Impact Factor ISSN: 1837-9664
J Cancer 2026; 17(3):614-625. doi:10.7150/jca.127511 This issue Cite
Review
Fluid-Derived Organoids from Pleural Effusion and Ascites: Emerging Models for Drug Resistance and Personalized Oncology
1. Department of Pulmonary Oncology, Affiliated Hospital of Guangdong Medical University, Zhanjiang, 524001, China.
2. Guangdong Medical University, Dongguan, 523000, China.
3. Dalian Medical University, Dalian, 116044, China.
4. Guangdong Provincial Key Laboratory of Autophagy and Major Chronic Non-Communicable Diseases, Affiliated Hospital of Guangdong Medical University, Zhanjiang, 524001, China.
5. Zhanjiang Key Laboratory of Tumor Microenvironment and Organoid Research, Zhanjiang, 524001, China.
6. Guangdong Provincial Key Laboratory of Medical Immunology and Molecular Diagnostics, Guangdong Medical University, Dongguan, 523000, China.
7. Guangdong Medical University, Zhanjiang, 524001, China.
†These authors contributed equally to this work.
Received 2025-10-29; Accepted 2026-2-10; Published 2026-3-4
Abstract
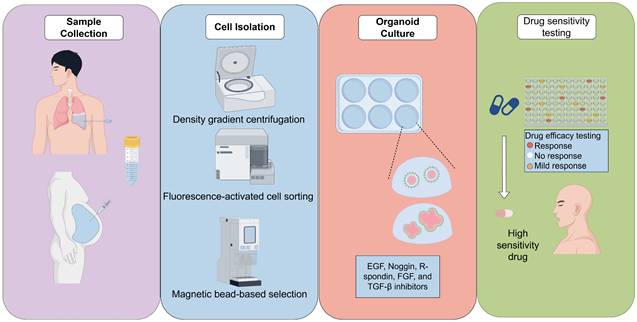
Malignant pleural effusion (MPE) and malignant ascites (MA) are common complications in advanced-stage cancers, often signifying disease progression and resistance to treatment. Compared to tissue biopsies or surgical specimens, materials derived from effusions offer advantages such as minimal invasiveness, ease of accessibility, and the feasibility of repeated collection during therapeutic interventions. Organoids generated from tumor cells in effusions, termed fluid-derived organoids (FDOs), have demonstrated the ability to maintain genetic heterogeneity and accurately replicate patient-specific tumor phenotypes. These characteristics position FDOs as promising models for investigating drug resistance mechanisms and informing personalized oncology strategies. In the context of lung cancer, organoids derived from pleural effusions have been employed to study acquired resistance to epidermal growth factor receptor (EGFR) tyrosine kinase inhibitors and immunotherapy. Similarly, in ovarian and gastrointestinal cancers, organoids derived from ascites have proven to be valuable platforms for examining chemotherapy resistance and conducting drug sensitivity testing. FDOs have shown significant potential for translational applications by effectively correlating ex vivo drug responses with clinical outcomes, thus facilitating real-time monitoring of resistance evolution. However, several challenges remain, such as achieving culture standardization, maintaining the integrity of tumor microenvironment components, and integrating with multi-omics approaches. This review provides a comprehensive overview of recent advancements in the use of pleural effusion- and ascites-derived organoids for drug resistance research, underscores their applications in personalized oncology, and explores future research directions.
Keywords: fluid-derived organoids, pleural effusion, ascites, drug resistance, personalized oncology
Introduction
Lung, ovarian, gastrointestinal, and pancreatic cancers continue to rank among the leading causes of cancer-related mortality globally [1]. A significant proportion of patients with advanced-stage disease present with malignant effusions, most frequently malignant pleural effusion (MPE) and malignant ascites (MA) [2, 3]. These effusions not only indicate disease progression but also reflect the onset of therapeutic resistance, presenting a substantial challenge in the field of oncology [2]. Clinically, the presence of malignant effusions is often correlated with a poor prognosis, limited therapeutic options, and diminished quality of life [4]. Therefore, the development of innovative strategies to investigate drug resistance and inform personalized therapy in this patient population is of critical importance.
Conventional preclinical models, such as two-dimensional (2D) cell lines, patient-derived xenografts (PDXs), and organoids derived from surgical or biopsy tissues, have significantly contributed to our understanding of cancer biology. Nonetheless, these models are insufficient in capturing the complexity of resistance mechanisms in advanced stages of the disease (Table 1) [5]. It has been shown that 2D cultures often fail to maintain intratumoral heterogeneity and lack the structural architecture of the native tumor microenvironment [6]. PDX models provide a more precise representation of tumor evolution; however, they are resource-intensive, time-consuming, and limited by the replacement of human stroma with murine stroma [7, 8]. Organoids derived from surgical tissues, despite their enhanced physiological relevance, depend on specimens obtained at the time of diagnosis or surgical intervention. As a result, these organoids may not accurately represent the development of resistance mechanisms that occur during treatment [9].
Comparison of FDOs with other preclinical models
| Advantage | Disadvantage | References | |
|---|---|---|---|
| 2D cell lines | Low cost, high throughput | Lack of heterogeneity and three-dimensional structure | [6] |
| PDXs | Preserve tumor complexity | Mouse-derived matrix interference, time-consuming | [7,8] |
| Organoids from surgical tissues | High physiological relevance | Relying on surgical samples, unable to monitor dynamically | [9] |
| FDOs | Minimally invasive, repeatable sampling, representative of advanced disease | The success rate varies greatly depending on the type of tumor | [12-14] |
| Co-culture system | Simulated immune microenvironment | Technically complex and low in standardization | [120,127] |
In contrast, malignant effusions provide an abundant and minimally invasive source of tumor cells [10]. Repeated thoracentesis and paracentesis procedures can be performed throughout the progression of the disease, allowing for real-time access to tumor material as resistance develops [11]. Organoids derived from effusion-derived tumor cells, collectively known as fluid-derived organoids (FDOs), constitute a promising model system [12]. The tumor cells present in malignant effusions are clones that have adapted to survive in the metastatic fluid environment. They often possess enhanced metastatic capacity, stem-like properties, and resistance to anoikis—a critical mechanism of cell death evasion linked to therapy resistance [13, 14]. Consequently, FDOs may be inherently predisposed to model drug resistance at a more advanced, metastatic disease stage. Furthermore, a malignant effusion is not merely a tumor cell suspension but a complex microenvironment. It is rich in immune cells, mesothelial cells, and fibroblasts [13]. Effusions frequently contain dysfunctional or suppressive immune populations, such as exhausted T cells and M2 macrophages [15]. If retained or reconstituted in co-culture, these cells could make FDOs excellent models for evaluating response or resistance to immune checkpoint inhibitors. Conversely, their loss during culture might lead to an overestimation of immunotherapy efficacy. Mesothelial cells and other cavity-resident cells interact with metastatic tumor cells, providing vital survival signals and forming a key component of the metastatic niche [14]. By secreting various protective factors, these cells can directly influence tumor cell sensitivity to chemotherapy or target agents. Thus, the distinct advantage of FDOs lies in their inherent complexity and context. They directly capture the cellular ecosystems of advanced metastatic sites, making them potentially more relevant than PDOs from primary tumors for modeling stage-specific drug resistance mechanisms, including metastatic adaptation, immune evasion, and microenvironment-mediated protection. In the realm of non-small cell lung cancer (NSCLC), the development of pleural effusion-derived organoids (PE-organoids) have become a focal point of research. These organoids have been instrumental in studies aimed at understanding the mechanisms underlying acquired resistance to epidermal growth factor receptor tyrosine kinase inhibitors (EGFR-TKIs). Investigations include the analysis of secondary mutations, such as T790M and C797S, as well as the exploration of bypass signaling pathways involving MET and HER2 amplification [16, 17]. Similarly, in the study of ovarian cancer, ascites-derived organoids (AS-organoids) have been extensively employed to elucidate the mechanisms of resistance to platinum and taxane therapies. Additionally, these organoids have been used to predict responsiveness to targeted therapies, such as PARP inhibitors [18]. In gastrointestinal and pancreatic cancers, AS-organoids have been utilized to evaluate chemotherapy efficacy and to explore personalized drug screening strategies [19].
