Impact Factor ISSN: 1837-9664
J Cancer 2026; 17(4):759-770. doi:10.7150/jca.132848 This issue Cite
Review
Roles of Cellular Neighborhoods in Lung Cancer
1. Department of Pulmonary Oncology, Zhongnan Hospital of Wuhan University, Wuhan, Hubei China.
2. Department of Radiation Oncology, The Affiliated Cancer Hospital of Zhengzhou University & Henan Cancer Hospital, Zhengzhou, Henan China.
Received 2026-2-8; Accepted 2026-3-13; Published 2026-3-25
Abstract
The occurrence and development of lung cancer (LC) involve complex interactions between various cell types in the tumor microenvironment (TME). Understanding the spatial distribution and interaction mechanisms of these cells may be the key to overcoming LC. The advancements of single-cell and spatial transcriptome techniques have promoted our understanding of cellular neighborhoods (CNs) and their functions in the pathogenesis of LC. In this review, we focus on the impact of different etiologies on LC CNs and the current research status of CNs in LC. This review may provide new insights into the molecular mechanisms of LC pathogenesis, develop more refined classification principles for LC diagnosis, and offer new perspectives for LC treatment.
Keywords: Cellular neighborhoods, Lung cancer, Spatial transcriptome, Immune microenvironment
Introduction
Thanks to the improvement of low-dose CT screening and advances in targeted therapy and immunotherapy, the mortality rate of lung cancer (LC) has significantly decreased in the past decade [1]. But LC remains the leading cause of cancer death. Non-small cell lung cancer (NSCLC) is the main type of LC, accounting for approximately 80-85% of all LC cases [2]. The development of LC is a heterogeneous and multi-step process involving multiple genetic and epigenetic changes which help tumor cells escape immune responses, promote cancer progression and metastasis [3].
Cellular neighborhoods (CNs) refer to the spatial relationships and interactions between specific cells and their surrounding cells in tissues or cell populations [4, 5]. This concept is commonly used to describe the distribution pattern of cells in a specific microenvironment, emphasizing the interactions and functional connections between cells [6-8]. Through CN analysis, researchers can better understand the complex structure and function of tissues [9, 10]. For example, in the tumor microenvironment (TME), different types of immune cells, tumor cells and their interactions form specific neighborhoods that may affect tumor growth and metastasis [11, 12]. CN analysis also helps identify intercellular signaling and interaction networks, which is crucial for understanding cancer progression. Because these interactions often determine the development and treatment response of cancer. A deeper understanding of the neighborhood relationship between immune cells and tumor cells in the TME can help develop new immune therapies.
The etiology of LC is still not fully understood, and the main pathogenic factors include smoking, occupational exposure, air pollution, ionizing radiation, pulmonary history, etc [13-17]. When pathogenic factors disrupt the homeostasis of the lungs, it ultimately leads to LC. The composition of LC is not simply the accumulation of tumor cells, but a microenvironment composed of fibroblasts, immune cells, extracellular matrix, and microvessels [18, 19]. There is a mutually restrictive relationship between tumor cells and the microenvironment [20]. The interaction between cells is crucial at every stage of LC development [21, 22]. Therefore, identifying LC-related CNs and understanding their functions are the key to deciphering the pathogenesis of LC. The advancement of technologies such as spatial transcriptome and single-cell sequencing has provided new avenues for studying CNs. Understanding the spatial environment and intercellular communication of cells in the microenvironment can reveal the complex pathogenesis of LC.
This article reviews the role of CNs in the pathogenesis of LC, analyzes the impact of different etiologies on CNs in LC, and the current research status of CNs in LC. This review describes the cell types involved in CNs of LC, the functions of CNs and their significance in the occurrence and development of LC. It also explores the key signaling pathways that may drive the formation of specific CNs. This review can improve our understanding of the pathogenesis of LC and provide new ideas for the treatment of LC by dissecting the CNs of LC.
