3.2
Impact Factor
ISSN: 1837-9664
J Cancer 2018; 9(24):4712-4717. doi:10.7150/jca.28636 This issue Cite
Research Paper
CLDN10 is Associated with Papillary Thyroid Cancer Progression
Department of Thyroid & Breast Surgery, The First Affiliated Hospital of Wenzhou Medical University, Wenzhou, Zhejiang, PR China
*Contributed Equally.
Received 2018-7-19; Accepted 2018-8-31; Published 2018-11-25
Abstract
The incidence of thyroid cancer is staying at a high level. Claudin family is a skelemin contacting with the intercellular junction and can keep a dynamic balance between cells. Recently, many types of research indicated that the expression level of claudins is closely related to various cancer types and they can be novel diagnostic markers. For instance, Claudin-10(CLDN10) is the high expression in primary hepatocellular carcinoma, papillary thyroid cancer (PTC) and so on. But the biological role and function of CLDN10 in PTC are unclear. In our study, we measured the expression of CLDN10 in human normal tissues and matched PTC tissues by quantitative real-time polymerase chain reaction (qRT-PCR) and this observation was consistent with that in the TCGA cohort. We discovered that high expression of CLDN10 was correlated with lymph node metastasis, age and Histological type in TCGA cohorts. Kaplan-Meier analysis showed that patients with higher CLDN10 expression had a worse overall survival. In vitro, CLDN10 could promote cellular proliferation, migration, and invasion in PTC cell lines. In a word, CLDN10 is a functionally gene facilitating tumorgenesis in PTC and acts as an oncogene in PTC.
Keywords: CLDN10, PTC, tumorigenesis
Introduction
Thyroid cancer is the most common cancers in the endocrine system and the occurrence rate of it tends to be stable in recent years [1]. Papillary thyroid cancer (PTC) consists of the largest group of thyroid cancer and it has few fatalities [2-4]. The five-year survival rate of PTC exceeds 95% [5]. PTC has the high risk of recurrence and hence grasp the tumorgenesis to give a precise treatment is important. However, the potential molecular mechanisms of PTC are still covered.
Claudin family is necessary for cell's tight junctions and it has more than 20 proteins [6-8]. A lot of claudins was found playing a significant role in tumorigeneses, such as CLDN1, CLDN3 and CLDN4 [9]. Plenty of researches explained that CLDN1, CLDN4, and CLDN7 are connected with breast carcinoma [10-12]. Primarily, CLDN1 and CLDN7 influenced tumor dissemination. In addition, CLDN4 was skilled in mammary glandular cell differentiation. Chao Yu-Chih et al discovered that CLDN1 can decrease metastasis in lung adenocarcinoma [13]. CLDN3 and CLDN7 were overexpressed in ovarian cancer cell [14, 15]. Specifically, CLDN3 with low DNA methylation promoted tumorgenesis by Sp1 and CLDN7 could be an antibody-mediated localized therapeutic site.
With the development of sequencing technology, we discovered that CLDN10 is upregulated in tumor tissues in our unpublished research. CLDN10 is a coding protein belonging to the claudin family. Aldred Micheala A et al. firstly demonstrated that CLDN10 is overexpressed in PTC and is one of five genes can separate PTC from follicular thyroid cancer (FTC) [16]. CITED1 and insulin-like growth factor binding protein 6 (IGFBP6) expression level were similar to CLDN10. But the specific biological role and function of CLDN10 in PTC are still unknown.
In our study, we obtained the expression level of CLDN10 in PTC tissues and cell lines by qRT-PCR. We detected that CLDN10 can influence cellular proliferation, migration, and invasion in vitro. According to these results, we certified CLDN10 is overexpressing in PTC again and it acts as a tumor promoter. CLDN10 might become a site for therapy in PTC in the future.
Materials and Methods
Tissue samples
All 55 tumor PTC tissues and matched adjacent tissues were gained during the surgery after patients' agreements at The First Affiliated Hospital of Wenzhou Medical University in recent years. And it got the consent of The Institute Research Medical Ethics Committee of The First Affiliated Hospital of Wenzhou Medical University. The fresh tissues were snap-frozen in liquid nitrogen immediately and stored at -80 °C until further RNA detection.