Beyond the realm of mechanistic studies, the translational applications of FDOs are gaining increasing recognition [20]. Numerous studies have demonstrated a significant concordance between the ex vivo drug responses of FDOs and clinical outcomes in corresponding patients, indicating that organoid-based assays may serve as predictive biomarkers for treatment efficacy [21, 22]. Their ability to model clonal evolution longitudinally is particularly valuable for understanding resistance dynamics [23]. However, the field is still in its early stages of development. Technical challenges associated with this approach include variable success rates across different tumor types, a lack of standardized protocols, and limited preservation of tumor microenvironment components, such as immune and stromal cells [24, 25].
The subsequent discourse will concentrate on the establishment and characteristics of FDOs, followed by an overview of the applications of pleural effusion- and ascites-derived organoids in resistance research across various cancer types. Finally, we will underscore the translational applications, current challenges, and future perspectives associated with integrating FDOs into clinical practice.
Establishment and Features of Fluid-Derived Organoids
Sample Collection and Cell Isolation
Effusion sample collection is generally conducted through thoracentesis or paracentesis under sterile conditions [26]. Compared to surgical resections, these procedures are less invasive, allow for repeated application, and yield adequate material even from patients with poor performance status [27]. The initial processing of these samples involves centrifugal separation to concentrate the cells, followed by red blood cell lysis and filtration to eliminate extraneous debris [28]. Numerous studies recommend the use of density gradient centrifugation (e.g., Ficoll) to enrich tumor cells and reduce contamination by immune or mesothelial cells [29]. In certain cases, fluorescence-activated cell sorting (FACS) or magnetic bead-based selection may be utilized to further enrich epithelial tumor cells. However, it is important to note that these methods may increase costs and complexity [30].
Organoid Culture Systems
Isolated cells are incorporated into extracellular matrix substitutes such as Matrigel, Cultrex, or other basement membrane extracts, which provide the essential three-dimensional scaffold [31]. Matrigel is a popular matrix for organoid culture but is unsuitable for clinical use due to its mouse sarcoma origin. Researchers have developed collagen-based matrices that support organoid growth like Matrigel and are more clinically viable [32]. Additionally, synthetic peptide hydrogels are used for kidney organoids, improving maturation and reducing unwanted cell differentiation [33]. Subsequently, these cells are cultured in optimized media formulations enriched with niche-specific growth factors and inhibitors. For lung cancer, the most prevalent media components include EGF, Noggin, R-spondin, FGF, and TGF-β inhibitors [34, 35]. In the context of organoids derived from ovarian cancer ascites, the addition of Wnt agonists and estrogen-related factors has been shown to enhance growth promotion [36]. The variability in media formulations across different laboratories presents a significant challenge, as it affects organoid morphology, growth efficiency, and drug response profiles (Fig. 1).
Schematic Diagram of Fluid-Derived Organoid Preparation
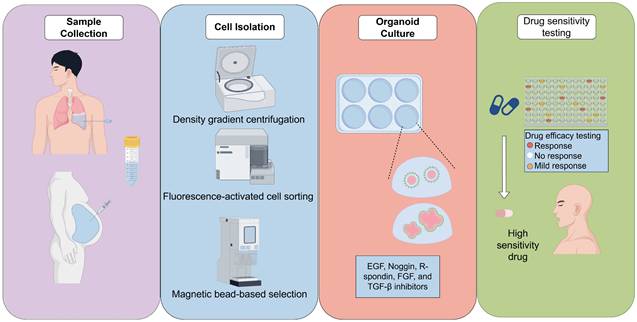
Success Rates Across Tumor Types
As evidenced by existing literature, the efficiency of FDO establishment exhibits significant variability across different tumor types [37]. The reported success rates are notably higher in ovarian cancer, ranging from 60% to 90%, attributed to the high concentration of viable tumor cells present in ascites [38]. Conversely, success rates for FDO establishment from pleural effusions in lung cancer range between 30% and 60%, influenced by factors such as tumor cell fraction and prior treatment exposure [39, 40]. Gastric and pancreatic cancers exhibit intermediate success rates, approximately 40% to 70%, whereas breast cancer and lymphoma demonstrate lower efficiencies [41]. Key factors impacting these success rates include tumor cellularity, the extent of immune cell infiltration, and previous chemotherapy exposure [42]. These findings indicate that while FDO generation is achievable across various cancer types, further optimization is necessary to ensure consistency [43].
Preservation of Tumor Characteristics
Several studies have corroborated that FDOs maintain critical histopathological and molecular characteristics of the original tumors [44]. Histological examination using hematoxylin and eosin staining frequently reveals glandular or papillary structures akin to those of the primary tumor [45]. Genomic analyses further confirm that FDOs retain essential driver mutations, copy number alterations, and mutational signatures present in the original tumors in patient samples [46]. Intra-patient heterogeneity is notably preserved, as organoids can reflect subclonal populations identified in sequencing studies [47]. Further evidence of their fidelity is demonstrated through transcriptomic profiling, which indicates that FDOs cluster closely with their corresponding parental tumors [48].
Characteristics of FDOs
Compared to tissue-derived patient-derived organoids (PDOs), FDOs present several advantages (Table 2). Firstly, they are minimally invasive and can be repeatedly sampled, facilitating longitudinal studies [49]. Secondly, effusions are often enriched with metastatic tumor cells, potentially offering a more accurate representation of advanced disease biology than resected primary tumors [50]. Lastly, the establishment of FDOs does not require surgical intervention, thereby broadening their applicability to patients with inoperable or late-stage disease [51].
Establishment and Characterization Comparison of Malignant Pleural and Ascitic FDOs
| Sample Collection | Cell Isolation and Enrichment | Key components in Culture Medium | Drug Resistance Mechanism Simulation | Sampling Feasibility | |
|---|---|---|---|---|---|
| PE-organoids | Thoracentesis | Density gradient centrifugation | EGF, FGF, TGF-β inhibitor | EGFR-TKI resistance mutations (T790M) | High (repeatable puncture) |
| AS-organoids | Paracentesis | Magnetic bead separation/FACS | Wnt agonist, estrogen factor | Platinum/Taxane Resistance | High (common in recurrent ascites) |
| PDOs | Surgical biopsy | Mechanical/enzymatic digestion method | Tissue-specific cytokines | Primary tumor drug resistance profile | Low (surgery-dependent) |
Despite the advantages, several challenges remain. Effusion samples are inherently heterogeneous, often comprising immune cells, fibroblasts, and mesothelial cells [52]. Although a certain level of heterogeneity is generally deemed beneficial, it is crucial to recognize that an overabundance of non-tumor cells can impede the establishment of organoids [53]. Long-term culture tends to select for epithelial cells, leading to the depletion of stromal and immune components that are essential for accurately modeling tumor-microenvironment interactions [54]. Additionally, the lack of standardized protocols across laboratories hampers reproducibility and complicates cross-study comparisons [55]. Scalability also presents a challenge: while individual FDO cultures can be expanded for drug testing, the creation of large biobanks for high-throughput screening demands substantial resources [56].
FDOs occupy a distinct position within the spectrum of preclinical models. While 2D cell lines are well-documented for their cost-effectiveness and scalability, they fall short in representing cellular heterogeneity [57]. PDX models, on the other hand, maintain tumor complexity but are hindered by lengthy establishment times and the confounding presence of murine stroma [58]. Although tissue-derived PDOs offer physiological relevance, their utility is constrained by challenges related to surgical access and the initial sampling process. In contrast, FDOs provide a unique combination of accessibility, fidelity to the original tumor, and the capacity for longitudinal sampling, thereby serving as a complementary asset in the array of preclinical models [59].
Pleural Effusion-Derived Organoids in Drug Resistance Research
MPE is a common complication associated with advanced-stage malignancies, with lung cancer being the most frequent underlying cause [60]. Approximately 40% of patients with advanced NSCLC experience pleural effusions during the course of their illness, particularly in the context of treatment resistance and disease progression [61]. This study highlights the potential of generating organoids from pleural effusion-derived tumor cells (PE-organoids), providing a unique platform to explore resistance mechanisms, monitor disease evolution over time, and assess personalized therapeutic strategies [62].