1. Etiology-related cellular neighborhoods in LC (Figure 1)
1.1 Smoking
Smoking is associated with the onset of metabolic syndrome [23]. Metabolic syndrome is generally accompanied by abnormalities in blood glucose, blood pressure, and blood lipids [24]. Especially high blood glucose and abnormal blood lipids are related to the formation and progression of LC [25, 26]. The changes in glucose metabolism and lipid metabolism mainly affect tumor associated macrophages (TAMs) [27]. TAM, as one of the main cell types of TME, plays an important role in tumor progression through various mechanisms by secreting proteases, inflammatory mediators, growth factors, and disrupting anti-tumor immunity [28]. Smoking not only increases the metabolism of LC cells, but also promotes the progression of LC by altering the immune regulatory function of TAMs [29]. Studies showed that exposure to cigarette smoke extract (CSE) could induce macrophage M2 polarization. CSE also promoted macrophage to secrete vesicles containing circEML4. After uptaking by NSCLC cells, it interacted with intracellular ALKBH5 to promote m6A modification of SOX2, ultimately promoting NSCLC progression [30]. In addition, the dysfunction of alveolar cells caused by smoking could promote the invasiveness of lung adenocarcinoma (LUAD).
Etiology-related CNs in LC. Etiology-related CNs, such as smoking, air pollution, ionizing radiation and lung injury, are shown. Air pollution promotes IL-1β secretion of macrophage, induces the malignant transformation of normal lung cells carrying carcinogenic mutations (EGFR or KRAS). Smoking promotes the aggregation of NK cells, B cells and TAM in LC CN, as well as the expression of HLA in AM. Lung injury promotes the formation of three types of niches. One enriches macrophages, one enriches DCs, B cells and T cells, and another enriches myofibroblast and abnormal basal cells. Radiotherapy regulates the immune balance of LC. Immunosuppressive CNs enrichment CAFs, MDSCs, TAMs and Tregs. Immunoenhancement CNs enrichment B cells, T cells, NK cells and neutrophils. All the above pathological factors will promote LC progression to some extent. Abbreviations: TAM, tumor associated macrophage; CAF, cancer associated fibroblast; MDSC, bone marrow-derived suppressor cell; NE, neutrophil; NK, natural killer; AM, alveolar macrophage; DC, dendritic cell; Treg, regulatory T cell.
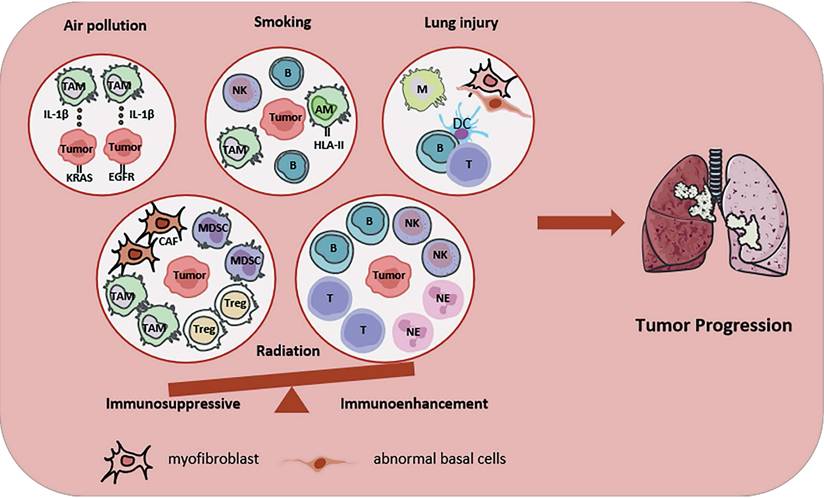
Single-cell profiling of tumor specimens and distal normal lung tissue from 22 LUAD patients (13 non-smokers and 9 smokers) demonstrated that compared to normal lung tissue, tumor tissue had a lower proportion of natural killer cells (NK cells) and a higher proportion of B lymphocytes [31]. Single-cell RNA sequencing (scRNA-seq) of normal lung cells adjacent to tumors showed that smoking led to upregulation of HLA-II and pro-inflammatory pathways (especially defense response to virus and response to type I interferon) in alveolar macrophages (AMs) [32]. Another study utilized single-cell transcriptome and epigenetic sequencing to establish a single-cell multi-omics map of human lung tissue. The study elucidated the key roles of epithelial cells and immune cells in the evolution of LC, and identified LC susceptibility genes, providing precise diagnostic targets and key molecular phenotypes with potential clinical application value for subsequent drug development and intervention [33]. An important study revealed how smoking drove the occurrence of early lung squamous cell carcinoma (LUSC) by disrupting the clonal dynamics of basal cells, providing a new perspective for understanding the “field cancerization” of LC [34].