Cell culture
Professor Mingzhao Xing of the Johns Hopkins University School of Medicine, Baltimore, MA, USA provided us the PTC cell lines (B-CPAP and KTC-1). The two cell lines were cultured in RPMI 1640 (Invitrogen, Carlsbad, CA, USA) having 10% fetal bovine serum (FBS; Invitrogen). The environment of culture was at 37℃ with 5% CO2 in a standard cell culture incubator (Thermo, Waltham, MA, USA).
Cell transfection
Shanghai Gene Pharma (Shanghai, China) designed and made the small interfering RNA (siRNA) targeting CLDN10. 100,000 (BCPAP), 80,000 cells (KTC1) or 60,000cells (TPC1) were plated in the 6-well plate. And we conducted the transfection after 24h. CLDN10 was knocked down using 10μl (TPC1), 7.5μl (KTC1) or 5μl (BCPAP) siRNA and joining the RNAiMAX (Life Technologies, Carlsbad, CA, USA). 4μl (TPC1), 3μl (KTC1) or 2μl (BCPAP). After 48 hours, we could do other experiments. CLDN10 siRNAs target the following sequences: si-CLDN10 #1, Forward: 5'- GCUGGUUGUUGUUGAAUAATT -3' and Reverse: 5'- UUAUUCAACAACAACCAGCTT-3'; si-CLDN10 #2, Forward: 5'- CCGUGAGGAUGAGCAGUUUTT-3' and Reverse: 5'-AAACUGCUCAUCCUCACGGTT -3'.
Quantitative real-time polymerase chain reaction (qRT-PCR)
Total RNA was using TRIZOL Reagent (Invitrogen) extracted from papillary thyroid cancer specimens and cells. Then RNA was reversed to the cDNA which runs the qRT-PCR with THUNDERBIRD SYBR qPCR Mix (Toyobo, Osaka, Japan) on Applied Biosystems 7500. The relative expression level was showed using the comparative cycle threshold (CT) (2-ΔΔCt) and the GAPHD as the endogenous control. The primer sequences were as follows: CLDN10 Forward: 5'- CTGTGGAAGGCGTGCGTTA-3' and Reverse: 5'- CAAAG AAGCC CAGGC TGACA -3'; GADPH Forward: 5'-GTCTCCTCTGACTTCAACAGCG-3' and Reverse: 5'-ACCACCCTGTTGCTGTAGCCAA-3'.
Colony formation assay
48 hours after transfection, putting B-CPAP (2,000 cells/wells), KTC-1 (1,500 cells/wells) and TPC-1 (1,000 cells/wells) in 6-well plates. Anchoring by 4% paraformaldehyde (PFA) and staining by Crystal violet dye approximately seven days culture. Performing it in triplicate and finally taking photos by camera.
CCK-8 proliferation assay
Cell-counting kit 8 (CCK8) can measure the number of cells. Seeding B-CPAP (2,000cells/wells), KTC-1 (1,500 cells/wells) and TPC-1 (1,000 cells/wells) in 96-well plates after transfection. After cells adhered, adding 10μl CCK8 every well and culturing 2-4 hours at 37℃ in the dark. Next, the absorbance was measured at 450 nm by a microplate reader. This experiment should take in four consecutive days.
Cell migration and invasion assays
For the migration assay, digesting the cells by trypsin and then put the cells (30,000 cells/well) into upper chamber with 1640 medium containing 10% FBS. The lower chamber was filled with the 600μl medium the same as upper chamber but without cells. Culturing around 21-24 hours, the membrane was washed by PBS, fixed with 4% PFA for 15 min and stained with 0.4% crystal violet solution for 30min. Wiping the crystal violet solution and selecting views under an inverted microscope at a magnification of x20. The invasion assay is similar to the migration assay.
Statistical analysis
All statistical analyses were using SPSS 22.0 software (SPSS Inc. Chicago, IL, USA). Student' st-test was used between a group with a value of P<0.05 which means the difference is statistically significant. The results were presented as mean ± SD. Odds ratios (ORs) and 95% confidence intervals (CIs) were calculated by the logistic regression model.