Lung Cancer: EGFR-TKI Resistance
NSCLC, especially the adenocarcinoma subtype, is strongly linked to activating mutations in the EGFR [63]. Although EGFR tyrosine kinase inhibitors (EGFR-TKIs) have significantly improved outcomes for patients with EGFR-mutant NSCLC, it is crucial to recognize that nearly all patients eventually develop acquired resistance to these therapies [64]. To investigate this phenomenon in greater detail, research has utilized patient-derived ex vivo PE-organoids obtained from pleural effusion samples. These models have been employed to elucidate resistance mechanisms [65]. The secondary EGFR T790M mutation is responsible for approximately 50% of resistance to first- and second-generation TKIs and has been accurately maintained in PE-organoids, facilitating the functional assessment of third-generation TKIs such as Osimertinib [66]. Prolonged treatment with osimertinib leads to the emergence of additional resistance mechanisms, including the EGFR C797S mutation and activation of bypass pathways such as MET or HER2 amplification and alterations in the PI3K pathway [67, 68]. The modeling of these resistance features in PE-organoids has been a productive endeavor, as demonstrated by the successful documentation of the molecular evolution of resistance [69]. Beyond EGFR, PE-organoids have also been utilized to explore resistance in ALK-rearranged NSCLC [70]. Organoids derived from EML4-ALK-positive effusions have revealed secondary ALK mutations (e.g., L1196M, G1202R) that confer resistance to second-generation ALK inhibitors [71]. These findings underscore the versatility of PE-organoids as platforms for investigating resistance across various oncogene-driven subsets of NSCLC.
Immune Resistance in NSCLC
Immunotherapy utilizing immune checkpoint inhibitors (ICIs), particularly those targeting the PD-1/PD-L1 axis, has emerged as a fundamental component in the treatment of NSCLC [72]. Nevertheless, therapeutic responses are limited to a subset of patients, and the phenomenon of acquired resistance is becoming increasingly prevalent [73, 74]. Although most PDO systems are devoid of immune elements, pleural effusions frequently contain tumor-associated lymphocytes and macrophages [75, 76]. By co-culturing pleural effusion-derived organoids with autologous immune cells from the same effusion, researchers have begun to investigate mechanisms of immune resistance, such as alterations in macrophage polarization that may contribute to sustained resistance to PD-1 blockade [77]. These co-culture models hold promise for evaluating combination therapies designed to overcome immune resistance.
Other Cancers with Malignant Pleural Effusion
While lung cancer is the predominant cause of MPE, such effusions also occur in breast cancer, gastric cancer, and hematologic malignancies [78, 79]. Although research on breast cancer pleural effusion-derived organoids is limited, organoid models have been effectively employed to study resistance to endocrine and HER2-targeted therapies in breast cancer, suggesting that effusion-derived organoids could be similarly utilized for these investigations [80]. In gastric cancer, PE-organoids have been developed to assess sensitivity to platinum-based chemotherapy, yielding results that align with patient responses and thereby underscoring their predictive utility [81-83]. Initial efforts to culture organoids from lymphoma-associated pleural fluid have also been documented [84].
Longitudinal Monitoring of Resistance Evolution
A distinct advantage of PE-organoids is their applicability for longitudinal monitoring [85]. Patients experiencing recurrent MPE can undergo multiple thoracenteses, providing sequential samples that capture tumor evolution under therapeutic pressure [86]. Serial PE-organoids have effectively documented the transition from EGFR exon 19 deletion-sensitive disease to T790M-positive resistance, and subsequently to C797S-mediated resistance [87]. Drug testing on these sequential organoids has mirrored clinical outcomes, demonstrating the feasibility of real-time resistance monitoring [88]. Longitudinal sampling not only corroborates clinical observations but also reveals novel resistance mechanisms [89]. Changes such as the activation of epithelial-mesenchymal transition (EMT), histological transformation from adenocarcinoma to small cell lung cancer, and modifications in the tumor microenvironment have been identified in serial PE-organoids, underscoring the value of effusion-derived models for studying tumor plasticity and heterogeneity [90].
Advantages and Limitations of PE-Organoids
PE-organoids present several advantages, including minimally invasive sampling, the capacity to investigate advanced stages of disease, and a high degree of concordance with clinical resistance phenotypes [80]. They serve as a complement to tissue-derived PDOs and circulating tumor cell (CTC)-derived models by providing an abundant and viable source of tumor cells directly from metastatic lesions [91]. However, PE-organoids are established with less efficiency than ascites-derived organoids, primarily due to the lower concentration of tumor cells present in pleural fluid [92]. Additionally, over time, the immune and stromal components within the culture tend to diminish [93]. To facilitate their integration into routine clinical workflows, there is a need for both standardized culture methods and extensive clinical validation (Table 3).
Application of PE-organoids and AS-organoids
| Application of PE-organoids in drug resistance research | |||||
|---|---|---|---|---|---|
| Main drug resistance mechanisms | Drug test results and clinical relevance | Microenvironment retention status | Culture success rate | References | |
| NSCLC | 1.EGFR T790M mutation 2. MET/HER2 amplification 3.PI3K pathway activation | 1. The prediction accuracy for osimertinib sensitivity is 85%. 2. The response rate to immunotherapy is related to PD-L1 expression. | Partial retention of lymphocytes and macrophages | 30%-60% | [15,16,62,69] |
| Breast cancer | Abnormal HER2 signaling pathway, endocrine therapy resistance (ESR1 mutation) | HER2-targeted drug response is consistent with patient clinical | Significant loss of stromal cells | 20%-40% | [77-79] |
| Lymphoma | 1.B cell receptor signaling pathway activation 2.Immune checkpoint inhibitor resistance | The results of chemotherapy drug screening are related to patient survival | Lack of B cell-T cell interaction | <20% | [40,48,83] |
| Gastric Cancer | 1.Loss of HER2 amplification 2. FGFR2 overexpression | Trastuzumab resistance is consistent with HER2 downregulation in organoids. | Co-culturing with fibroblasts can enhance model stability. | 40%-50% | [77,78,102] |
| Application of AS-organoids in drug resistance research | |||||
| Ovarian cancer | 1.BRCA reversion mutation 2.Homologous recombination repair restoration 3.Upregulation of platinum efflux pumps | PARP inhibitor sensitivity prediction accuracy is 80% | Preserve CA125 secretion function | 60%-90% | [17,35,37] |
| Colorectal cancer | 1.KRAS/BRAF mutations 2.EGFR monoclonal antibody resistance (MET amplification) | Cetuximab resistance is consistent with KRAS mutations in organoids. | Co-culture tumor-associated macrophages | 50%-70% | [101,111] |
| Pancreatic cancer | 1.Stroma-mediated chemotherapy barrier 2.Deficiency of gemcitabine-metabolizing enzymes | FOLFIRINOX response prediction is associated with patients' progression-free survival | Stromal cells need to be added to maintain the viability of the model. | 40%-60% | [18,103-106] |
Ascites-Derived Organoids in Drug Resistance Research
MA is commonly observed in individuals with advanced stages of ovarian, gastric, pancreatic, and other abdominal cancers [94, 95]. The ascitic fluid is a rich reservoir of exfoliated tumor cells, immune cells, and stromal components, rendering it particularly advantageous for the cultivation of organoids [96]. In comparison to pleural effusions, malignant ascites typically exhibits higher tumor cellularity, which contributes to increased success rates in the establishment of organoids [97]. Organoids derived from ascites (AS-organoids) have been extensively utilized to model chemotherapy resistance, evaluate sensitivity to targeted therapies, and investigate personalized treatment strategies [18].
Ovarian Cancer
Among various cancers, ovarian cancer exhibits the strongest association with malignant ascites [98]. Although platinum- and taxane-based regimens remain the standard therapeutic approach, the majority of patients experience relapse due to the development of acquired resistance [99]. AS-organoids derived from ovarian cancer ascites effectively replicate the phenotypes associated with chemotherapy resistance and maintain critical genomic alterations pertinent to therapeutic interventions [18]. Furthermore, they facilitate the assessment of sensitivity to targeted therapies, such as PARP inhibitors, thus serving as ex vivo platforms for predicting treatment response [100, 101].
Gastrointestinal and Pancreatic Cancers
In gastrointestinal malignancies, such as metastatic colorectal cancer, organoids derived from ascitic fluid have been employed for drug sensitivity testing of fluoropyrimidines and platinum-based agents, demonstrating a correlation between organoid responses and patient outcomes [102]. Additionally, ascites-derived organoids have been utilized to explore resistance mechanisms to HER2-targeted therapies in HER2-positive gastric cancer [103]. In the realm of pancreatic cancer, organoids derived from ascites have been instrumental in drug screening efforts, including the assessment of FOLFIRINOX and gemcitabine/nab-paclitaxel treatment regimens, although research in this area remains limited [104-106].
