3.2
Impact Factor
ISSN: 1837-9664
J Cancer 2019; 10(6):1453-1465. doi:10.7150/jca.28919 This issue Cite
Research Paper
Genetic variants and Expression of Cytochrome p450 Oxidoreductase Predict Postoperative Survival in Patients with Hepatitis B Virus-Related Hepatocellular Carcinoma
1. Department of Hepatobiliary Surgery, The First Affiliated Hospital of Guangxi Medical University, Nanning, 530021, Guangxi Zhuang Autonomous Region, People's Republic of China;
2. Department of Hepatobiliary Surgery, Affiliated Hospital of Guangdong Medical University, Zhanjiang, 524001, Guangdong Province, People's Republic of China;
3. Department of Hepatobiliary and Pancreatic Surgery, The First Affiliated Hospital of Zhengzhou University, Zhengzhou, 450000, Henan Province, People's Republic of China;
4. Department of Hepatobiliary Surgery, The Third Affiliated Hospital of Guangxi Medical University, Nanning, 530031, Guangxi Zhuang Autonomous Region, People's Republic of China;
5. Key Laboratory of Combined Multi-Organ Transplantation, Ministry of Public Health and Key Laboratory of Organ Transplantation of Zhejiang Province, Hangzhou, 310003, Zhejiang Province, People's Republic of China;
6. Science for Life Laboratory, KTH-Royal Institute of Technology, Stockholm, SE-171 21, Sweden;
7. Department of Hepatobiliary Surgery, The Sixth Affiliated Hospital of Guangxi Medical University, Yulin, 537000, Guangxi Zhuang Autonomous Region, People's Republic of China.
*These authors contributed equally to this work.
Received 2018-8-1; Accepted 2018-12-14; Published 2019-2-23
Abstract
Our current study investigates the prognostic values of genetic variants and mRNA expression of cytochrome p450 oxidoreductase (POR) in hepatitis B virus (HBV)-related hepatocellular carcinoma (HCC). A total of 19 candidate single nucleotide polymorphisms (SNPs) located in the exons of POR were genotyped using Sanger sequencing from 476 HBV-related HCC patients who underwent hepatectomy between 2003 and 2013. The mRNA expression of POR in 212 patients with HBV-related HCC was obtained from GSE14520 dataset. Survival analysis was performed to investigate the association of POR variants and mRNA expression with overall survival (OS) and recurrence-free survival (RFS). Nomograms were used to predict the prognosis of HBV-related HCC patients. Gene set enrichment analysis (GSEA) was used to investigate the mechanism of POR in HBV-related HCC prognosis. The polymorphism POR-rs1057868 was significantly associated with HBV-related HCC OS (CT/TT vs. CC, hazard ratio [HR] = 0.69, 95% confidence interval [CI] = [0.54, 0.88], P = 0.003), but not significantly associated with RFS (CT/TT vs. CC, P = 0.378). POR mRNA expression was also significantly associated with HBV-related HCC OS (high vs. low, HR = 0.61, 95% CI = [0.38, 0.97], P = 0.036), but not significantly associated with the RFS (high vs. low, P = 0.201). Two nomograms were developed to predict the HBV-related HCC OS. Furthermore, GSEA suggests that multiple gene sets were significantly enriched in liver cancer survival and recurrence, as well as POR-related target therapy in the liver. In conclusion, our study suggests that POR-rs1057868 and mRNA expression may serve as a potential postoperative prognosis biomarker in HBV-related HCC.
Keywords: hepatocellular carcinoma, cytochrome p450 oxidoreductase, prognosis, hepatitis B virus, hepatectomy
Introduction
Liver cancer is the seventh most common cancer and the third leading cause of cancer-related death worldwide, and approximately 70%-90% of primary liver cancer is hepatocellular carcinoma (HCC) [1, 2]. Hepatitis B virus (HBV) promotes cirrhosis, which is found in approximately half of the patients with HCC worldwide and this rate is higher in China[3]. Although the rapid development of a comprehensive surgery-based treatment has improved the clinical outcome for HCC patients greatly, the long-term clinical outcome after hepatectomy of HCC remains unsatisfactory[4, 5]. Multiple clinical indicators of HCC such as tumor size, vascular invasion, portal vein tumor thrombus (PVTT), serum alpha-fetoprotein (AFP) levels, Barcelona Clinic Liver Cancer (BCLC) stage, and antiviral therapy have been indicated as useful predictors for HCC patient prognosis after hepatectomy[6-10]. However, these factors still cannot accurately predict HCC prognosis. Considering that prognosis might be affected by genetic and clinicopathological factors, it would be rational to identify potential biomarkers for a more effective prognosis prediction, thus improving clinical outcomes in HCC patients.
During the past few years, an increasing number of genes that encode metabolic enzymes have been confirmed as tumor susceptibility genes[11, 12]. Cytochrome p450 oxidoreductase (POR) encodes a metabolic enzyme, which is expressed extensively in multiple normal and tumor tissues. POR is known as the unique electron donor for all microsomal cytochrome p450 (CYP) enzymes, which play a critical role in the metabolism of steroid hormones, fatty acids, bile acids, and drugs[13, 14]. Variants in POR have also been reported as risk factors in multiple cancers. Haiman et al. demonstrated that synonymous variants of POR G5G might increase the risk of breast cancer in African American populations[15]. Xiao et al. reported that the POR variant A503V is correlated with a risk of bladder cancer in a Chinese population[16]. However, the association of POR variants and expression with HCC prognosis remains unclear. Therefore, our current study is to investigate the prognostic values of genetic variants and expression of POR in HBV-related HCC prognosis after hepatectomy.
Materials and Methods
Study population
This study was approved by the Ethics Committee of The First Affiliated Hospital of Guangxi Medical University (Approval number: 2015[KY-E-032]). The inclusion criteria of patients in the present study were listed as follows: 1) Chinese population; 2) Patients with HBV-related HCC; 3) Patients underwent hepatectomy. The exclusion criteria of patients in the present study were listed as follows: 1) Patients with missing clinical features and prognostic data; 2) Patients survived less than one month. To explore the application value of POR genetic variation in prognosis of HCC, a total of 476 HBV-related HCC patients who underwent hepatectomy between 2003 and 2013 in The First Affiliated Hospital of Guangxi Medical University, Guangxi, China, and identified as cohort 1. To explore the application value of POR mRNA expression in prognosis of HCC, by retrieving the public database, only the whole genome expression profile data set of GSE14520 compliance with the above inclusion conditions, and identified as cohort 2. Cohort 2 consisted of 212 HBV-related HCC patients who underwent hepatectomy from the GSE14520 dataset (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE14520)[17, 18].
Clinical data and definitions
The clinical features were retrospectively collected from medical records. HBV-related HCC patients were defined as patients with a seropositive of the HBV surface antigen (HBsAg) and with a postoperative sample pathological diagnosis of HCC. Tumor stage was classified according to the BCLC staging system[19]. PVTT was classified according to the classification system of the Liver Cancer Study Group of Japan[20]. Liver reserve function was measured according to the Child-Pugh score[21]. Preoperative AFP was classified into two groups with a cut-off value of 400 ng/ml[22]. The degree of cirrhosis was determined according to the pathological report. Antiviral therapy is defined here as receiving anti-HBV treatment systematically after surgery. Recurrence-free survival (RFS) was defined as the interval between surgery and the date of diagnosis of the first recurrence or the date of the last follow-up. Overall survival (OS) was defined as the interval between surgery and the date of death, or the date of the last follow-up.