Results
CLDN10 is up-regulated in both PTC tissues and cell lines
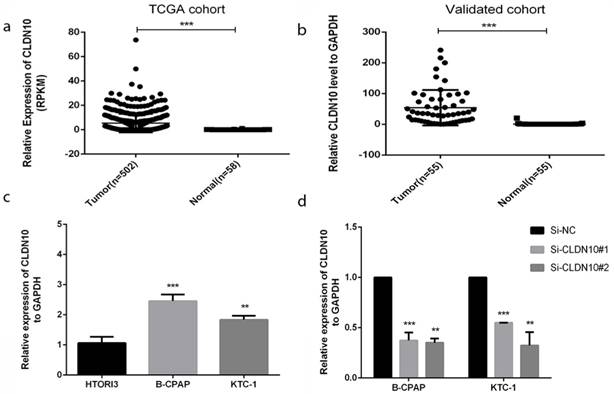
In order to realize the role of CLDN10 in PTC, we sought it in The Cancer Genome Atlas (TCGA) (Fig.1a) and found that the expression in tumor tissues is much larger than in normal tissues. To confirm this discovery, we measured the expression level in 55 tumors PTC tissues and matched adjacent tissues by qRT-PCR. As is shown in Fig.1b, the expression of CLDN10 in PTC tissues is higher than matched adjacent tissues (p<0.001). In addition, we continued to test the expression level in normal thyroid cell line (HTORI3) and PTC cell lines (BCPAP and KTC-1) by qRT-PCR. As Fig.1c shows, we noticed that CLDN10 is highly expressed in PTC cell lines rather than normal thyroid cell line (p<0.01). These results demonstrated that CLDN10 is up-regulated in PTC tissues and cell lines.
Thus, we chose BCPAP and KTC-1 to further experiments. We used the small interfering RNAs (siRNAs) (Si-NC, Si-CLDN10#1 and Si-CLDN10#2) to knock down the CLDN10 expression in BCPAPA and KTC-1 cells. Obviously, siRNA could knock off more than half (Fig.1d).
The relationship between CLDN10 expression and clinicopathologic features
In order to know if the CLDN10 expression is related to tumorigenesis and progression of PTC, we counted the clinicopathologic features of CLDN10. Firstly, we collected 55 clinicopathologic information of PTC patients. Next, we divided the patients into low-expression group (n =27; T/N<130) and high expression group (n =28; T/N > 130) on the basis of CLDN10 expression. The results from the validation cohort showed that the relationship of CLDN10 and lymph node metastasis is more closely than other features (Table 1). Furthermore, we obtained clinicopathologic information of PTC from TCGA and we divided the patients with PTC into low and high expression groups in accordance with the CLDN10 expression of the median value. The analysis indicated that CLDN10 expression is prominently associated with age (P=0.012<0.05), histological type (P=0.000 < 0.05) and lymph node metastasis (P=0.000 <0.05) (Table 2). But the remains such as gender, tumor size, extrathyroidal invasion and the like we can't demonstrate their relationship with CLDN10 expression.
The relationship between CLDN10 and clinicopathologic characteristics in validated cohort.
| Clinicopathologic characteristics | Low expression (%) | High expression (%) | X2 | P |
|---|---|---|---|---|
| Age | 0.150 | 0.698 | ||
| ≤45 | 12(44.4) | 11(30.3) | ||
| >45 | 15(55.6) | 17(69.7) | ||
| Gender | 4.199 | 0.400 | ||
| Female | 13(48.1) | 21(75) | ||
| Male | 14(51.9) | 7(25) | ||
| Tumor size | 0.034 | 0.853 | ||
| ≤10mm | 9(33.3) | 10(35.7) | ||
| >10mm | 18(66.7) | 18(64.3) | ||
| Unilateral or Bilateral | 0.181 | 0.671 | ||
| Unilateral | 11(40.7) | 13(46.4) | ||
| Bilateral | 16(59.3) | 15(53.6) | ||
| Extrathyroidal invasion | 0.392 | 0.531 | ||
| YES | 2(7.4) | 1(3.6) | ||
| NO | 25(92.6) | 27(96.4) | ||
| Lymph node metastasis | 3.104 | 0.078* | ||
| YES | 13(48.1) | 20(71.4) | ||
| NO | 14(51.9) | 8(28.6) | ||
| AJCC | 0.437 | 0.509 | ||
| I+II | 15(55.6) | 18(64.3) | ||
| III+IV | 12(44.4) | 10(35.7) | ||
* P-value < 0.05
The relationship between CLDN10 and clinicopathologic characteristics in TCGA cohort.