Longitudinal Monitoring and Tumor Evolution
The recurrent nature of ascites allows for repeated sampling, facilitating the establishment of sequential organoids from the same patient [92]. This methodology has been applied to track the evolution of resistance in ovarian cancer, capturing transitions from platinum-sensitive to platinum-resistant states and identifying secondary BRCA mutations that restore homologous recombination [107, 108]. Furthermore, longitudinal monitoring of patient-derived pancreatic ductal adenocarcinoma (PDAC) organoids, utilizing live-cell imaging and deep learning techniques, has enabled dynamic evaluation of treatment responses, thereby providing a framework for capturing tumor evolution under therapeutic pressure [109].
Advantages and Limitations of AS-Organoids
AS-organoids typically demonstrate higher establishment rates and extended culture viability compared to PE-organoids, primarily due to the increased tumor cellularity present in ascites samples [110]. This characteristic is particularly advantageous in the context of ovarian cancer, where malignant cells are abundant. Additionally, the larger volumes of these samples facilitate higher-throughput drug screening [111]. However, it is important to note that stromal and immune components are gradually lost during prolonged culture periods, and reactive mesothelial cells may occasionally proliferate more rapidly than tumor cells, thereby complicating the culture process [112]. Furthermore, organoids derived from gastrointestinal and pancreatic cancers remain relatively underexplored, necessitating further validation of their clinical relevance. AS-organoids serve as ex vivo platforms for evaluating chemotherapies, targeted agents, and novel drug combinations, with the potential to personalize therapeutic approaches for patients with recurrent ascites [113]. The integration of genomic and transcriptomic analyses can enhance patient stratification and facilitate the identification of resistance biomarkers [114]. Ultimately, AS-organoids hold the potential to be incorporated into prospective clinical trials as predictive assays, thereby guiding individualized treatment strategies (Table 3) [115].
Resistance Mechanisms Across Cancer Types
Despite originating from different anatomical sites (pleural cavity vs. peritoneal cavity) and diverse cancer types, the acquired resistance mechanisms revealed by FDOs demonstrate remarkable commonalities. These shared mechanisms transcend specific histological origins, reflecting universal adaptive strategies of tumors under therapeutic pressure. The activation of EMT has been identified in longitudinal FDO models of both lung and ovarian cancers, associating with a more aggressive and therapy-resistant phenotype. Secondary driver gene mutations (e.g., EGFR T790M/C797S and ALK L1196M in lung cancer; BRCA reversion mutations in ovarian cancer) represent a key pathway for evading targeted therapies. The activation of bypass signaling pathways (e.g., MET/HER2 amplification, PI3K pathway alterations) is a common strategy for resistance to targeted agents across multiple cancers. Furthermore, alterations in the tumor microenvironment, including immune cell dysfunction (e.g., T-cell exhaustion, M2 macrophage polarization) and protective signals from stromal cells (e.g., mesothelial cells), collectively constitute a niche that supports tumor survival and mediates resistance to chemotherapy, targeted therapy, and immunotherapy. Highlighting these commonalities underscores the value of FDOs as powerful tools for systematically studying the universal principles of resistance biology across cancer types (Table 4).
Resistance mechanisms across cancer types modeled by FDOs
| Resistance Mechanism | Pleural Effusion-Derived Organoids | Ascites-Derived Organoids | References |
|---|---|---|---|
| Secondary Driver Gene Mutations | EGFR T790M/C797S (vs. EGFR-TKIs); ALK L1196M/G1202R (vs. ALK inhibitors) | BRCA1/2 reversion mutations (restoring HR, vs. PARP inhibitors) | [65-68, 70, 71, 106, 107] |
| Bypass Pathway Activation | MET/HER2 amplification, PI3K pathway alterations (vs. EGFR/ALK inhibitors) | Understudied (likely involves RTK reprogramming) | [66-68] |
| Histological / Cell Fate Transition | Adenocarcinoma to SCLC transformation; EMT activation | EMT activation; Dedifferentiation | [89, 108] |
| Altered DNA Damage Response | Possibly involved (linked to SCLC transformation) | Restored homologous recombination | [89, 106, 107] |
| Microenvironment-Mediated Protection | Immune suppression (exhausted T cells, M2 macrophages); Survival signals from mesothelial cells | Immune suppression; Stromal cell interactions; Unique cytokine milieu of ascites | [74-76, 95, 111] |
Translational Applications of FDOs in Personalized Oncology
Ex vivo Drug Sensitivity Testing
Ex vivo drug screening using organoids represents a promising approach for guiding therapeutic interventions [65]. are particularly advantageous due to the ease of obtaining effusion samples, even from patients with advanced-stage disease [116]. By cultivating tumor cells from pleural effusion or ascitic fluid within three-dimensional systems, FDOs can be evaluated against various therapeutic agents, with several studies demonstrating a correlation between these responses and patient outcomes [117]. The ability to repeatedly sample effusions facilitate the generation of sequential organoids from the same patient throughout the course of treatment. This capability allows for the dynamic investigation of resistance evolution and treatment adaptation, as serial pleural effusion- and ascites-derived organoids reflect clinical progression [118]. Sequencing FDOs offers insights into the molecular drivers of resistance and supports patient stratification for targeted therapies [119]. Furthermore, transcriptomic and single-cell analyses elucidate pathway reprogramming, immune evasion, and metabolic alterations, which can guide the development of rational therapeutic combinations [120]. Effusion samples frequently contain immune cells, enabling the establishment of co-culture systems with autologous lymphocytes or macrophages to explore mechanisms of immune resistance [121]. These models may improve prediction of ICI efficacy and support testing of combinations with targeted agents or PARP inhibitors [122].
Biobanking and Clinical Trials
Biobanks of FDO derived from pleural effusion and ascites samples serve as valuable resources for translational research and precision medicine trials [123]. Emerging prospective studies are integrating organoid-based assays into clinical decision-making processes, thereby establishing a foundation for large-scale validation [124]. Prior to widespread implementation, several challenges must be addressed, including the standardization of protocols, optimization of assay turnaround times, development of regulatory frameworks, and assessment of cost-effectiveness. Adequately powered prospective trials are necessary to substantiate that FDO-guided treatments enhance patient outcomes compared to standard care.
Technical and Standardization Challenges
The absence of standardized protocols for the establishment of FDOs leads to significant variability across laboratories in terms of sample processing, matrix selection, and media composition. The success rates of these protocols are contingent upon the type of tumor, which is further influenced by factors such as tumor cellularity, immune infiltration, previous treatments, and handling conditions [92, 125]. For the advancement of multi-center studies, it is imperative to develop reproducible and cost-effective protocols [126]. The immune, stromal, and mesothelial components present in effusions are frequently lost during culture, which restricts the ability to model interactions between tumors and their microenvironment [127]. To address this limitation, co-culture strategies involving autologous immune cells or cancer-associated fibroblasts, as well as organoid-on-chip platforms, are being developed to more accurately replicate the native microenvironment [128]. The integration of functional organoid data with genomic, transcriptomic, proteomic, and metabolomic analyses presents a complex challenge that necessitates robust bioinformatic infrastructure. Although artificial intelligence and machine learning approaches have the potential to synthesize high-dimensional data to predict drug responses, they require extensive validation [129, 130]. The expansion from single-patient cultures to large-scale biobanks demands coordinated infrastructure, standardized protocols, and adequate resources [131]. Comprehensive repositories of PE and AS organoids will significantly enhance drug discovery and preclinical validation [132].
The translation of FDO-based functional diagnostic tests from research tools to clinically actionable assays face significant regulatory hurdles, primarily centered on CLIA (Clinical Laboratory Improvement Amendments) certification and analytical validation [133]. For an FDO drug sensitivity test to be used in guiding patient therapy in the United States, it must be performed in a CLIA-certified laboratory [133, 134]. This mandates the development of and adherence to stringent, standardized protocols for every step-from sample processing and organoid culture to drug exposure and viability readout-ensuring rigorous quality control, personnel competency, and assay reproducibility [133].