Follow-up
The patients were followed up via telephone or hospital visit after surgery until patients died or up to the date of the last follow-up. The last follow-up was in January 2018.
Identification of POR variants
The 19 previously reported candidate single nucleotide polymorphisms (SNPs) located in the exons of POR were selected to investigate the prognostic values in HBV-related HCC. All SNPs were genotyped in HCC tumor tissues using the Sanger sequencing method[23]. The primers used for amplification of the POR SNPs were according to a previous study (Supplementary Table S1)[16]. As shown in supplementary Table S2, two POR gene variants (rs1135612, rs1057868) were observed to have a polymorphic distribution in the cohort 1 population. The SNPs with a minor allele frequency (MAF) < 10% were excluded from the present study, and only the data of rs1135612 and rs1057868 were included in further study. The MAF of rs1135612 and rs1057868 was 43.8% and 35.9%, respectively. Both of these SNPs met the Hardy-Weinberg equilibrium (rs1135612: 0.619; rs1057868: 0.378). Rs1135612 is a synonymous mutation (P130P) and rs1057868 is a missense mutation (A503V).
Gene expression profile
The gene expression dataset was obtained from the Gene Expression Omnibus public database with a serial number of GSE14520[17, 18]. In order to avoid the batch effect between different platforms, only the dataset of the Affymetrix HT Human Genome U133A Array (GPL3921) of GSE14520 was included in the present study. Processing of raw data was according to the manufacturer's guidelines and shown in https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSM362960. For multiple probe sets, the average value corresponding to the same gene was regarded as the expression value of the gene and normalized by the limma package in the R platform. Only 212 HBV-related HCC patients who underwent hepatictomy were analyzed further in the present study.
Gene set enrichment analysis
To investigate the potential mechanism of POR in HCC prognosis, a gene set enrichment analysis (GSEA, https://software.broadinstitute.org/gsea/) [24] was conducted using the gene expression dataset of GSE14520 based on the molecular signature database (MSigDB) of c2 (c2.all.v6.1.symbols.gmt) and c5 (c5.all.v6.1.symbols.gmt) gene sets[25]. A P-value < 0.05 and false discovery rate (FDR) < 0.25 were considered statistically significant.
Statistical analysis
Survival was calculated using the Kaplan- Meier method and compared with a log-rank test. Cox proportional hazards model was used to identify independent predictors of survival. All clinical features with a P-value < 0.1 in the univariate analysis were filtered into the multivariate analysis. A nomogram was developed to predict the survival based on Cox regression. The significant factors associated with prognosis in the multivariate analysis were fitted into nomograms. The nomogram was validated using the concordance index (c-index)[26]. All analyses were performed using R version 3.3.2 (http://www.R-project.org). A P-value < 0.05 was considered statistically significant.
Results
Characteristics of the study population
Cohort 1 consisted of 476 HBV-related HCC patients from The First Affiliated Hospital of Guangxi Medical University. The median age was 46 years. The relationship between POR variants and clinicopathological factors are summarized in Table 1. No significant correlation with rs1057868 and rs1135612 was found with gender, age, smoking, drinking, Child-Pugh, AFP, PVTT, BCLC, cirrhosis or antiviral therapy (all P> 0.05). The OS follow-up rate was 100%, while the RFS follow-up rate was 72%.
Cohort 2 consisted of 212 HBV-related HCC patients from GSE14520. The median age was 50 years. The patients were divided into low- and high-expression groups according to the median value of POR mRNA expression. The relationship between POR expression and clinicopathological factors are summarized in Table 2. No significant correlation was found with gender, age, alanine aminotransferase (ALT), and cirrhosis (all P > 0.05). However, POR expression was significantly associated with gender (P = 0.046), AFP (P< 0.001), and BCLC (P = 0.023). Both OS and RFS follow-up rate were 100%.
The relationship between POR variants and clinicopathological factors in the cohort 1.
| Variable | rs1057868 | P-value* | rs1135612 | P-value* | |||||
|---|---|---|---|---|---|---|---|---|---|
| CC | CT | TT | AA | AG | GG | ||||
| Gender | Male | 172 | 199 | 50 | 0.665 | 137 | 199 | 85 | 0.584 |
| Female | 19 | 29 | 7 | 16 | 30 | 9 | |||
| Age | ≤46years | 107 | 126 | 26 | 0.360 | 78 | 125 | 56 | 0.419 |
| >46years | 84 | 102 | 31 | 75 | 104 | 38 | |||
| Smoking | No | 122 | 157 | 34 | 0.330 | 94 | 160 | 59 | 0.186 |
| Yes | 69 | 71 | 23 | 59 | 69 | 35 | |||
| Drinking | No | 111 | 143 | 34 | 0.624 | 87 | 144 | 57 | 0.499 |
| Yes | 80 | 85 | 23 | 66 | 85 | 37 | |||
| Child-Pugh | A | 164 | 197 | 52 | 0.563 | 132 | 205 | 76 | 0.110 |
| B | 27 | 31 | 5 | 21 | 24 | 18 | |||
| AFP | <400ng/ml | 98 | 131 | 35 | 0.284 | 87 | 133 | 44 | 0.165 |
| ≥400ng/ml | 93 | 97 | 22 | 66 | 96 | 50 | |||
| PVTT | No | 155 | 195 | 50 | 0.343 | 132 | 191 | 77 | 0.621 |
| vp1-4 | 36 | 33 | 7 | 21 | 38 | 17 | |||
| BCLC | 0/A | 110 | 136 | 35 | 0.945 | 93 | 135 | 53 | 0.900 |
| B | 29 | 37 | 9 | 25 | 36 | 14 | |||
| C | 52 | 55 | 13 | 35 | 58 | 27 | |||
| Cirrhosis | No | 20 | 30 | 7 | 0.699 | 18 | 29 | 10 | 0.874 |
| Yes | 171 | 198 | 50 | 135 | 200 | 84 | |||
| Antiviral | No | 116 | 153 | 39 | 0.326 | 101 | 148 | 59 | 0.874 |
| Yes | 75 | 75 | 18 | 52 | 81 | 35 | |||
Abbreviations: AFP, alpha-fetoprotein; PVTT, portal vein tumor thrombus; BCLC, Barcelona Clinic Liver Cancer stage. Note: *P-value was calculated using two-sided χ2 test.
The relationship between POR expression and clinicopathological factors in the cohort 2.
| Variable | POR expression | P-value* | ||
|---|---|---|---|---|
| Low | High | |||
| Gender | Male | 86 | 97 | 0.046 |
| Female | 20 | 9 | ||
| Age | ≤50years | 51 | 58 | 0.410 |
| >50years | 55 | 48 | ||
| ALT | ≤50U/L | 59 | 65 | 0.486 |
| >50U/L | 47 | 41 | ||
| AFP | ≤300ng/ml | 40 | 75 | <0.001 |
| >300ng/ml | 65 | 29 | ||
| BCLC | 0/A | 74 | 89 | 0.023 |
| B/C | 32 | 17 | ||
| Cirrhosis | No | 6 | 11 | 0.312 |
| Yes | 100 | 95 | ||
Abbreviations: ALT, alanine aminotransferase; AFP, alpha-fetoprotein; BCLC, Barcelona Clinic Liver Cancer stage. Note: *P-value was calculated using two-sided χ2 test.