| Clinicopathologic characteristics | Low expression (%) | High expression (%) | X2 | P |
|---|---|---|---|---|
| Age | 6.269 | 0.012* | ||
| ≤45 | 104(41.4) | 132(52.6) | ||
| >45 | 147(58.6) | 119(47.4) | ||
| Gender | 0.01 | 0.92 | ||
| Female | 184(73.3) | 183(72.9) | ||
| Male | 67(26.7) | 68(27.1) | ||
| Histological type | 91.03 | 0.000* | ||
| Classical | 148(59.0) | 208(83.2) | ||
| Follicular | 92(36.7) | 9(3.6) | ||
| Tall Cell | 7(2.8) | 28(11.2) | ||
| other | 4(1.6) | 5(2.0) | ||
| Tumor size | 1.653 | 0.199 | ||
| ≤20mm | 78(31.1) | 65(25.9) | ||
| >20mm | 173(68.9) | 186(74.1) | ||
| T stage | 1.653 | 0.199 | ||
| T1 | 78(31.1) | 65(25.9) | ||
| >T1 | 173(68.9) | 186(74.1) | ||
| Extrathyroidal invasion | 0.113 | 0.737 | ||
| YES | 5(2) | 4(1.6) | ||
| NO | 246(98) | 247(98.4) | ||
| Lymph node metastasis | 52.938 | 0.000* | ||
| YES | 71(28.3) | 152(60.6) | ||
| NO | 180(71.7) | 99(39.4) | ||
| AJCC | 1.079 | 0.299 | ||
| I+II | 172(68.5) | 161(64.1) | ||
| III+IV | 79(31.5) | 90(35.9) | ||
* P-value < 0.05
To get further knowledge of risk factors of lymph node metastasis, we used Univariate logistic regression analysis and Multivariate logistic regression analysis. Univariate logistic regression analysis showed that CLDN10 expression, age, gender, primary neoplasm focus type and tumor size are all risk factors of lymph node metastasis (P<0.05) (Table 3). As a consequence, we selected these factors for Multivariate logistic regression analysis. Table 4 showed that CLDN10 expression, tumor size and age influence the occurrence of lymph node metastasis. Mostly, CLDN10 expression promoted lymph node metastasis (OR=3.079).
Univariate logistic regression analysis for the lymph node metastatic risk.
| Clinicopathologic features | OR | 95% CI | P |
|---|---|---|---|
| CLDN10 expression | 3.892 | 2.679-5.655 | 0.000* |
| Age | 0.652 | 0.458-0.93 | 0.018* |
| Gender | 1.569 | 1.055-2.333 | 0.026* |
| Primary Neoplasm Focus Type | 1.483 | 1.037-2.122 | 0.031* |
| Tumor size | 2.237 | 1.483-3.375 | 0.000* |
* P-value < 0.05.
Multivariate logistic regression analysis for the lymph node metastatic risk
| Clinicopathologic features | OR | 95% CI | P |
|---|---|---|---|
| CLDN10 expression | 3.079 | 2.514-5.472 | 0.000* |
| Age | 0.673 | 0.456-0.673 | 0.046* |
| Gender | 1.509 | 0.972-2.344 | 0.067 |
| Primary Neoplasm Focus Type | 1.531 | 1.034-2.268 | 0.071 |
| Tumor size | 2.253 | 1.438-3.529 | 0.014* |
* P-value < 0.05
Knockdown of CLDN10 restrains proliferation in PTC cell lines
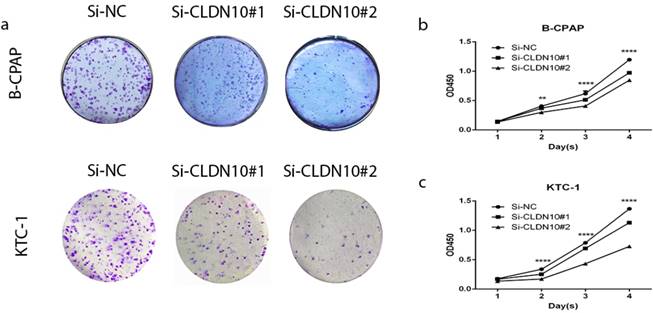
BCPAP and KTC-1 were selected as representtative of PTC cell lines to explore the function of CLDN10. To prove if knockdown of CLDN10 represses the proliferation in BCPAP and KTC-1, colony formation assay and CCK-8 assay were applied. In fig 2a, cell colony formation was significantly inhibited in BCPAP and KTC-1 by knocking down CLDN10 expression compared with control. And further CCK-8 assay was inhibited in PTC cell lines by knocking down CLDN10 expression compared with control (fig 2b and c). Colony formation assay and CCK-8 assay indicated that knockdown of CLDN10 restrains proliferation in BCPAP and KTC-1 cells over time (Fig.2a-2d).