The current lack of standardized protocols for FDO generation poses a fundamental challenge to meeting these requirements. Beyond laboratory standards, the assay itself must undergo extensive analytical validation to demonstrate its reliability. This involves formally establishing key parameters such as precision, accuracy, sensitivity, sample stability, and assay robustness [135]. Ultimately, establishing clinical validity-proving that the test result predicts patient outcome through large-scale prospective trials-is the critical final step. The path to regulatory approval thus requires a coordinated process of protocol standardization, analytical validation within a CLIA framework, and subsequent clinical validation, which represents a substantial but necessary undertaking for integrating FDO-based functional diagnostics into routine clinical practice.
Substantial evidence from prospective trials is essential to confirm the efficacy of FDO-guided treatment [136]. Critical considerations encompass turnaround time, assay thresholds for sensitivity and resistance, and the integration of findings into tumor boards [137]. The direct application of FDOs to guide clinical therapy selection or monitor resistance remains predominantly at the preclinical validation or early-stage clinical research phase, while the field is promising, studies providing definitive clinical predictive data are still limited. A clinical study specifically designed for FDOs, illustrating the complete pathway from research objectives to clinical application validation (NCT06658080), To evaluate the accuracy of FDOs in predicting the efficacy of chemotherapy/targeted therapy in patients with advanced breast cancer. The treatment decision window for advanced cancer is typically short (approximately 2-4 weeks). While traditional organoid culture requires 4-8 weeks, integrating new technologies like microfluidics has the potential to shorten the entire workflow to 1-2 weeks, which is crucial for real-time treatment guidance. Not all effusion samples can successfully yield FDOs, as this depends on the viability and quantity of tumor cells in the sample. Furthermore, the ability to retain key tumor microenvironment components during culture directly impacts the predictive accuracy for therapies like immunotherapy. Currently, there is a lack of uniform standards for FDO culture, drug testing, and result interpretation, making data comparison across different laboratories difficult. Additionally, the high cost of complex 3D culture and personalized testing presents a barrier to widespread adoption. Most current supporting evidence comes from retrospective or small-scale prospective studies. To establish the clinical utility of FDOs, it is essential to demonstrate through large-scale, prospective, interventional clinical trials that their use ultimately improves patient survival or quality of life. In summary, FDOs represent a highly promising tool for achieving precision cancer therapy, particularly in modeling late-stage metastatic foci and guiding treatment for refractory patients. However, their clinical application still faces practical challenges such as long turnaround times and a lack of standardization. This field is in a period of rapid development, and the integration of new technologies is key to overcoming existing bottlenecks.
Regulatory frameworks must address issues of reproducibility, quality control, and patient safety [138]. Promising avenues for advancement include the development of organoid-on-chip systems, immune-integrated organoids, AI-driven analytics, global biobanking networks, and the incorporation of these technologies into clinical trials. The convergence of technological innovations with rigorous clinical validation has the potential to transform FDOs from experimental tools into vital components of precision oncology.
Conclusion
Organoids derived from malignant effusions-pleural fluid and ascites-have established a powerful and clinically accessible platform for studying drug resistance. These FDOs complement existing models by capturing dynamic tumor evolution through serial sampling and preserving patient-specific resistance profiles. Although challenges such as variable culture success, protocol standardization, and loss of native microenvironment components remain, ongoing technological advances-in co-culture systems, organoid-on-chip platforms, and multi-omics integration-are actively addressing these gaps. Moving forward, establishing well-characterized FDO biobanks and incorporating organoid-guided assays into prospective clinical trials will be essential to validate their predictive utility. In summary, FDOs offer a minimally invasive, physiologically relevant, and highly adaptable model system that holds strong potential to illuminate resistance mechanisms and inform personalized treatment strategies in advanced cancer.
Acknowledgements
We thank Figdraw (www.figdraw.com) for expert assistance in the pattern drawing.
Funding
This work was supported by the National Natural Science Foundation of China (82073388); The Affiliated Hospital of Guangdong Medical University Clinical Research Program (LCYJ2020B005 and LCY2022DL002); the Natural Outstanding Youth Fund of Guangdong Province (2022B1515020090).
Author Contributions
Y.C., X.X., and J.C. contributed equally to this work. They performed the literature search, evidence synthesis, and drafted the manuscript. M.Y. and J.C. participated in reference curation and manuscript editing. Z.Q. assisted with critical revision. M.S. prepared the figures and tables, conceived the topic, and supervised the overall work together with W.S. All authors read and approved the final version of the manuscript.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Bray F, Laversanne M, Sung H. et al. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: a cancer journal for clinicians. 2024;74:229-63
2. Gonnelli F, Hassan W, Bonifazi M. et al. Malignant pleural effusion: current understanding and therapeutic approach. Respiratory research. 2024;25:47
3. Wang Y, Sun Y, Li X. et al. Progress in the treatment of malignant ascites. Critical reviews in oncology/hematology. 2024;194:104237
4. Yang Y, Du J, Wang YS. et al. Prognostic impact of pleural effusion in patients with malignancy: A systematic review and meta-analysis. Clinical and translational science. 2022;15:1340-54
5. Lin D, Wyatt AW, Xue H. et al. High fidelity patient-derived xenografts for accelerating prostate cancer discovery and drug development. Cancer research. 2014;74:1272-83
6. Marconi GD, Mazzone A, Della Rocca Y. et al. Three-Dimensional Culture System: A New Frontier in Cancer Research, Drug Discovery, and Stem Cell-Based Therapy. Biology. 2025 14
7. Meehan TF, Conte N, Goldstein T. et al. PDX-MI: Minimal Information for Patient-Derived Tumor Xenograft Models. Cancer research. 2017;77:e62-e6
8. Blomme A, Van Simaeys G, Doumont G. et al. Murine stroma adopts a human-like metabolic phenotype in the PDX model of colorectal cancer and liver metastases. Oncogene. 2018;37:1237-50
9. Rassomakhina NV, Ryazanova AY, Likhov AR. et al. Tumor Organoids: The Era of Personalized Medicine. Biochemistry Biokhimiia. 2024;89:S127-s47
10. Szulkin A, Szatmári T, Hjerpe A. et al. Chemosensitivity and resistance testing in malignant effusions with focus on primary malignant mesothelioma and metastatic adenocarcinoma. Pleura and peritoneum. 2016;1:119-33
11. De Wolf AM. Preoperative optimization of patients with liver disease. Current opinion in anaesthesiology. 2005;18:325-31
12. Chen J, Ma S, Yang H. et al. Generation and metabolomic characterization of functional ductal organoids with biliary tree networks in decellularized liver scaffolds. Bioactive materials. 2023;26:452-64
13. Rickard BP, Conrad C, Sorrin AJ. et al. Malignant Ascites in Ovarian Cancer: Cellular, Acellular, and Biophysical Determinants of Molecular Characteristics and Therapy Response. Cancers. 2021 13
