 and is overexpressed in castration-resistant prostate cancer. Oncogene. 2012;31(41):4460-4471
8. Prensner JR, Iyer MK, Sahu A, Asangani IA, Cao Q, Patel L, Vergara IA, Davicioni E, Erho N, Ghadessi M. et al. The long noncoding RNA SChLAP1 promotes aggressive prostate cancer and antagonizes the SWI/SNF complex. Nature genetics. 2013;45(11):1392-1398
9. Ameres SL, Zamore PD. Diversifying microRNA sequence and function. Nature reviews Molecular cell biology. 2013;14(8):475-488
10. Ke J, Zhao Z, Hong SH, Bai S, He Z, Malik F, Xu J, Zhou L, Chen W, Martin-Trevino R. et al. Role of microRNA221 in regulating normal mammary epithelial hierarchy and breast cancer stem-like cells. Oncotarget. 2015;6(6):3709-3721
11. Sun T, Wang X, He HH, Sweeney CJ, Liu SX, Brown M, Balk S, Lee GS, Kantoff PW. MiR-221 promotes the development of androgen independence in prostate cancer cells via downregulation of HECTD2 and RAB1A. Oncogene. 2014;33(21):2790-2800
12. Fu A, Hoffman AE, Liu R, Jacobs DI, Zheng T, Zhu Y. Targetome profiling and functional genetics implicate miR-618 in lymphomagenesis. Epigenetics: official journal of the DNA Methylation Society. 2014;9(5):730-737
13. Fassan M, Baffa R, Palazzo JP, Lloyd J, Crosariol M, Liu CG, Volinia S, Alder H, Rugge M, Croce CM. et al. MicroRNA expression profiling of male breast cancer. Breast cancer research: BCR. 2009;11(4):R58
14. Hui L, Wu H, Yang N, Guo X, Jang X. Identification of prognostic microRNA candidates for head and neck squamous cell carcinoma. Oncology reports. 2016;35(6):3321-3330
15. Abdalla MA, Haj-Ahmad Y. Promising Candidate Urinary MicroRNA Biomarkers for the Early Detection of Hepatocellular Carcinoma among High-Risk Hepatitis C Virus Egyptian Patients. Journal of Cancer. 2012;3:19-31
16. Cheng Q, Zhang X, Xu X, Lu X. MiR-618 inhibits anaplastic thyroid cancer by repressing XIAP in one ATC cell line. Annales d'endocrinologie. 2014;75(4):187-193
17. Yan X, Zhou H, Zhang T, Xu P, Zhang S, Huang W, Yang L, Gu X, Ni R, Zhang T. Downregulation of FOXP2 promoter human hepatocellular carcinoma cell invasion. Tumour biology: the journal of the International Society for Oncodevelopmental Biology and Medicine. 2015
18. Cuiffo BG, Campagne A, Bell GW, Lembo A, Orso F, Lien EC, Bhasin MK, Raimo M, Hanson SE, Marusyk A. et al. MSC-regulated microRNAs converge on the transcription factor FOXP2 and promote breast cancer metastasis. Cell stem cell. 2014;15(6):762-774
19. Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods. 2001;25(4):402-408
20. Drake JM, Barnes JM, Madsen JM, Domann FE, Stipp CS, Henry MD. ZEB1 coordinately regulates laminin-332 and {beta}4 integrin expression altering the invasive phenotype of prostate cancer cells. The Journal of biological chemistry. 2010;285(44):33940-33948
21. Song LB, Li J, Liao WT, Feng Y, Yu CP, Hu LJ, Kong QL, Xu LH, Zhang X, Liu WL. et al. The polycomb group protein Bmi-1 represses the tumor suppressor PTEN and induces epithelial-mesenchymal transition in human nasopharyngeal epithelial cells. The Journal of clinical investigation. 2009;119(12):3626-3636
22. Cai J, Guan H, Fang L, Yang Y, Zhu X, Yuan J, Wu J, Li M. MicroRNA-374a activates Wnt/beta-catenin signaling to promote breast cancer metastasis. The Journal of clinical investigation. 2013;123(2):566-579
23. Tsai JH, Yang J. Epithelial-mesenchymal plasticity in carcinoma metastasis. Genes & development. 2013;27(20):2192-2206
24. Dweep H, Sticht C, Pandey P, Gretz N. miRWalk-database: prediction of possible miRNA binding sites by "walking" the genes of three genomes. Journal of biomedical informatics. 2011;44(5):839-847