Knockdown of CLDN10 represses migration and invasion in PTC cell lines
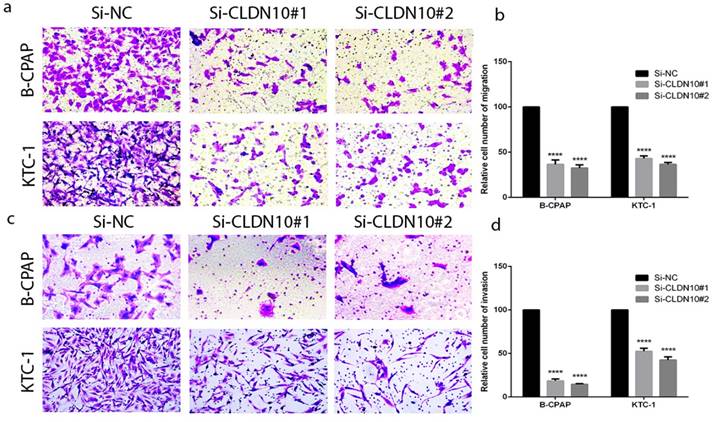
According to the relationship of CLDN10 expression and clinicopathologic features, lymph node metastasis is a significant feature. To further explore the migratory and invasive abilities of CLDN10 in PTC, we did the Transwell assays in BCPAP and KTC-1. As predicted, the migratory ability of siCLDN10 cells was decreased compared with that of si-NC cells. The same phenomenon also occurs in invasion assay (Fig.3a-3d). Transwell assays indicated that CLDN10 knockdown suppressed migration and invasion of BCPAP and KTC-1 cells compared with Si-NC group in vitro.
CLDN10 is up-regulated in human PTC tissues and cell lines. (a) Relationship of expression levels of CLDN10 in thyroid tumor tissues and normal tissues in TCGA. (b) CLDN10 is significantly increased in 55 human PTC tissues in comparison to matched adjacent tissues. (c) The relative expression of CLDN10 to GAPDH using qRT-PCR. Both B-CPAP and KTC-1 cell lines are overexpressed. (d) The efficiency of siRNAs (Si-NC, Si-CLDN10#1 and Si-CLDN10#2) was assayed by qRT-PCR in B-CPAPA and KTC-1 cells. *P < .05; **P < .01; ***P < .001 in comparison with the control group using Student's t-test. 2-ΔΔCt is used to represent the fold change in qRT-PCR detection.
Effect of CLDN10 on proliferation in PTC cell lines. (a) Colon formation assay indicated that knockdown of CLDN10 represses cell proliferation in B-CPAPA and KTC-1 cells. (b, c) CCK-8 assay indicated that knockdown of CLDN10 represses cell proliferation in B-CPAP and KTC-1 cells. **P < .01; ****P < .0001 in comparison with the Si-NC group using Student's t-test.
CLDN10 regulates PTC cells migration and invasion. (a, b) Migration assay in B-CPAP and KTC-1 showed that knockdown of CLDN10 decrease the cell migration compared with that in Si-NC group. (c, d) knockdown of CLDN10 decrease the cell invasion compared with that in Si-NC group. ****P < .0001 in comparison with the Si-NC group using Student's t-test.
Discussion
Papillary thyroid cancer has an extremely high prevalence, but the 30-year survival rate is more than 90% if patients receive regular therapy such as thyroidectomy [17]. For all this, PTC still recurs easily after conventional therapy. Therefore, many researchers explore targeted therapy for accurate treatment. Lei Hongwei et al. found that lncRNA TUG1 influences PTC cell proliferation, migration and EMT formation by miR-145 [18]. Huang Lei-Lei et al. discovered that AEG-1 is associated with metastasis in PTC by MMP2/9 [19]. Wang Xiaoming et al. detected that lncRNA PTCSC3 has a relationship with tumor progression in PTC via Wnt/β-Catenin Signaling [20].
Claudin family has been found playing an important role in tumorigenesis and becoming emerging targets for therapy [9, 21]. CLDN6 and CLDN18.2 were in adenocarcinomas and large-cell carcinomas which could become a potential therapeutic target in non-small-cell lung cancer [22]. And claudin-4 brought cytotoxic effect and also could be therapy target for prostate cancer [23]. CLDN10 was connected with the recurrence of primary hepatocellular carcinoma [24] and progression of gastric cancer [25]. But the specific function of CLDN10 and if it can be a novel therapeutic target in PTC is unknown.