Association of POR variants with OS
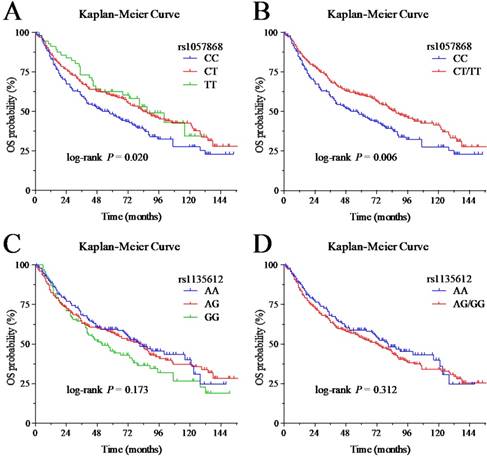
The result of univariate and multivariate analysis of POR variants with OS are presented in Table 3 and Figure 1A-D. The median OS of the POR-rs1057868 CC, CT, and TT group was 57, 84, and 87 months, respectively. In the univariate analysis, we found patients with the POR-rs1057868 CT genotype had better OS than patients with the CC genotype (CT vs. CC, hazard ratio [HR] = 0.73, 95% confidence interval [CI] = [0.56, 0.94], P = 0.014; TT vs. CC, HR = 0.65, 95% CI = [0.43, 0.99], P = 0.041). Furthermore, we found a greater significant difference in the merge model of POR-rs1057868 (CT/TT vs. CC, HR = 0.71, 95% CI = [0.56, 0.91], P=0.006). The median OS of the POR-rs1135612 AA, AG, and GG group was 81, 82, and 52 months, respectively. There was no significant difference among groups with different genotypes of POR-rs1135612 in the univariate analysis (AG vs. AA, GG vs. AA, and AG/GG vs. AA, all P > 0.05).
Kaplan-Meier curves for OS among different POR variants groups. (A) OS stratified by POR-rs1057868 CC, CT and TT genotypes; (B) OS stratified by POR-rs1057868 CC and CT/TT genotypes; (C) OS stratified by POR-rs1135612 AA, AG and GG genotypes; (D) OS stratified by POR- rs1135612 AA and AG/GG genotypes.
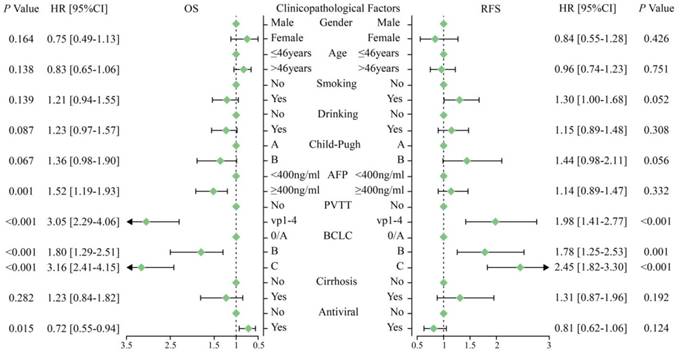
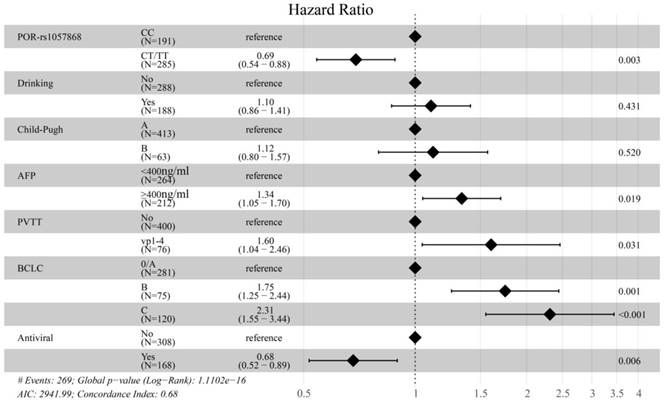
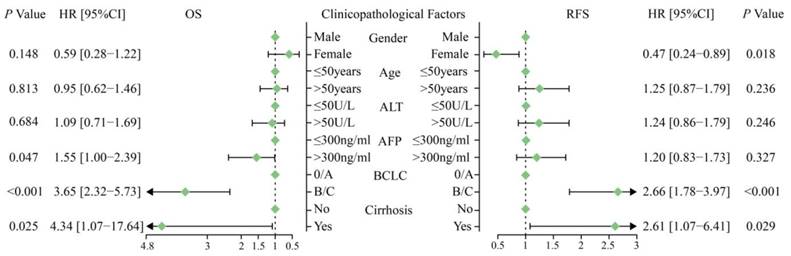
We also performed univariate analysis to investigate the association of clinicopathological factors with OS. As shown in Figure 2, the factors drinking (yes vs. no, P = 0.087), Child-Pugh (B vs. A, P = 0.067), AFP (≥400 ng/ml vs. < 400 ng/ml, P = 0.001), PVTT (vp1-4 vs. no, P < 0.001), BCLC (B vs. 0/A, P < 0.001; C vs. 0/A, P < 0.001), and antiviral therapy (yes vs. no, P = 0.015) with a P-value < 0.1 were entered into the multivariate analysis. As shown in Table 3 and Figure 3, the multivariate analysis, adjusted by drinking, Child-Pugh, AFP, PVTT, BCLC, and antiviral therapy, showed a similar result. Compared with the CC genotype, patients with the CT or TT genotype of POR-rs1057868 had a significant association with a better OS (CT vs. CC, HR = 0.70, 95% CI = [0.54, 0.90], P = 0.006; TT vs. CC, HR = 0.67, 95% CI = [0.45, 1.02], P = 0.062; CT/TT vs. CC, HR = 0.69, 95% CI = [0.54,0.88], P = 0.003). However, there was still no significant association between POR-rs1135612 and OS in the multivariate analysis (AG vs. AA, GG vs. AA, and AG/GG vs. AA, all P > 0.05).
Association of POR variants with RFS
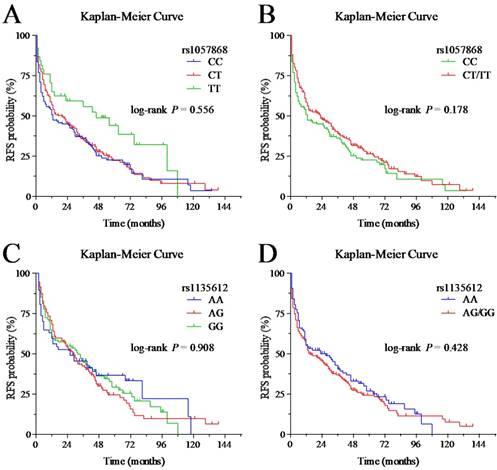
The result of univariate and multivariate analysis of POR variants with RFS was present in Table 4 and Figure 4A-D. The median RFS of the POR-rs1057868 CC, CT, and TT group was 13, 20, and 46 months, respectively. The median RFS of the POR-rs1135612 AA, AG, and GG group was 27, 14, and 16 months, respectively. In the univariate analysis, no significant difference in RFS was found among groups with different genotypes in POR-rs1057868 (CT vs. CC, CT/TT vs. CC, all P > 0.05) and POR-rs1135612 (AG vs. AA, GG vs. AA, and AG/GG vs. AA, all P > 0.05), except in POR-rs1057868 TT vs. CC (P = 0.024).