14. Ahmed N, Stenvers KL. Getting to know ovarian cancer ascites: opportunities for targeted therapy-based translational research. Frontiers in oncology. 2013;3:256
15. Alvarez-Silva C, Schierwagen R, Pohlmann A. et al. Compartmentalization of Immune Response and Microbial Translocation in Decompensated Cirrhosis. Frontiers in immunology. 2019;10:69
16. Peng KC, Su JW, Xie Z. et al. Clinical outcomes of EGFR+/METamp+ vs. EGFR+/METamp- untreated patients with advanced non-small cell lung cancer. Thoracic cancer. 2022;13:1619-30
17. Wang CG, Zeng DX, Huang JA. et al. Effective assessment of low times MET amplification in pleural effusion after epidermal growth factor receptor-tyrosine kinase inhibitors (EGFR-TKIs) acquired resistance: Cases report. Medicine. 2018;97:e9021
18. Arias-Diaz AE, Ferreiro-Pantin M, Barbazan J. et al. Ascites-Derived Organoids to Depict Platinum Resistance in Gynaecological Serous Carcinomas. International journal of molecular sciences. 2023 24
19. Lau HCH, Kranenburg O, Xiao H. et al. Organoid models of gastrointestinal cancers in basic and translational research. Nature reviews Gastroenterology & hepatology. 2020;17:203-22
20. Yue L, Lan Z, Liu YJ. The Theoretical Estimation of the Bioluminescent Efficiency of the Firefly via a Nonadiabatic Molecular Dynamics Simulation. The journal of physical chemistry letters. 2015;6:540-8
21. Grossman JE, Muthuswamy L, Huang L. et al. Organoid Sensitivity Correlates with Therapeutic Response in Patients with Pancreatic Cancer. Clinical cancer research: an official journal of the American Association for Cancer Research. 2022;28:708-18
22. Liu L, Yu L, Li Z. et al. Patient-derived organoid (PDO) platforms to facilitate clinical decision making. Journal of translational medicine. 2021;19:40
23. Minoli M, Cantore T, Hanhart D. et al. Bladder cancer organoids as a functional system to model different disease stages and therapy response. Nature communications. 2023;14:2214
24. Anderson NM, Simon MC. The tumor microenvironment. Current biology: CB. 2020;30:R921-r5
25. Zhao Y, Shen M, Wu L. et al. Stromal cells in the tumor microenvironment: accomplices of tumor progression? Cell death & disease. 2023;14:587
26. Shidham VB. Collection and processing of effusion fluids for cytopathologic evaluation. CytoJournal. 2022;19:5
27. Jouneau S, Letheulle J, Desrues B. Repeated therapeutic thoracentesis to manage complicated parapneumonic effusions. Current opinion in pulmonary medicine. 2015;21:387-92
28. Che J, Mach AJ, Go DE. et al. Microfluidic purification and concentration of malignant pleural effusions for improved molecular and cytomorphological diagnostics. PloS one. 2013;8:e78194
29. Górny MK, Micksche M, Wolf A. Radioimmunometric demonstration of immunoglobulin G on cancer cells derived from malignant pleural effusions. Oncology. 1982;39:6-12
30. Johnson S, Chen H, Lo PK. Isolation of Cancer Epithelial Cells from Mouse Mammary Tumors. Bio-protocol. 2013 3
31. Schuger L, O'Shea KS, Nelson BB. et al. Organotypic arrangement of mouse embryonic lung cells on a basement membrane extract: involvement of laminin. Development (Cambridge, England). 1990;110:1091-9
32. Jee JH, Lee DH, Ko J. et al. Development of Collagen-Based 3D Matrix for Gastrointestinal Tract-Derived Organoid Culture. Stem cells international. 2019;2019:8472712
33. Treacy NJ, Clerkin S, Davis JL. et al. Growth and differentiation of human induced pluripotent stem cell (hiPSC)-derived kidney organoids using fully synthetic peptide hydrogels. Bioactive materials. 2023;21:142-56
34. Zhou T, Reji R, Kairon RS. et al. A review of algorithmic approaches for cell culture media optimization. Frontiers in bioengineering and biotechnology. 2023;11:1195294
35. Rahimmanesh I, Tavangar M, Zahedi SN. et al. Optimization of Culture Media for Ex vivo T-Cell Expansion for Adoptive T-Cell Therapy. Advanced biomedical research. 2022;11:94
36. Kopper O, de Witte CJ, Lõhmussaar K. et al. An organoid platform for ovarian cancer captures intra- and interpatient heterogeneity. Nature medicine. 2019;25:838-49
37. Kwan HCK. Reconsideration of the safety and effectiveness of human oocyte cryopreservation. Reproductive biology and endocrinology: RB&E. 2023;21:22
38. Hamilton TC, Ozols RF, Longo DL. Biologic therapy for the treatment of malignant common epithelial tumors of the ovary. Cancer. 1987;60:2054-63
39. Bielsa S, Hernández P, Rodriguez-Panadero F. et al. Tumor type influences the effectiveness of pleurodesis in malignant effusions. Lung. 2011;189:151-5
40. Aydogmus U, Ozdemir S, Cansever L. et al. Bedside talc pleurodesis for malignant pleural effusion: factors affecting success. Annals of surgical oncology. 2009;16:745-50
41. Huguet F, Rivin Del Campo E, Labidi M. et al. [Gastric and pancreatic cancers: Will neoadjuvant (chemo)radiotherapy replace adjuvant chemoradiotherapy?]. Cancer radiotherapie: journal de la Societe francaise de radiotherapie oncologique. 2020;24:493-500
42. Zulkowski K, Kath R, Semrau R. et al. Regression of brain metastases from breast carcinoma after chemotherapy with bendamustine. Journal of cancer research and clinical oncology. 2002;128:111-3
43. Snyder A, Nathanson T, Funt SA. et al. Contribution of systemic and somatic factors to clinical response and resistance to PD-L1 blockade in urothelial cancer: An exploratory multi-omic analysis. PLoS medicine. 2017;14:e1002309
44. Kazama A, Anraku T, Kuroki H. et al. Development of patient-derived tumor organoids and a drug testing model for renal cell carcinoma. Oncology reports. 2021 46
45. Vander Griend DJ, Konishi Y, De Marzo AM. et al. Dual-label centromere and telomere FISH identifies human, rat, and mouse cell contribution to Multispecies recombinant urogenital sinus xenografts. The Prostate. 2009;69:1557-64
46. Barzilai O, Martin A, Reiner AS. et al. Clinical reliability of genomic data obtained from spinal metastatic tumor samples. Neuro-oncology. 2022;24:1090-100
47. Bolck HA, Corrò C, Kahraman A. et al. Tracing Clonal Dynamics Reveals that Two- and Three-dimensional Patient-derived Cell Models Capture Tumor Heterogeneity of Clear Cell Renal Cell Carcinoma. European urology focus. 2021;7:152-62
48. Cai EY, Garcia J, Liu Y. et al. A bladder cancer patient-derived xenograft displays aggressive growth dynamics in vivo and in organoid culture. Scientific reports. 2021;11:4609
49. Fernandes D. Minimally invasive percutaneous collagen induction. Oral and maxillofacial surgery clinics of North America. 2005;17:51-63 vi
50. Harada K, Sakamoto N, Ukai S. et al. Establishment of oxaliplatin-resistant gastric cancer organoids: importance of myoferlin in the acquisition of oxaliplatin resistance. Gastric cancer: official journal of the International Gastric Cancer Association and the Japanese Gastric Cancer Association. 2021;24:1264-77
51. Geisbüsch A, Auer C, Dickhaus H. et al. Electromagnetic bone segment tracking to control femoral derotation osteotomy-A saw bone study. Journal of orthopaedic research: official publication of the Orthopaedic Research Society. 2017;35:1106-12
52. Verga L, Leni D, Cazzaniga G. et al. The spectrum of the cytopathological features of primary effusion lymphoma and human herpes virus 8-related lymphoproliferative disorders. Cytopathology: official journal of the British Society for Clinical Cytology. 2020;31:541-6
53. Inda MM, Bonavia R, Mukasa A. et al. Tumor heterogeneity is an active process maintained by a mutant EGFR-induced cytokine circuit in glioblastoma. Genes & development. 2010;24:1731-45
54. Gu Z, Wu Q, Shang B. et al. Organoid co-culture models of the tumor microenvironment promote precision medicine. Cancer innovation. 2024;3:e101
55. Novak V, Andeer PF, King E. et al. Breaking the reproducibility barrier with standardized protocols for plant-microbiome research. PLoS biology. 2025;23:e3003358
56. Luo Z, Wang B, Luo F. et al. Establishment of a large-scale patient-derived high-risk colorectal adenoma organoid biobank for high-throughput and high-content drug screening. BMC medicine. 2023;21:336