Our study confirmed that CLDN10 is overexpressed in PTC tissues and cell lines. And the relationship of CLDN10 expression and clinicopathologic features analyzed by Kaplan-Meier analysis and logistic regression analysis in TCGA expounded that CLDN10 is conspicuously relevant with age and histological type, especially lymph node metastasis. In vitro, knockdown of CLDN10 could influence the cellular proliferation, migration, and invasion in B-CPAP and KTC-1. All in all, CLDN10 could affect tumorigenesis in PTC.
However, our study remains many inadequacies. On the one hand, we should obtain more PTC tissues and do an experiment in vivo to replenish our study thus become convective. On the other hand, the concrete mechanism of CLDN10 in PTC is still to be found.
Conclusion
In summary, we proved that CLDN10 up-regulated in PTC tissues and cell lines. And clinicopathologic analysis testified that lymph node metastasis is the most important feature of CLDN10. We also found that knockdown of CLDN10 restrains cellular proliferation, migration, and invasion in vitro. These results indicated that CLDN10 plays a vital role in tumorigenesis in PTC and can be a site in targeted therapy.
Acknowledgements
This study was funded by National Natural Science Foundation of China (NO.81572291) and Natural Science Foundation of Zhejiang province (LY17H160053, LGF18H160031, GF18H160071, and LGF18H160032) and the Medical and Health Technology Projects of Zhejiang province (NO.2017187475) and the Science and Technology Project of Wenzhou (Y20170030).
Informed consent
Written informed consent was obtained from each individual participant.
Ethical approval
Ethical approval for this study was obtained from the Ethics Committee of the First Affiliated Hospital of Wenzhou Medical University.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Siegel RL, Miller KD, Jemal A. Cancer statistics, 2018. CA Cancer J Clin. 2018;68:7-30
2. Burns WR, Zeiger MA. Differentiated thyroid cancer. Semin Oncol. 2010;37:557-66
3. Hinson AM, Massoll NA, Jolly LA, Stack BC Jr, Bodenner DL, Franco AT. Structural alterations in tumor-draining lymph nodes before papillary thyroid carcinoma metastasis. Head Neck. 2017;39:1639-46
4. Ansari M, Babaei AA, Shafiei B, Javadi H, Assadi M, Nabipour I. et al. Pathological evaluation of differentiated thyroid cancer in patients with positive serum thyroglobulin and negative iodine scan. Eur Rev Med Pharmacol Sci. 2014;18:1925-9
5. Hay ID, Thompson GB, Grant CS, Bergstralh EJ, Dvorak CE, Gorman CA. et al. Papillary thyroid carcinoma managed at the Mayo Clinic during six decades (1940-1999): temporal trends in initial therapy and long-term outcome in 2444 consecutively treated patients. World J Surg. 2002;26:879-85
6. Mineta K, Yamamoto Y, Yamazaki Y, Tanaka H, Tada Y, Saito K. et al. Predicted expansion of the claudin multigene family. FEBS Lett. 2011;585:606-12
7. Tsukita S, Furuse M. Pores in the wall: claudins constitute tight junction strands containing aqueous pores. J Cell Biol. 2000;149:13-6
8. Zhang L, Feng T, Spicer LJ. The role of tight junction proteins in ovarian follicular development and ovarian cancer. Reproduction. 2018;155:R183-R98
9. Swisshelm K, Macek R, Kubbies M. Role of claudins in tumorigenesis. Adv Drug Deliv Rev. 2005;57:919-28
10. Kominsky SL, Argani P, Korz D, Evron E, Raman V, Garrett E. et al. Loss of the tight junction protein claudin-7 correlates with histological grade in both ductal carcinoma in situ and invasive ductal carcinoma of the breast. Oncogene. 2003;22:2021-33
11. Tokes AM, Kulka J, Paku S, Szik A, Paska C, Novak PK. et al. Claudin-1, -3 and -4 proteins and mRNA expression in benign and malignant breast lesions: a research study. Breast Cancer Res. 2005;7:R296-305