We also performed univariate analysis to investigate the association of clinicopathological factors with RFS. As shown in Figure 2, the factors smoking (yes vs. no, P = 0.052), Child-Pugh (B vs. A, P = 0.056), PVTT (vp1-4 vs. no, P < 0.001), and BCLC (B vs. 0/A, P = 0.001; C vs. 0/A, P < 0.001) with a P-value < 0.1 were entered into the multivariate analysis. As shown in Table 4, adjusted by smoking, Child-Pugh, PVTT, and BCLC, there was still no significant difference in RFS among groups with different genotypes in POR-rs1057868 (CT vs. CC, TT vs. CC, and CT/TT vs. CC, all P > 0.05) and POR-rs1135612 (AG vs. AA, GG vs. AA, and AG/GG vs. AA, all P > 0.05) in the multivariate analysis.
Association of POR mRNA expression with OS
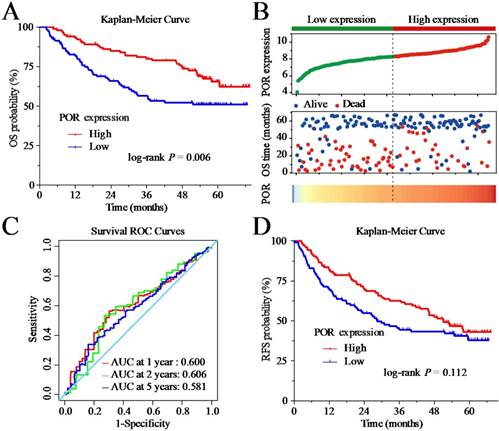
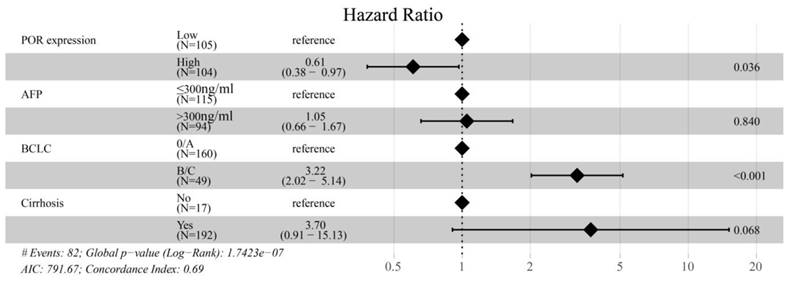
Furthermore, we also investigated the prognostic values of POR mRNA expression by using the 212 HBV-related HCC patients from the GSE14520 dataset. The results of univariate analysis of POR mRNA expression with OS are presented in Table 5 and Figure 5A and B. The median OS of the POR low- and high-expression group were not available because more than half of the patients survived up to the end of the follow-up. In the univariate analysis, we found patients with higher mRNA expression of POR in HBV-related HCC had better OS than the patients with lower expression (high vs. low: HR = 0.54, 95% CI = [0.35, 0.84], P =0.006). Time-dependent receiver operating characteristic (ROC) analysis substantiated that the POR mRNA expression showed a good performance in the HBV-related HCC OS prediction, as the area under curve (AUC) of the ROC curve was 0.600, 0.606, and 0.581 for 1-, 2-, and 5-year survival (Figure 5C), respectively. As shown in Figure 6, the clinicopathological factors (AFP > 300 ng/ml vs. ≤ 300 ng/ml, P = 0.047; BCLC: B/C vs. 0/A, P < 0.001; cirrhosis: yes vs. no, P = 0.025) with a P-value < 0.1 in the univariate analysis were entered into the multivariate analysis. Furthermore, as shown in Table 5 and Figure 7, in the Cox regression model, adjusted by AFP, BCLC, and cirrhosis, patients with high mRNA expression of POR in HCC also had better OS than the patients with low expression (high vs. low: HR = 0.61, 95% CI = [0.38,0.97], P = 0.036).
The result of univariate and multivariate analysis of POR variants with OS.
| Variable | N | Dead | MST (months) | Univariate analysis | Multivariate analysis* | |||
|---|---|---|---|---|---|---|---|---|
| HR(95%CI) | P-value | HR(95%CI) | P-value | |||||
| rs1057868 | ||||||||
| CC | 191 | 120 | 57 | 1.00 | 1.00 | |||
| CT | 228 | 121 | 84 | 0.73(0.56, 0.94) | 0.014 | 0.70(0.54, 0.90) | 0.006 | |
| TT | 57 | 28 | 87 | 0.65(0.43, 0.99) | 0.041 | 0.67(0.45, 1.02) | 0.062 | |
| CT/TT | 285 | 149 | 87 | 0.71(0.56, 0.91) | 0.006 | 0.69(0.54, 0.88) | 0.003 | |
| rs1135612 | ||||||||
| AA | 153 | 81 | 81 | 1.00 | 1.00 | |||
| AG | 229 | 126 | 82 | 1.06(0.80, 1.41) | 0.669 | 1.08(0.81, 1.43) | 0.607 | |
| GG | 94 | 62 | 52 | 1.35(0.97, 1.88) | 0.077 | 1.31(0.94, 1.83) | 0.106 | |
| AG/GG | 323 | 188 | 71 | 1.14(0.88, 1.49) | 0.312 | 1.15(0.88, 1.49) | 0.302 | |
Abbreviations: MST, median survival time; HR, hazard ratio; CI, confidence interval. Note: *Multivariate analysis was adjusted by drinking, Child-Pugh, AFP, PVTT, BCLC and antiviral therapy.
The result of univariate and multivariate analysis of POR variants with RFS.
| Variable | N£ | Recurrent | MRT (months) | Univariate analysis | Multivariate analysis* | |||
|---|---|---|---|---|---|---|---|---|
| HR(95%CI) | P-value | HR(95%CI) | P-value | |||||
| rs1057868 | ||||||||
| CC | 134 | 100 | 13 | 1.00 | 1.00 | |||
| CT | 169 | 125 | 20 | 0.91(0.70, 1.19) | 0.503 | 0.95(0.73, 1.24) | 0.708 | |
| TT | 38 | 23 | 46 | 0.59(0.38, 0.94) | 0.024 | 0.66(0.41, 1.05) | 0.076 | |
| CT/TT | 207 | 148 | 23 | 0.84(0.65, 1.08) | 0.178 | 0.89(0.69, 1.15) | 0.378 | |
| rs1135612 | ||||||||
| AA | 107 | 78 | 27 | 1.00 | 1.00 | |||
| AG | 165 | 123 | 14 | 1.20(0.90, 1.60) | 0.212 | 1.11(0.83, 1.48) | 0.476 | |
| GG | 69 | 47 | 16 | 0.95(0.66, 1.38) | 0.786 | 0.91(0.63, 1.31) | 0.597 | |
| AG/GG | 234 | 170 | 15 | 1.12(0.85, 1.46) | 0.428 | 1.05(0.80, 1.37) | 0.751 | |
Abbreviations: MRT, median recurrent time; HR, hazard ratio; CI, confidence interval. Notes: *Multivariate analysis was adjusted by smoking, Child-Pugh, PVTT, and BCLC; £ The information of RFS was unavailable in 135 patients.