57. Braun FK, Rothhammer-Hampl T, Lorenz J. et al. Scaffold-Based (Matrigel™) 3D Culture Technique of Glioblastoma Recovers a Patient-like Immunosuppressive Phenotype. Cells. 2023 12
58. Jiang W, Xie S, Liu Y. et al. The Application of Patient-Derived Xenograft Models in Gynecologic Cancers. Journal of Cancer. 2020;11:5478-89
59. Trujillo CA, Muotri AR. Brain Organoids and the Study of Neurodevelopment. Trends in molecular medicine. 2018;24:982-90
60. Penz E, Watt KN, Hergott CA. et al. Management of malignant pleural effusion: challenges and solutions. Cancer management and research. 2017;9:229-41
61. Ibrahim FM, Helal DS, Ali DA. et al. Prognostic role of annexin A2 and cancer-associated fibroblasts in advanced non-small cell lung cancer: Implication in epithelial-mesenchymal transition and gefitinib resistance. Pathology, research and practice. 2023;241:154293
62. Pan B, Zhao D, Liu Y. et al. Breast cancer organoids from malignant pleural effusion-derived tumor cells as an individualized medicine platform. In vitro cellular & developmental biology Animal. 2021;57:510-8
63. Cortes-Funes H, Gomez C, Rosell R. et al. Epidermal growth factor receptor activating mutations in Spanish gefitinib-treated non-small-cell lung cancer patients. Annals of oncology: official journal of the European Society for Medical Oncology. 2005;16:1081-6
64. Du Z, Kan H, Sun J. et al. Molecular mechanisms of acquired resistance to EGFR tyrosine kinase inhibitors in non-small cell lung cancer. Drug resistance updates: reviews and commentaries in antimicrobial and anticancer chemotherapy. 2025;82:101266
65. Taverna JA, Hung CN, Williams M. et al. Ex vivo drug testing of patient-derived lung organoids to predict treatment responses for personalized medicine. Lung cancer (Amsterdam, Netherlands). 2024;190:107533
66. Soejima K, Yasuda H, Hirano T. Osimertinib for EGFR T790M mutation-positive non-small cell lung cancer. Expert review of clinical pharmacology. 2017;10:31-8
67. Zeng Y, Yu D, Tian W. et al. Resistance mechanisms to osimertinib and emerging therapeutic strategies in nonsmall cell lung cancer. Current opinion in oncology. 2022;34:54-65
68. Wu Z, Zhao W, Yang Z. et al. Novel Resistance Mechanisms to Osimertinib Analysed by Whole-Exome Sequencing in Non-Small Cell Lung Cancer. Cancer management and research. 2021;13:2025-32
69. Le Compte M, De La Hoz EC, Peeters S. et al. Single-organoid analysis reveals clinically relevant treatment-resistant and invasive subclones in pancreatic cancer. NPJ precision oncology. 2023;7:128
70. Peters S, Taron M, Bubendorf L. et al. Treatment and detection of ALK-rearranged NSCLC. Lung cancer (Amsterdam, Netherlands). 2013;81:145-54
71. Lin YT, Yu CJ, Yang JC. et al. Anaplastic Lymphoma Kinase (ALK) Kinase Domain Mutation Following ALK Inhibitor(s) Failure in Advanced ALK Positive Non-Small-Cell Lung Cancer: Analysis and Literature Review. Clinical lung cancer. 2016;17:e77-e94
72. Xia L, Liu Y, Wang Y. PD-1/PD-L1 Blockade Therapy in Advanced Non-Small-Cell Lung Cancer: Current Status and Future Directions. The oncologist. 2019;24:S31-s41
73. Lynes JP, Nwankwo AK, Sur HP. et al. Biomarkers for immunotherapy for treatment of glioblastoma. Journal for immunotherapy of cancer. 2020 8
74. Goggi JL, Khanapur S, Ramasamy B, et al. Imaging Kv1.3 Expressing Memory T Cells as a Marker of Immunotherapy Response. Cancers. 2022; 14
75. Yang R, Wang S, Li Z. et al. Patient-derived organoid co-culture systems as next-generation models for bladder cancer stem cell research. Cancer letters. 2025;625:217793
76. Chan WS, Mo X, Ip PPC. et al. Patient-derived organoid culture in epithelial ovarian cancers-Techniques, applications, and future perspectives. Cancer medicine. 2023;12:19714-31
77. Troiani T, Giunta EF, Tufano M. et al. Alternative macrophage polarisation associated with resistance to anti-PD1 blockade is possibly supported by the splicing of FKBP51 immunophilin in melanoma patients. British journal of cancer. 2020;122:1782-90
78. Pemmaraju N, Chang E, Daver N. et al. Extramedullary acute myeloid leukemia: leukemic pleural effusion, case report and review of the literature. Frontiers in oncology. 2014;4:130
79. Alexandrakis MG, Passam FH, Kyriakou DS. et al. Pleural effusions in hematologic malignancies. Chest. 2004;125:1546-55
80. Kim SC, Kim GH, Park JH. et al. Establishment and characterization of 24 breast cancer cell lines and 3 breast cancer organoids reveals molecular heterogeneity and drug response variability in malignant pleural effusion-derived models. Breast cancer research: BCR. 2025;27:66
81. McDonald HG, Harper MM, Hill K. et al. Creation of EGD-Derived Gastric Cancer Organoids to Predict Treatment Responses. Cancers. 2023 15
82. Zu M, Hao X, Ning J. et al. Patient-derived organoid culture of gastric cancer for disease modeling and drug sensitivity testing. Biomedicine & pharmacotherapy = Biomedecine & pharmacotherapie. 2023;163:114751
83. Zhao Y, Li S, Zhu L. et al. Personalized drug screening using patient-derived organoid and its clinical relevance in gastric cancer. Cell reports Medicine. 2024;5:101627
84. Chang L, Li L, Han Y. et al. Organoids in Haematologic Research: Advances and Future Directions. Cell proliferation. 2025;58:e13806
85. Wang Y, Guo Y, Wang P. et al. An engineered human placental organoid microphysiological system in a vascular niche to model viral infection. Communications biology. 2025;8:669
86. Ost DE, Goldblatt C, Jung M. et al. The Clinical and Economic Implications of Different Treatment Pathways for Patients With Rapidly Recurrent Malignant Pleural Effusion. Chest. 2024;166:867-81
87. Kobayashi N, Katakura S, Fukuda N. et al. The Impact of Bevacizumab and miR200c on EMT and EGFR-TKI Resistance in EGFR-Mutant Lung Cancer Organoids. Genes. 2024 15
88. Del Re M, Rofi E, Cappelli C. et al. The increase in activating EGFR mutation in plasma is an early biomarker to monitor response to osimertinib: a case report. BMC cancer. 2019;19:410
89. Mackenzie IZ, Roseman F, Findlay J. et al. Clinical validation of routine antenatal anti-D prophylaxis questions the modelling predictions adopted by NICE for Rhesus D sensitisation rates: results of a longitudinal study. European journal of obstetrics, gynecology, and reproductive biology. 2008;139:38-42
90. Chakraborty S, Coleman C, Manoj P. et al. De Novo and Histologically Transformed Small-Cell Lung Cancer Is Sensitive to Lurbinectedin Treatment Through the Modulation of EMT and NOTCH Signaling Pathways. Clinical cancer research: an official journal of the American Association for Cancer Research. 2023;29:3526-40
91. Farshbaf A, Lotfi M, Zare R. et al. The organoid as reliable cancer modeling in personalized medicine, does applicable in precision medicine of head and neck squamous cell carcinoma? The pharmacogenomics journal. 2023;23:37-44
92. Choi W, Kim YH, Woo SM. et al. Establishment of Patient-Derived Organoids Using Ascitic or Pleural Fluid from Cancer Patients. Cancer research and treatment. 2023;55:1077-86
93. Xue B, Schüler J, Harrod CM. et al. A Novel Hydrogel-Based 3D In vitro Tumor Panel of 30 PDX Models Incorporates Tumor, Stromal and Immune Cell Compartments of the TME for the Screening of Oncology and Immuno-Therapies. Cells. 2023 12
94. Huang Y, Zou K, Jiang H. et al. The complex role of IL-10 in malignant ascites: a review. Cancer immunology, immunotherapy: CII. 2024;73:32
95. Cavazzoni E, Bugiantella W, Graziosi L. et al. Malignant ascites: pathophysiology and treatment. International journal of clinical oncology. 2013;18:1-9
96. Kumar V, Mukhopadhyay A. Monitoring Natural Killer Cell Function in Human Ovarian Cancer Cells of Ascitic Fluid. Bio-protocol. 2018;8:e3124
97. Kalantri Y, Naik G, Joshi SP. et al. Role of cancer antigen-125 from pleural & ascitic fluid samples in non malignant conditions. The Indian journal of medical research. 2007;125:25-30
98. Lane D, Matte I, Rancourt C. et al. The prosurvival activity of ascites against TRAIL is associated with a shorter disease-free interval in patients with ovarian cancer. Journal of ovarian research. 2010;3:1