12. Kominsky SL, Vali M, Korz D, Gabig TG, Weitzman SA, Argani P. et al. Clostridium perfringens enterotoxin elicits rapid and specific cytolysis of breast carcinoma cells mediated through tight junction proteins claudin 3 and 4. Am J Pathol. 2004;164:1627-33
13. Chao YC, Pan SH, Yang SC, Yu SL, Che TF, Lin CW. et al. Claudin-1 is a metastasis suppressor and correlates with clinical outcome in lung adenocarcinoma. Am J Respir Crit Care Med. 2009;179:123-33
14. Honda H, Pazin MJ, D'Souza T, Ji H, Morin PJ. Regulation of the CLDN3 gene in ovarian cancer cells. Cancer Biol Ther. 2007;6:1733-42
15. Tassi RA, Bignotti E, Falchetti M, Ravanini M, Calza S, Ravaggi A. et al. Claudin-7 expression in human epithelial ovarian cancer. Int J Gynecol Cancer. 2008;18:1262-71
16. Aldred MA, Huang Y, Liyanarachchi S, Pellegata NS, Gimm O, Jhiang S. et al. Papillary and follicular thyroid carcinomas show distinctly different microarray expression profiles and can be distinguished by a minimum of five genes. J Clin Oncol. 2004;22:3531-9
17. Markovina S, Grigsby PW, Schwarz JK, DeWees T, Moley JF, Siegel BA. et al. Treatment approach, surveillance, and outcome of well-differentiated thyroid cancer in childhood and adolescence. Thyroid. 2014;24:1121-6
18. Lei H, Gao Y, Xu X. LncRNA TUG1 influences papillary thyroid cancer cell proliferation, migration and EMT formation through targeting miR-145. Acta Biochim Biophys Sin (Shanghai). 2017;49:588-97
19. Huang LL, Wang Z, Cao CJ, Ke ZF, Wang F, Wang R. et al. AEG-1 associates with metastasis in papillary thyroid cancer through upregulation of MMP2/9. Int J Oncol. 2017;51:812-22
20. Wang X, Lu X, Geng Z, Yang G, Shi Y. LncRNA PTCSC3/miR-574-5p Governs Cell Proliferation and Migration of Papillary Thyroid Carcinoma via Wnt/beta-Catenin Signaling. J Cell Biochem. 2017;118:4745-52
21. Kominsky SL. Claudins: emerging targets for cancer therapy. Expert Rev Mol Med. 2006;8:1-11
22. Micke P, Mattsson JS, Edlund K, Lohr M, Jirstrom K, Berglund A. et al. Aberrantly activated claudin 6 and 18.2 as potential therapy targets in non-small-cell lung cancer. Int J Cancer. 2014;135:2206-14
23. Maeda T, Murata M, Chiba H, Takasawa A, Tanaka S, Kojima T. et al. Claudin-4-targeted therapy using Clostridium perfringens enterotoxin for prostate cancer. Prostate. 2012;72:351-60
24. Cheung ST, Leung KL, Ip YC, Chen X, Fong DY, Ng IO. et al. Claudin-10 expression level is associated with recurrence of primary hepatocellular carcinoma. Clin Cancer Res. 2005;11:551-6
25. Gao M, Li W, Wang H, Wang G. The distinct expression patterns of claudin-10, -14, -17 and E-cadherin between adjacent non-neoplastic tissues and gastric cancer tissues. Diagn Pathol. 2013;8:205
Author contact
![]() Corresponding authors: Dr. Yinghao Wang, Department of Thyroid and Breast Surgery, The First Affiliated Hospital of Wenzhou Medical University, Wenzhou, Zhejiang, People's Republic of China, 325000. Contact number: +86 577 5557 8527; Email: yinghao_necom and Dr. Ouchen Wang, Department of Thyroid and Breast Surgery, The First Affiliated Hospital of Wenzhou Medical University, Wenzhou, Zhejiang, People's Republic of China, 325000. Contact number: +86 577 5557 8527; Email: woc863com
Corresponding authors: Dr. Yinghao Wang, Department of Thyroid and Breast Surgery, The First Affiliated Hospital of Wenzhou Medical University, Wenzhou, Zhejiang, People's Republic of China, 325000. Contact number: +86 577 5557 8527; Email: yinghao_necom and Dr. Ouchen Wang, Department of Thyroid and Breast Surgery, The First Affiliated Hospital of Wenzhou Medical University, Wenzhou, Zhejiang, People's Republic of China, 325000. Contact number: +86 577 5557 8527; Email: woc863com

 Global reach, higher impact
Global reach, higher impact