The results of univariate analysis of POR mRNA expression with OS.
| Variable | N | Dead | MST (months) | Univariate analysis | Multivariate analysis* | |||
|---|---|---|---|---|---|---|---|---|
| HR(95%CI) | P-value | HR(95%CI) | P-value | |||||
| POR expression | ||||||||
| Low | 106 | 49 | NA | 1.00 | 1.00 | |||
| High | 106 | 33 | NA | 0.54(0.35, 0.84) | 0.006 | 0.61(0.38, 0.97) | 0.036 | |
Abbreviations: MST, median survival time; HR, hazard ratio; CI, confidence interval. Note: *Multivariate analysis was adjusted by AFP, BCLC and cirrhosis.
The result of univariate analysis of clinicopathological factors with OS and RFS in the cohort 1.
The result of multivariate analysis of POR-rs1057868 combined with clinicopathological factors with OS.
Association of POR mRNA expression with RFS
The results of the univariate analysis of POR mRNA expression with RFS are presented in Table 6 and Figure 5D. The median RFS of the POR low- and high-expression group was 28 and 51 months, respectively. In the univariate analysis, no significant difference in RFS was found between POR low- and high-expression groups (high vs. low: HR = 0.74, 95% CI = [0.52, 1.07], P = 0.112). As shown in Figure 6, the clinicopathological factors (gender: female vs. male, P = 0.018; BCLC: B/C vs. 0/A, P < 0.001; cirrhosis: yes vs. no, P = 0.029) with a P-value < 0.1 in the univariate analysis were entered into the multivariate analysis. Furthermore, as shown in Table 6, in the Cox regression model, adjusted by gender, BCLC, and cirrhosis, there was also no significant difference in RFS between POR low- and high-expression groups (high vs. low: HR = 0.78, 95% CI = [0.54, 1.14], P = 0.201).
Nomograms and stratified analysis
Multivariate Cox analysis suggested that POR-rs1057868 and mRNA expression were significantly associated with HBV-related HCC OS. To further investigate the prognostic values of POR-rs1057868 and mRNA expression in HBV-related HCC prognosis, we performed a stratified analysis to assess prognosis values in different strata of HCC and developed two nomograms for individual prognostic evaluation based on POR-rs1057868 or mRNA expression combined with other clinical features.
Kaplan-Meier curves for RFS among different POR variants groups. (A) RFS stratified by POR-rs1057868 CC, CT and TT genotypes; (B) RFS stratified by POR-rs1057868 CC and CT/TT genotypes; (C) RFS stratified by POR-rs1135612 AA, AG and GG genotypes; (D) RFS stratified by POR- rs1135612 AA and AG/GG genotypes.
Prognosis values of POR mRNA expression in HBV-related HCC prognosis prediction. (A) OS stratified by POR mRNA expression levels; (B) From top to bottom are the POR mRNA expression, patients' survival status distribution, and POR mRNA expression heat map for low- and high-expression groups. (C) ROC curve for predicting OS in HBV-related HCC patients by POR mRNA expression; (D) RFS stratified by POR mRNA expression.
The result of univariate analysis of clinicopathological factors with OS and RFS in the cohort 2.
The result of multivariate analysis of POR mRNA expression combined with clinicopathological factors with OS.
The results of univariate analysis of POR mRNA expression with RFS.
| Variable | N | Recurrent | MRT (months) | Univariate analysis | Multivariate analysis* | |||
|---|---|---|---|---|---|---|---|---|
| HR(95%CI) | P-value | HR(95%CI) | P-value | |||||
| POR expression | ||||||||
| Low | 106 | 61 | 28 | 1.00 | 1.00 | |||
| High | 106 | 55 | 51 | 0.74(0.52, 1.07) | 0.112 | 0.78(0.54, 1.14) | 0.201 | |
Abbreviation: MRT, median recurrent time; HR, hazard ratio; CI, confidence interval. Note: *Multivariate analysis was adjusted by gender, BCLC, and cirrhosis.
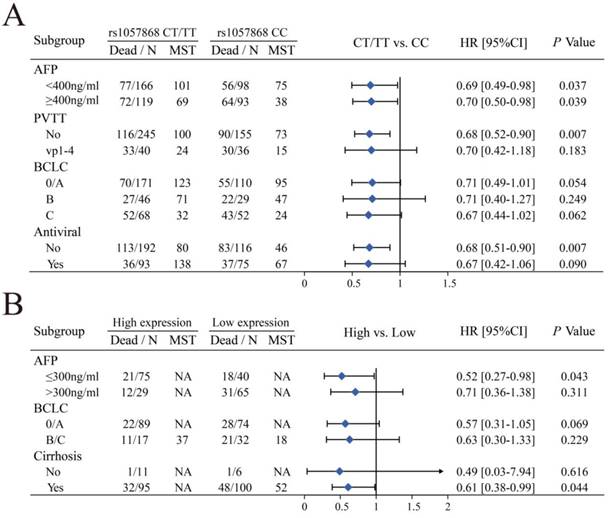
The stratified analysis suggested that patients in both AFP values, without PVTT and antiviral therapy, with a T allele of POR-rs1057868 had a significantly reduced risk of death in HBV-related HCC compared with the CC genotypes (Figure 8A). We also performed a stratified analysis for POR mRNA expression in the GSE14520 cohort and suggested that high expression of POR had a significantly reduced risk of death of HBV-related HCC in patients with AFP ≤ 300 ng/ml and cirrhosis, compared with low expression (Figure 8B).
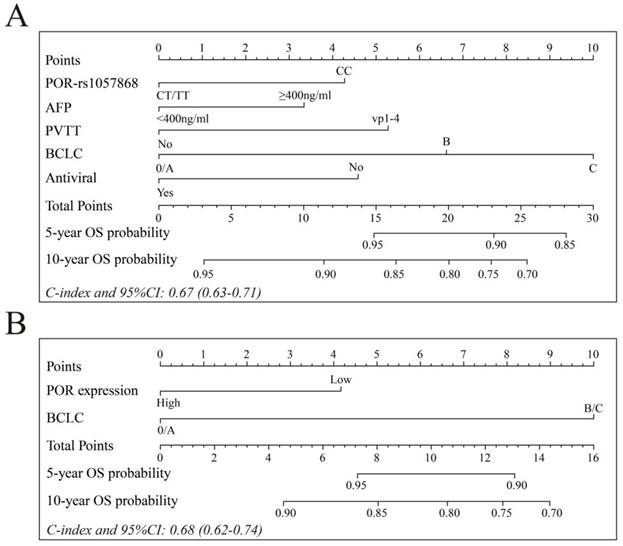
Nomograms base on POR-rs1057868 or mRNA expression combine with other clinical features are shown in Figure 9A and B. The c-index (95% CI) of the nomogram of POR-rs1057868 combined with AFP, PVTT, BCLC, and antiviral therapy was 0.67 (0.63 to 0.71). The c-index (95% CI) of the nomogram of POR mRNA expression combined with BCLC was 0.68 (0.62 to 0.74).