1.2 Air pollution
Particulate matter (PM) is a variety of solid and liquid particles in the atmosphere, which has been identified as the main cause of LC [35]. PM2.5 is a particle with aerodynamic size ≤ 2.5 μm, which can lead to an increase in the incidence rate and mortality of LC in the population [36]. Engine exhaust and air pollution induce LC through genotoxicity, oxidative stress, and inflammation [37, 38]. Research revealed that air pollution significantly promoted the accumulation of mutations in the LC genome, and the mutation load was dose-dependent with PM2.5 concentration, suggesting that air pollution was not only a mutagenic source of LC, but also a promoting factor for tumor development [39]. A study found a new mechanism by which PM2.5 drove LC. PM2.5 did not directly cause cancer mutations in lung cells, but rather promoted the entry of a large number of macrophages into lung tissue, releasing IL-1β to create an inflammatory environment, leading to the malignant transformation of normal lung cells carrying carcinogenic mutations (EGFR or KRAS), thereby causing cancer [40].
1.3 Ionizing radiation
Ionizing radiation can induce the formation of immunogenic or immunosuppressive TME [41]. If the balance shifts towards an immunogenic phenotype, radiation will cause tumor cells to release new antigens and damage associated molecular patterns (DAMPs) [42]. These signals in turn lead to an increase in antigen presentation, thereby activating the innate immune system, increasing CD8+ cytotoxic T cell infiltration, and inhibiting immune suppressive cells [43]. If the balance tilts towards the immunosuppressive phenotype, radiation will kill T lymphocytes in the TME, increase infiltration of bone marrow-derived suppressor cells (MDSCs) and regulatory T cells (Tregs) and promote the activation of cancer associated fibroblasts (CAFs), thereby promoting tumor growth [44, 45]. Radiation not only has a regulatory effect on TME, but can also alter the immune characteristics of patients. As shown in the meta-analysis of different cancer types, patients experienced a systemic decrease in CD3+ and CD4+ peripheral T cells one month after the last radiotherapy [46]. Another retrospective study evaluated the effect of RT on programmed cell death-ligand 1 (PD-L1) expression and CD8+ T cell infiltration in NSCLC patients. While PD-L1 expression in cancer cells showed no consistent trend, the density of CD8+ T cells increased after radiation [47]. In addition, research showed that T cells, B cells, neutrophils and NK cells were enriched in the radiation sensitive group [48].
Radiation also reshape vascular system in TME. Radiation-induced endothelial cell dysfunction is characterized by increased permeability and apoptosis. Damage to vascular endothelial cells leads to increased expression of intercellular adhesion molecule (ICAM) and vascular cell adhesion molecule (VCAM), which play a key role in leukocyte migration and activation [49]. Endothelial cell damage can also attract natural immune cells to the TME. Radiotherapy causes hypoxia by damaging blood vessels, and further exacerbates TME hypoxia by consuming oxygen and producing toxic substances [50]. Hypoxia activates the hypoxia-inducible factor-1 (HIF-1) signaling pathway and promotes angiogenesis through vascular endothelial growth factor (VEGF) [51]. Researchers analyzed single-cell data from NSCLC patients and found that macrophages with hypoxia had pro-angiogenic activity and may be potential therapeutic targets for NSCLC patients [52].
On the other hand, the activation of CAFs after radiation leads to changes in growth factor secretion and the release of numerous extracellular matrix (ECM) regulators which can promote angiogenesis, invasion, and metastasis of cancer cells [53]. Research showed that alveolar epithelial cells type II underwent early apoptosis in direct response to ionizing radiation to clear damaged cells. Epithelial cell apoptosis destroyed the alveolar structure, causing the body to be unable to effectively clear damaged cells in a timely manner. At the same time, it was accompanied by a strong inflammatory response, exacerbating radiation-induced lung injury (RILI) and promoting the occurrence of LC [54].
1.4 Lung injury
Most NSCLC patients have a smoking history, which can lead to fibrosis and scar formation in lung tissue. In addition, diseases with increased pulmonary fibrosis, such as idiopathic pulmonary fibrosis (IPF), are associated with an increased incidence rate of LC [55, 56]. Researchers used spatial single-cell proteomics to analyze 6 cases of IPF and 3 cases of normal lung samples, successfully identifying multiple cell types in lung tissue. Among them, CSMD1+ fibroblasts were enriched in fibrotic areas and co-localized with pro-fibrotic molecules such as TGF-β1 and CTHRC1. CD248+ fibroblasts existed in low inflammatory areas and were associated with the anti-fibrotic factor MGP [57].