99. Webber K, Friedlander M. Chemotherapy for epithelial ovarian, fallopian tube and primary peritoneal cancer. Best practice & research Clinical obstetrics & gynaecology. 2017;41:126-38
100. Majumder B, Baraneedharan U, Thiyagarajan S. et al. Predicting clinical response to anticancer drugs using an ex vivo platform that captures tumour heterogeneity. Nature communications. 2015;6:6169
101. Goh J, De Mel S, Hoppe MM. et al. An ex vivo platform to guide drug combination treatment in relapsed/refractory lymphoma. Science translational medicine. 2022;14:eabn7824
102. Smabers LP, Wensink E, Verissimo CS. et al. Organoids as a biomarker for personalized treatment in metastatic colorectal cancer: drug screen optimization and correlation with patient response. Journal of experimental & clinical cancer research: CR. 2024;43:61
103. Zou Y, Zheng S, Xie X. et al. N6-methyladenosine regulated FGFR4 attenuates ferroptotic cell death in recalcitrant HER2-positive breast cancer. Nature communications. 2022;13:2672
104. Kim JS, Park CH, Kim E. et al. Establishing 3D organoid models from patient-derived conditionally reprogrammed cells to bridge preclinical and clinical insights in pancreatic cancer. Molecular cancer. 2025;24:162
105. Springfeld C, Jäger D, Büchler MW. et al. Chemotherapy for pancreatic cancer. Presse medicale (Paris, France: 1983). 2019;48:e159-e74
106. Seppälä TT, Zimmerman JW, Suri R. et al. Precision Medicine in Pancreatic Cancer: Patient-Derived Organoid Pharmacotyping Is a Predictive Biomarker of Clinical Treatment Response. Clinical cancer research: an official journal of the American Association for Cancer Research. 2022;28:3296-307
107. Khalique S, Pettitt SJ, Kelly G. et al. Longitudinal analysis of a secondary BRCA2 mutation using digital droplet PCR. The journal of pathology Clinical research. 2020;6:3-11
108. Ng CK, Cooke SL, Howe K. et al. The role of tandem duplicator phenotype in tumour evolution in high-grade serous ovarian cancer. The Journal of pathology. 2012;226:703-12
109. Ferreira N, Kulkarni A, Agorku D. et al. OrganoIDNet: a deep learning tool for identification of therapeutic effects in PDAC organoid-PBMC co-cultures from time-resolved imaging data. Cellular oncology (Dordrecht, Netherlands). 2025;48:101-22
110. Chen W, Fang PH, Zheng B. et al. Effective Treatment for Recurrent Ovarian Cancer Guided by Drug Sensitivity from Ascites-Derived Organoid: A Case Report. International journal of women's health. 2023;15:1047-57
111. Duraki D, Jabeen M, Mao C. et al. A necrosis inducer promotes an immunogenic response and destroys ovarian cancers in mouse xenografts and patient ascites organoids. Cancer letters. 2025;625:217738
112. Li X, Larsson P, Ljuslinder I. et al. Ex vivo Organoid Cultures Reveal the Importance of the Tumor Microenvironment for Maintenance of Colorectal Cancer Stem Cells. Cancers. 2020 12
113. Chen H, Gotimer K, De Souza C. et al. Short-term organoid culture for drug sensitivity testing of high-grade serous carcinoma. Gynecologic oncology. 2020;157:783-92
114. Nagasawa S, Zenkoh J, Suzuki Y. et al. Spatial omics technologies for understanding molecular status associated with cancer progression. Cancer science. 2024;115:3208-17
115. Soto-Gamez A, Gunawan JP, Barazzuol L. et al. Organoid-based personalized medicine: from tumor outcome prediction to autologous transplantation. Stem cells (Dayton, Ohio). 2024;42:499-508
116. Nishie R, Tanaka T, Hirosuna K. et al. Creation and Validation of Patient-Derived Cancer Model Using Peritoneal and Pleural Effusion in Patients with Advanced Ovarian Cancer: An Early Experience. Journal of clinical medicine. 2024 13
117. Hamilton G, Rath B. Applicability of tumor spheroids for in vitro chemosensitivity assays. Expert opinion on drug metabolism & toxicology. 2019;15:15-23
118. Boretto M, Maenhoudt N, Luo X. et al. Patient-derived organoids from endometrial disease capture clinical heterogeneity and are amenable to drug screening. Nature cell biology. 2019;21:1041-51
119. Hocking AJ, Mortimer LA, Farrall AL. et al. Establishing mesothelioma patient-derived organoid models from malignant pleural effusions. Lung cancer (Amsterdam, Netherlands). 2024;191:107542
120. Chen SL, Fei YR, Cai XX. et al. Exploring the role of metabolic pathways in TNBC immunotherapy: insights from single-cell and spatial transcriptomics. Frontiers in endocrinology. 2024;15:1528248
121. Halla JT, Schrohenloher RE, Volanakis JE. Immune complexes and other laboratory features of pleural effusions: a comparison of rheumatoid arthritis, systemic lupus erythematosus, and other diseases. Annals of internal medicine. 1980;92:748-52
122. Takigami A, Mato N, Hagiwara K. et al. A novel bioassay reflecting response to immune checkpoint inhibitor therapy in non-small cell lung cancer with malignant pleural effusion. Translational lung cancer research. 2024;13:3267-77
123. Huang CY, Huang WK, Yeh KY. et al. Integrating comprehensive genomic profiling in the management of oncology patients: applications and challenges in Taiwan. Biomedical journal. 2025: 100851.
124. Kim SY, van de Wetering M, Clevers H. et al. The future of tumor organoids in precision therapy. Trends in cancer. 2025;11:665-75
125. Navarro P, Grazioso TP, Barquín A. et al. Multicenter study correlating molecular characteristics and clinical outcomes of cancer cases with patient-derived organoids. Journal of experimental & clinical cancer research: CR. 2025;44:182
126. Rieck S, Sharma K, Altringer C. et al. Forward programming of human induced pluripotent stem cells via the ETS variant transcription factor 2: rapid, reproducible, and cost-effective generation of highly enriched, functional endothelial cells. Cardiovascular research. 2024;120:1472-84
127. Jagirdar RM, Rouka E, Pitaraki E. et al. Effects of patient pleural effusion fluids on the BBSome components expression of human benign mesothelial cells. American journal of physiology Lung cellular and molecular physiology. 2025;328:L105-l12
128. Veith I, Mencattini A, Picant V. et al. Apoptosis mapping in space and time of 3D tumor ecosystems reveals transmissibility of cytotoxic cancer death. PLoS computational biology. 2021;17:e1008870
129. Ferreira FJN, Carneiro AS. AI-Driven Drug Discovery: A Comprehensive Review. ACS omega. 2025;10:23889-903
130. Zou X, Liu Y, Ji L. Review: Machine learning in precision pharmacotherapy of type 2 diabetes-A promising future or a glimpse of hope? Digital health. 2023;9:20552076231203879
131. Smith ME, Sanderson SC, Brothers KB. et al. Conducting a large, multi-site survey about patients' views on broad consent: challenges and solutions. BMC medical research methodology. 2016;16:162
132. Zhou Z, Cong L, Cong X. Patient-Derived Organoids in Precision Medicine: Drug Screening, Organoid-on-a-Chip and Living Organoid Biobank. Frontiers in oncology. 2021;11:762184
133. Zhou L, Huang J, Li C. et al. Organoids and organs-on-chips: Recent advances, applications in drug development, and regulatory challenges. Med (New York, NY). 2025;6:100667
134. Kanagal-Shamanna R, Singh RR, Routbort MJ. et al. Principles of analytical validation of next-generation sequencing based mutational analysis for hematologic neoplasms in a CLIA-certified laboratory. Expert review of molecular diagnostics. 2016;16:461-72
135. Li S, Looby N, Chandran V. et al. Challenges in the Metabolomics-Based Biomarker Validation Pipeline. Metabolites. 2024 14
136. He W, Xie J, Wang Z. et al. Neoadjuvant treatment patterns and biomarker selection in muscle-invasive bladder cancer. Discover oncology. 2025;16:1197
137. Dive L, Critchley C, Otlowski M. et al. Public trust and global biobank networks. BMC medical ethics. 2020;21:73
138. Luchini C, Lawlor RT, Milella M. et al. Molecular Tumor Boards in Clinical Practice. Trends in cancer. 2020;6:738-44
Author contact
Corresponding authors: suwenmei123com (W.S.), ming1899edu.cn (M.S.).

 Global reach, higher impact
Global reach, higher impact