GSEA
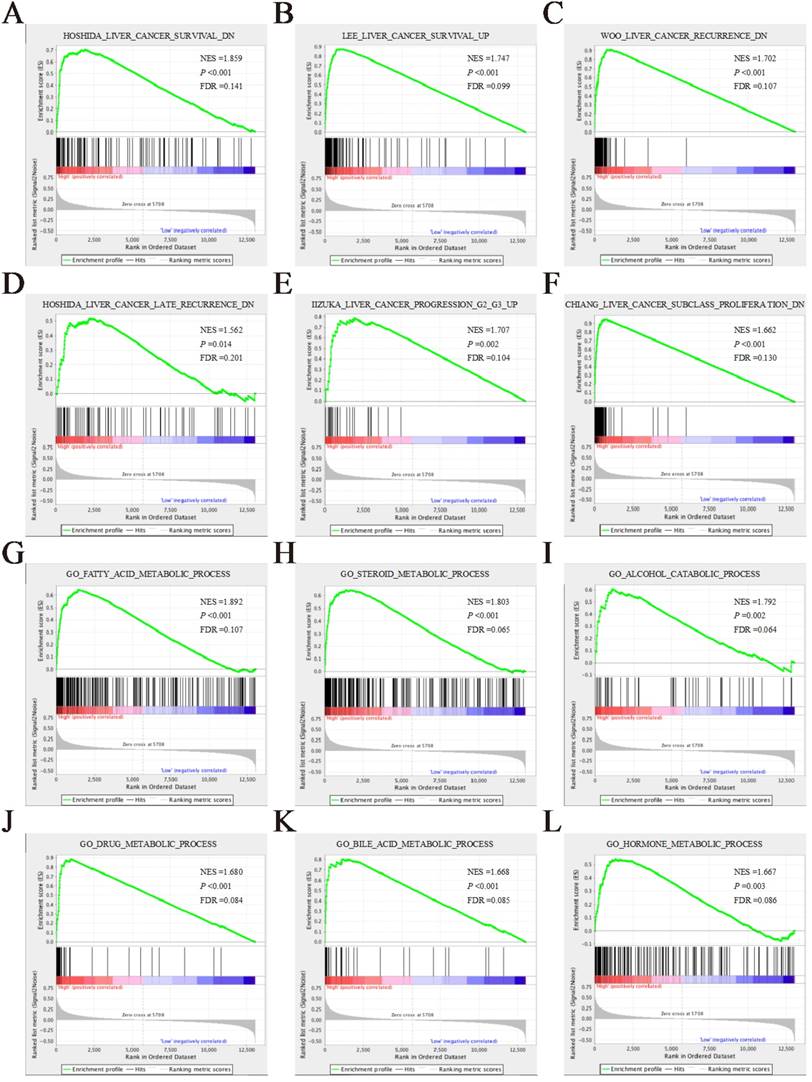
In addition, we conducted a GSEA to identify gene sets that contributed to the HCC prognosis between POR high- and low-expression groups. The enrichment results of the high POR expression group are summarized in supplementary Tables S3 and S4. The enrichment results of the high POR expression group in the c2 gene set suggested that there were multiple gene sets were found to significantly correlate with liver cancer survival, recurrence, and development[27-31] (Figure 10 A-F). For the c5 gene set, we observed that high POR expression was significantly enriched in the metabolism of multiple substances in biological processes (Figure 10 G-L).
(A) Stratified analysis of POR-rs1057868 in different clinical features; (B) Stratified analysis of POR mRNA expression in different clinical features.
(A) Nomograms for predicting HBV-related HCC OS with POR-rs1057868 genotypes and clinical information; (B) Nomograms for predicting HBV-related HCC OS with POR mRNA expression and clinical information.
GSEA results of POR in GSE14520 HBV-related HCC patients. (A-F) GSEA results of c2 reference gene sets for high POR expression groups; (G-L) GSEA results of c5 reference gene sets for high POR expression groups.
Discussion
The human POR gene, located on chromosome 7q11.23, is highly polymorphic[13, 32]. To date, more than 50 POR SNPs have been reported[33]. In a study population of 476 HBV-related HCC patients from south China, we observed two SNPs, rs1057868 and rs1135612, in POR exons. The MAF of rs1057868 and rs1135612 was 35.9% and 43.8%, respectively, which was close to the MAF in the HapMap database (39.6% and 45.8%, respectively)[16]. POR-rs1057868 is a missense mutation (A503V), which has been identified as the most common POR variant in all racial/ethnic populations (about 28% of human alleles)[33]. It has been shown that the variant A503V is a conservative change in an unstructured loop of the FAD-binding domain, which may affect the activity of POR[34]. Hart et al. demonstrated that variant A503V was significantly related to POR activity (P = 0.003)[35]. As an obligate electron donor for CYP enzymes, POR plays a critical role in the metabolic process of CYPs. Many previous studies have reported that POR-rs1057868 was associated with the activities of CYPs. Agrawal et al. reported that POR A503V caused a 15% decrease in CYP1A2 activity, 13% increase in CYP2C19, and a 33%-49% decrease in CYP3A4 in vitro[36, 37]. Hylke et al. reported that POR A503V was associated with increased CYP3A5 activity in 298 patients[38]. Additionally, multiple studies observed that the A503V variant decreased cytochrome c reduction activity[33, 39]. Most of the previous studies suggested that POR variants were associated with certain genetic disorders, such as ambiguous genitalia, Antley-Bixler skeletal malformation syndrome, and congenital adrenal hyperplasia[40, 41]. Recently, a few studies have focused on the association between the polymorphisms in A503V and cancer risk. Xiao et al. reported that the POR-rs1057868 was correlated with the risk of bladder cancer in a Chinese population[16]. However, no study has investigated the association between POR-rs1057868 and the risk and prognosis of HCC. In the present study, we are first to report that POR-rs1057868 was significantly associated with OS in HBV-related HCC. The patients with rs1057868 CT or TT genotype had a better OS after hepatectomy compared to those with the CC genotype. However, no significant association was found between POR-rs1057868 and RFS. The other POR variant rs1135612 is a synonymous mutation (P130P). To date, no study has reported that POR-rs1135612 was associated with the activities of POR and CYPs or the risk and prognosis of cancer. In the present study, no significant association was found between POR-rs1135612 and OS and RFS in HBV-related HCC.
With the hypothesis that the mRNA expression level of POR might affect the activity of POR and in turn, the prognosis of HBV-related HCC, we also investigated the association between POR mRNA expression and the prognosis of HBV-related HCC using the GSE14520 dataset. We found that HBV-related HCC patients with a high-expression of POR have a better OS after hepatectomy than those with low-expression. GSEA is a powerful analytical method for interpreting gene expression data and identifying biological pathways in cancer[24]. In the present study, we also performed a GSEA to identify biological pathways that correlated with HBV-related HCC prognosis. We found that multiple gene sets were significantly correlated with liver cancer survival, recurrence, and HCC development and with the metabolism of multiple substances in biological processes. Therefore, the results of the GSEA suggested that the mRNA expression of POR might play an important role in the prognosis of HBV-related HCC.
In the present study, we found that variants POR-rs1057868 and mRNA expression were associated with the OS in HBV-related HCC. Nomograms are widely used as prognostic devices in oncology with the ability to generate an individual probability of a clinical event[26]. Considering OS might be affected by genetic and clinicopathological factors, we also developed two novel nomograms to predict OS by enrolling variants POR-rs1057868 and mRNA expression combined with clinicopathological factors. It might be useful in the clinic to identify high-risk patients.