To unravel the immune cell landscape of the lung and identify the mechanisms driving injury-associated immune pathology, spatial transcriptome analysis of lung tissues from healthy individuals and those with IPF was conducted, revealing that the majority of differentially expressed genes were located in the fibrotic region, with multiple chemokines highly expressed at the fibrotic edge and reduced expression levels in the alveoli. On the contrary, ECM related genes (COL1A1, COL1A2, and LUM) were lowly expressed at the edge of fibrosis and highly expressed in fibrosis and adjacent alveoli [58]. Another study used spatial transcriptomics and the scRNA-seq to determine the unique cellular composition and localization of three disease-related niches. Fibrotic niche was composed of myofibroblasts and abnormal basal cells. Macrophage niche was located around the airway and adjacent to the airway lumen. Immune niche was characterized by obvious lymphocyte foci in fibrotic tissue. This spatial characterization of IPF niche will help identify drug targets and aid in the development of disease-related in vitro models [59]. The study used image-based spatial transcriptomics techniques to explore the evolution of alveolar niche dysbiosis in IPF and identified 12 ecological niches with unique gene expression characteristics and cell type composition, including cell ecological niche C11 associated with macrophage accumulation in the airspace, with FABP4+ and SPP1+ two main populations. In the IPF sample, the distribution and proportion of these two types of macrophages changed with disease progression, indicating that the evolution of macrophage phenotype was related to the progression of IPF [60].
2. Cellular neighborhoods in LC
Tumor heterogeneity refers to the molecular biology or genetic changes that occur during the progression of tumors, resulting in differences in the growth rate, invasion ability, and drug sensitivity of different tumor cells [61, 62]. It is a crucial characteristic in the process of tumor occurrence and development, a supporting point for tumor progression, and a key theory discovered through continuous exploration in recent years [63-65]. Tumor heterogeneity not only affects diagnosis, but also has an impact on treatment, efficacy, disease monitoring, drug resistance, and prognosis [66, 67]. The TME is the main source of heterogeneity in LC [68, 69]. Therefore, in-depth analysis of the spatial landscape of the immune microenvironment in LC is crucial. Such analysis can aid in understanding its pathogenesis, screening disease biomarkers, and developing novel treatment strategies. With the application of technologies such as spatial transcriptomics and single-cell sequencing, we have gained a deeper understanding of the heterogeneity of CNs in LC [70].
2.1 Immune cell-related cellular neighborhoods (Figure 2)
2.1.1 T cell-related cellular neighborhoods
T cells are an important component of the immune system in combating cancer, and the degree of T cell infiltration is positively correlated with the patient's treatment response and prognosis [71, 72]. The spatial distribution of T cells in tumor tissue is closely related to the TME, and the microenvironment characteristics of different regions determine the functional status of T cells [73, 74].
Representative immune cell-related CNs in LC. T cell-associated CNs, macrophage-related CNs, TLSs and cancer associated fibroblast-related CNs, reprogramming of the TME and interactions with neighboring cells, are shown. Abbreviations: Trm, tissue-resident memory T cells; Tem, effective memory T Cells; DC, dendritic cell; CAF, cancer associated fibroblast; TLS, tertiary lymphoid structure; Treg, regulatory T cell.
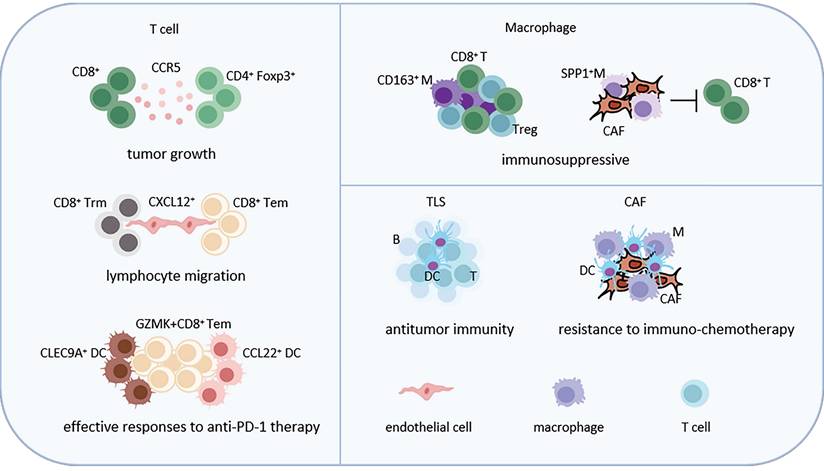
CD8+ T cells promoted the infiltration of CD4+Foxp3+ Tregs and accelerated the growth of murine LUAD. This was achieved by increasing the levels of CCR5 chemokines in the TME in an IFN-γ and TNF-α dependent manner [75]. A study used scRNA-seq to map the TME of NSCLC patients after receiving PD-1 combined chemotherapy, revealing that the CD4+ Tregs count was higher in the group that did not reach pathological remission, and the distribution of these cell types varied among the different groups. ZNF683+CD8+ tissue-resident memory T cells (Trm) and S100B+ NK cells were enriched in the non-pathological remission group, while GZMK+CD8+ effective memory T Cells (Tem) were enriched in the pathological remission group. Cell-chat analysis showed that in major pathological response (MPR) and non MPR tumors, CXCL12+ endothelial cells strongly interacted with CD8+ Trm and CD8+ Tem cells, and might participate in lymphocyte migration through the CXC12-CXCR4 axis in the MPR group. The study also found that CLEC9A+ cDC1 and CCL22+ mDC were highly correlated with GZMK+CD8+ Tem cells through specific ligand receptor interactions [76]. These observations provide a deeper understanding of how the cellular niches influence T cell functions in LC.