25. Konopka G, Bomar JM, Winden K, Coppola G, Jonsson ZO, Gao F, Peng S, Preuss TM, Wohlschlegel JA, Geschwind DH. Human-specific transcriptional regulation of CNS development genes by FOXP2. Nature. 2009;462(7270):213-217
26. Massague J. TGFbeta in Cancer. Cell. 2008;134(2):215-230
27. Zavadil J, Bottinger EP. TGF-beta and epithelial-to-mesenchymal transitions. Oncogene. 2005;24(37):5764-5774
28. Koo BH, Kim Y, Je Cho Y, Kim DS. Distinct roles of transforming growth factor-beta signaling and transforming growth factor-beta receptor inhibitor SB431542 in the regulation of p21 expression. European journal of pharmacology. 2015
29. Heldin CH, Miyazono K, ten Dijke P. TGF-beta signalling from cell membrane to nucleus through SMAD proteins. Nature. 1997;390(6659):465-471
30. Stumm L, Burkhardt L, Steurer S, Simon R, Adam M, Becker A, Sauter G, Minner S, Schlomm T, Sirma H. et al. Strong expression of the neuronal transcription factor FOXP2 is linked to an increased risk of early PSA recurrence in ERG fusion-negative cancers. Journal of clinical pathology. 2013;66(7):563-568
31. Hsieh IS, Chang KC, Tsai YT, Ke JY, Lu PJ, Lee KH, Yeh SD, Hong TM, Chen YL. MicroRNA-320 suppresses the stem cell-like characteristics of prostate cancer cells by downregulating the Wnt/beta-catenin signaling pathway. Carcinogenesis. 2013;34(3):530-538
32. Chang YL, Zhou PJ, Wei L, Li W, Ji Z, Fang YX, Gao WQ. MicroRNA-7 inhibits the stemness of prostate cancer stem-like cells and tumorigenesis by repressing KLF4/PI3K/Akt/p21 pathway. Oncotarget. 2015
33. Chen CD, Welsbie DS, Tran C, Baek SH, Chen R, Vessella R, Rosenfeld MG, Sawyers CL. Molecular determinants of resistance to antiandrogen therapy. Nature medicine. 2004;10(1):33-39
34. Zhu ML, Kyprianou N. Androgen receptor and growth factor signaling cross-talk in prostate cancer cells. Endocrine-related cancer. 2008;15(4):841-849
35. Pramanik D, Campbell NR, Karikari C, Chivukula R, Kent OA, Mendell JT, Maitra A. Restitution of tumor suppressor microRNAs using a systemic nanovector inhibits pancreatic cancer growth in mice. Molecular cancer therapeutics. 2011;10(8):1470-1480
36. Hu QL, Jiang QY, Jin X, Shen J, Wang K, Li YB, Xu FJ, Tang GP, Li ZH. Cationic microRNA-delivering nanovectors with bifunctional peptides for efficient treatment of PANC-1 xenograft model. Biomaterials. 2013;34(9):2265-2276
37. Deng L, Shang L, Bai S, Chen J, He X, Martin-Trevino R, Chen S, Li XY, Meng X, Yu B. et al. MicroRNA100 inhibits self-renewal of breast cancer stem-like cells and breast tumor development. Cancer research. 2014;74(22):6648-6660
Author contact
![]() Corresponding authors: Ziqing Wu, Department of Pathology, Integrated Hospital of Traditional Chinese Medicine, Southern Medical University, Shiliugang Road, Haizhu District. Guangzhou 510315, China. E-mail: hailiancom. Shan-Chao Zhao, Department of Urology, Nanfang Hospital, Southern Medical University/The First School of Clinical Medicine, Southern Medical University, No.1023, South Shatai Road, Baiyun District, Guangzhou, Guangdong, 510515, China. E-mail: zhaoshanchaonet.
Corresponding authors: Ziqing Wu, Department of Pathology, Integrated Hospital of Traditional Chinese Medicine, Southern Medical University, Shiliugang Road, Haizhu District. Guangzhou 510315, China. E-mail: hailiancom. Shan-Chao Zhao, Department of Urology, Nanfang Hospital, Southern Medical University/The First School of Clinical Medicine, Southern Medical University, No.1023, South Shatai Road, Baiyun District, Guangzhou, Guangdong, 510515, China. E-mail: zhaoshanchaonet.

 Global reach, higher impact
Global reach, higher impact