This study has some limitations that should be pointed out. First, since the HCC specimens from our cohort were all collected before 2013, the mRNA of these HCC had been degraded. Therefore, we could not evaluate the association of POR-rs1057868 and mRNA expression with clinical outcome of HBV-related HCC in the same cohort population. Secondly, the patient's clinical information obtained from the GSE14520 dataset was limited, therefore some factors were missing and some cut-off factors were inconsistent with data from cohort 1. Thirdly, since this is a retrospective study, some postoperative reexaminations and treatment are not controllable. Some patients had not been examined for imaging up until death, and consequently, it was uncertain whether HCC had recurred. Therefore, the recurrence rate of this study was low. Fourth, due to the difficult in HCC tumor tissues, both the results of POR variants and mRNA expression level in HBV-related HCC prognosis were generated from single cohort, and lack of verification cohort. As well as, the present study is a clinical prognosis biomarker investigation study and bioinformatics analysis study. Therefore, the present study was lack of in vivo and in vitro functional experiment verification.
Despite these limitations, we are first to report the prognostic application of POR-rs1057868 and mRNA expression in patients with HBV-related HCC. In addition, we investigated the potential mechanism of different POR mRNA expression levels in HCC prognosis through a GSEA approach. Once these results are confirmed, POR may have potential in the clinical application of prognostic monitoring, cancer management, and targeted therapy of HCC.
Conclusions
In summary, our study suggests that POR-rs1057868 (A503V) and mRNA expression were significantly associated with OS in HBV-related HCC patients and may serve as potential biomarkers for predicting OS in HBV-related HCC after hepatectomy. We also developed two novel nomograms consisting of traditional clinical parameters and POR-rs1057868 or mRNA expression to predict OS in HBV-related HCC. However, additional HBV-related HCC cohorts are needed to verify our results.
Abbreviations
POR: cytochrome p450 oxidoreductase; HCC: hepatocellular carcinoma; HBV: hepatitis B virus; HBsAg: hepatitis B virus surface antigen; PVTT: portal vein tumor thrombus; AFP: alpha-fetoprotein; ALT: alanine aminotransferase; BCLC: Barcelona Clinic Liver Cancer; SNP: single nucleotide polymorphisms; OS: overall survival; RFS: recurrence-free survival; MST: median survival time; MRT: median recurrent time; HR: hazard ratio; CI: confidence interval; MAF: minor allele frequency; HWE: Hardy-Weinberg equilibrium; FDR: false discovery rate; c-index: concordance index; ROC: receiver operating characteristic; AUC: area under curve; GSEA: gene set enrichment analysis.
Supplementary Material
Supplementary tables.
Acknowledgements
This work was supported in part by the National Nature Science Foundation of China (Grant No. 81560535, 81072321, 30760243, 30460143 and 30560133), 2009 Program for New Century Excellent Talents in University (NCET), Guangxi Nature Sciences Foundation (GuiKeGong 1104003A-7), and Guangxi Health Ministry Medicine Grant (Key- Scientific Research-Grant Z201018). The present study is also partly supported by Self-raised Scientific Research Fund of the Health and Family Planning Commission of Guangxi Zhuang Autonomous Region (Z2016318), The Basic Ability Improvement Project for Middle-aged and Young Teachers in Colleges and Universities in Guangxi (2018KY0110), and Innovation Project of Guangxi Graduate Education (JGY2018037). As well as, the present study is also partly supported by Research Institute of Innovative Think-tank in Guangxi Medical University (The gene-environment interaction in hepatocarcinogenesis in Guangxi HCCs and its translational applications in the HCC prevention). We would also acknowledge the supported by the National Key Clinical Specialty Programs (General Surgery & Oncology) and the Key Laboratory of Early Prevention & Treatment for Regional High-Incidence-Tumor (Guangxi Medical University), Ministry of Education, China. The authors thank the contributors of GSE14520 (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE14520) for sharing the HBV-related HCC data on open access.
Authors' Contributions
Ketuan Huang, Xiwen Liao and Tao Peng designed this manuscript; Ketuan Huang, Xiwen Liao, Chuangye Han, Xiangkun Wang, Tingdong Yu, Chengkun Yang, Xiaoguang Liu, Long Yu, Zhiwei Chen, Wei Qin, Guangzhi Zhu, Hao Su, Zhengqian Liu, Xianmin Zeng, Xin Zhou, Sicong Lu, Jianlv Huang, Yu Liang, Zhengtao Liu, Jianlong Deng, Xinping Ye, and Tao Peng conducted the study, collected and analyzed the data. Ketuan Huang wrote and revised the manuscript, Xiwen Liao contributed to the data interpretation, and Tao Peng guided the writing. All authors read and approved the final manuscript.
Ethical approval
The samples from human in this study were obtained in accordance with the ethical standards of the ethics committee of The First Affiliated Hospital of Guangxi Medical University.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Yang JD, Roberts LR. Hepatocellular carcinoma: A global view. Nature reviews Gastroenterology & hepatology. 2010;7:448-58
2. Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A. Global cancer statistics, 2012. CA: a cancer journal for clinicians. 2015;65:87-108
3. El-Serag HB. Epidemiology of viral hepatitis and hepatocellular carcinoma. Gastroenterology. 2012;142:1264-73 e1
4. Gluer AM, Cocco N, Laurence JM, Johnston ES, Hollands MJ, Pleass HC. et al. Systematic review of actual 10-year survival following resection for hepatocellular carcinoma. HPB: the official journal of the International Hepato Pancreato Biliary Association. 2012;14:285-90
5. Zhong JH, Ke Y, Gong WF, Xiang BD, Ma L, Ye XP. et al. Hepatic resection associated with good survival for selected patients with intermediate and advanced-stage hepatocellular carcinoma. Annals of surgery. 2014;260:329-40
6. Ma WJ, Wang HY, Teng LS. Correlation analysis of preoperative serum alpha-fetoprotein (AFP) level and prognosis of hepatocellular carcinoma (HCC) after hepatectomy. World journal of surgical oncology. 2013;11:212
7. Nathan H, Schulick RD, Choti MA, Pawlik TM. Predictors of survival after resection of early hepatocellular carcinoma. Annals of surgery. 2009;249:799-805
8. Sakamoto K, Nagano H. Surgical treatment for advanced hepatocellular carcinoma with portal vein tumor thrombus. Hepatology research: the official journal of the Japan Society of Hepatology. 2017;47:957-62
9. Huang G, Lau WY, Wang ZG, Pan ZY, Yuan SX, Shen F. et al. Antiviral therapy improves postoperative survival in patients with hepatocellular carcinoma: a randomized controlled trial. Annals of surgery. 2015;261:56-66
10. Vitale A, Burra P, Frigo AC, Trevisani F, Farinati F, Spolverato G. et al. Survival benefit of liver resection for patients with hepatocellular carcinoma across different Barcelona Clinic Liver Cancer stages: a multicentre study. Journal of hepatology. 2015;62:617-24
11. Yuan JM, Chan KK, Coetzee GA, Castelao JE, Watson MA, Bell DA. et al. Genetic determinants in the metabolism of bladder carcinogens in relation to risk of bladder cancer. Carcinogenesis. 2008;29:1386-93
12. Wu K, Wang X, Xie Z, Liu Z, Lu Y. Glutathione S-transferase P1 gene polymorphism and bladder cancer susceptibility: an updated analysis. Molecular biology reports. 2013;40:687-95
13. Pandey AV, Fluck CE. NADPH P450 oxidoreductase: structure, function, and pathology of diseases. Pharmacology & therapeutics. 2013;138:229-54
14. Hamdane D, Xia C, Im SC, Zhang H, Kim JJ, Waskell L. Structure and function of an NADPH-cytochrome P450 oxidoreductase in an open conformation capable of reducing cytochrome P450. The Journal of biological chemistry. 2009;284:11374-84
15. Haiman CA, Setiawan VW, Xia LY, Le Marchand L, Ingles SA, Ursin G. et al. A variant in the cytochrome p450 oxidoreductase gene is associated with breast cancer risk in African Americans. Cancer research. 2007;67:3565-8
16. Xiao X, Ma G, Li S, Wang M, Liu N, Ma L. et al. Functional POR A503V is associated with the risk of bladder cancer in a Chinese population. Scientific reports. 2015;5:11751
17. Roessler S, Jia HL, Budhu A, Forgues M, Ye QH, Lee JS. et al. A unique metastasis gene signature enables prediction of tumor relapse in early-stage hepatocellular carcinoma patients. Cancer research. 2010;70:10202-12
18. Roessler S, Long EL, Budhu A, Chen Y, Zhao X, Ji J. et al. Integrative genomic identification of genes on 8p associated with hepatocellular carcinoma progression and patient survival. Gastroenterology. 2012;142:957-66 e12
19. Ayuso C, Rimola J, Vilana R, Burrel M, Darnell A, Garcia-Criado A. et al. Diagnosis and staging of hepatocellular carcinoma (HCC): current guidelines. European journal of radiology. 2018;101:72-81
20. Kudo M, Kitano M, Sakurai T, Nishida N. General Rules for the Clinical and Pathological Study of Primary Liver Cancer, Nationwide Follow-Up Survey and Clinical Practice Guidelines: The Outstanding Achievements of the Liver Cancer Study Group of Japan. Digestive diseases. 2015;33:765-70
21. Pugh RN, Murray-Lyon IM, Dawson JL, Pietroni MC, Williams R. Transection of the oesophagus for bleeding oesophageal varices. The British journal of surgery. 1973;60:646-9
22. Zheng SS, Xu X, Wu J, Chen J, Wang WL, Zhang M. et al. Liver transplantation for hepatocellular carcinoma: Hangzhou experiences. Transplantation. 2008;85:1726-32
23. Sanger F, Coulson AR. A rapid method for determining sequences in DNA by primed synthesis with DNA polymerase. Journal of molecular biology. 1975;94:441-8
24. Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proceedings of the National Academy of Sciences of the United States of America. 2005;102:15545-50
25. Liberzon A, Birger C, Thorvaldsdottir H, Ghandi M, Mesirov JP, Tamayo P. The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell systems. 2015;1:417-25
26. Balachandran VP, Gonen M, Smith JJ, DeMatteo RP. Nomograms in oncology: more than meets the eye. The Lancet Oncology. 2015;16:e173-80
27. Hoshida Y, Villanueva A, Kobayashi M, Peix J, Chiang DY, Camargo A. et al. Gene expression in fixed tissues and outcome in hepatocellular carcinoma. The New England journal of medicine. 2008;359:1995-2004
28. Lee JS, Chu IS, Heo J, Calvisi DF, Sun Z, Roskams T. et al. Classification and prediction of survival in hepatocellular carcinoma by gene expression profiling. Hepatology. 2004;40:667-76
29. Woo HG, Park ES, Cheon JH, Kim JH, Lee JS, Park BJ. et al. Gene expression-based recurrence prediction of hepatitis B virus-related human hepatocellular carcinoma. Clinical cancer research: an official journal of the American Association for Cancer Research. 2008;14:2056-64
30. Iizuka N, Oka M, Yamada-Okabe H, Mori N, Tamesa T, Okada T. et al. Self-organizing-map-based molecular signature representing the development of hepatocellular carcinoma. FEBS letters. 2005;579:1089-100
31. Chiang DY, Villanueva A, Hoshida Y, Peix J, Newell P, Minguez B. et al. Focal gains of VEGFA and molecular classification of hepatocellular carcinoma. Cancer research. 2008;68:6779-88
32. Miller WL, Agrawal V, Sandee D, Tee MK, Huang N, Choi JH. et al. Consequences of POR mutations and polymorphisms. Molecular and cellular endocrinology. 2011;336:174-9
33. Huang N, Agrawal V, Giacomini KM, Miller WL. Genetics of P450 oxidoreductase: sequence variation in 842 individuals of four ethnicities and activities of 15 missense mutations. Proceedings of the National Academy of Sciences of the United States of America. 2008;105:1733-8
34. Scott RR, Gomes LG, Huang N, Van Vliet G, Miller WL. Apparent manifesting heterozygosity in P450 oxidoreductase deficiency and its effect on coexisting 21-hydroxylase deficiency. The Journal of clinical endocrinology and metabolism. 2007;92:2318-22
35. Hart SN, Wang S, Nakamoto K, Wesselman C, Li Y, Zhong XB. Genetic polymorphisms in cytochrome P450 oxidoreductase influence microsomal P450-catalyzed drug metabolism. Pharmacogenetics and genomics. 2008;18:11-24
36. Agrawal V, Huang N, Miller WL. Pharmacogenetics of P450 oxidoreductase: effect of sequence variants on activities of CYP1A2 and CYP2C19. Pharmacogenetics and genomics. 2008;18:569-76
37. Agrawal V, Choi JH, Giacomini KM, Miller WL. Substrate-specific modulation of CYP3A4 activity by genetic variants of cytochrome P450 oxidoreductase. Pharmacogenetics and genomics. 2010;20:611-8
38. de Jonge H, Metalidis C, Naesens M, Lambrechts D, Kuypers DR. The P450 oxidoreductase *28 SNP is associated with low initial tacrolimus exposure and increased dose requirements in CYP3A5-expressing renal recipients. Pharmacogenomics. 2011;12:1281-91
39. Huang N, Pandey AV, Agrawal V, Reardon W, Lapunzina PD, Mowat D. et al. Diversity and function of mutations in p450 oxidoreductase in patients with Antley-Bixler syndrome and disordered steroidogenesis. American journal of human genetics. 2005;76:729-49
40. Adachi M, Tachibana K, Asakura Y, Yamamoto T, Hanaki K, Oka A. Compound heterozygous mutations of cytochrome P450 oxidoreductase gene (POR) in two patients with Antley-Bixler syndrome. American journal of medical genetics Part A. 2004;128A:333-9
41. Ko JM, Cheon CK, Kim GH, Yoo HW. A case of Antley-Bixler syndrome caused by compound heterozygous mutations of the cytochrome P450 oxidoreductase gene. European journal of pediatrics. 2009;168:877-80
Author contact
![]() Corresponding author: Prof. Tao Peng, Department of Hepatobiliary Surgery, The First Affiliated Hospital of Guangxi Medical University, Shuang_Yong Rd. 6#, Nanning, 530021, Guangxi Zhuang Autonomous Region, People's Republic of China. Tel: (+86)-771-5356528. Fax: (+86)-771-5350031. E-mail: pengtaogmucom; pengtaoedu.cn. ORCID ID: http://orcid.org/0000-0001-6133-7078.
Corresponding author: Prof. Tao Peng, Department of Hepatobiliary Surgery, The First Affiliated Hospital of Guangxi Medical University, Shuang_Yong Rd. 6#, Nanning, 530021, Guangxi Zhuang Autonomous Region, People's Republic of China. Tel: (+86)-771-5356528. Fax: (+86)-771-5350031. E-mail: pengtaogmucom; pengtaoedu.cn. ORCID ID: http://orcid.org/0000-0001-6133-7078.

 Global reach, higher impact
Global reach, higher impact