2.1.2 Macrophage cell-related cellular neighborhoods
TAMs have multiple functions, including promoting tumor growth, invasion, metastasis, and immune escape, and their functional diversity reflects the heterogeneity of cells [77, 78]. The functional diversity of TAMs is not only influenced by their origin, but also regulated by local factors such as tumor type, organs, and microenvironment [79, 80].
The results of single-cell sequencing showed that macrophages were the most common cell population in the immune microenvironment of myeloid cells and macrophages with CD163 positivity had the highest enrichment level in solid tumors. In addition, CD163+ macrophages were strongly correlated with Treg and CD8+ T cells, indicating a potential interaction between macrophages and T cell populations in the immune microenvironment of NSCLC [81]. The spatial transcriptome data revealed that tumor cells interact with SPP1+ macrophages and COL11A1+ CAFs, promoting the deposition and entanglement of collagen fibers at the tumor boundary, hindering T cell infiltration, and leading to adverse outcomes [82]. By using scRNA-seq to analyze macrophages in NSCLC tissues, further investigation found that they could be subdivided into seven main subgroups, including two monocyte macrophage subgroups and five macrophage subgroups. Each subgroups had different inflammation related gene expression characteristics. Among them, IL1B+and FCN1+ subgroups were more abundant in adjacent cancer tissues, while LGMN+ and APOE+ subgroups had a higher degree of infiltration in tumor tissues. Macrophages derived from tumor tissue exhibit an anti-inflammatory phenotype and are associated with poorer progression free survival (PFS) in patients [83].
In addition, research showed that pseudomonas aeruginosa promoted the secretion of CXCL9 by TAMs and enhances CD8+ T cell activity, thereby enhancing the immune response of NSCLC to anti-PD-1 therapy [84]. Understanding the mechanism of immune suppression mediated by macrophages can develop the targeted intervention strategies aimed at macrophages, thereby enhancing the efficacy of LC immunotherapy.
2.1.3 Tertiary lymphoid structures
There are different states of tertiary lymphoid structures (TLSs) in the TME, among which activated TLSs are associated with good prognosis, while the hypoxic microenvironment inhibits the development of TLSs and is associated with poor prognosis in LC [85-87].
Single-cell sequencing of NSCLC patients receiving PD-1 combined chemotherapy identified several unique subtypes of memory B cells: IGHG4+ plasma cells, SCGB3A1+, IFITM3+, RBFOX2+ memory B cells. Based on the TLSs gene set, TLS scores were calculated for each type of B cell. The TLS score in tumors was significantly higher than that in adjacent normal tissues, especially in MPR tumors. The highest TLS scores were observed in IFITM3+ memory B cells, HLA-DQA2+ memory B cells and RBFOX2+ memory B cells, indicating their predominance within TLSs [76]. Through spatial transcriptome analysis, the study further revealed the close correlation between TLS distribution and microenvironment status. The activation level of TLS was significantly reduced in low oxygen regions, while TLS exhibited higher maturity in invasion boundary regions with sufficient oxygen supply [82]. A study conducted a combined analysis of scRNA-seq and spatial transcriptomics on tumor tissue samples from 12 NSCLC patients before and after neoadjuvant immunotherapy combined with chemotherapy. Research found that treatment significantly enhanced the infiltration of immune cells such as T/NK cells and plasma cells and increased fibroblast activity. The formation of TLS was observed in patients with pathological remission, suggesting that enhanced immune response and spatial tissue remodeling were closely related to treatment efficacy [88]. These findings underscore the function of TLSs in LC.
2.1.4 Cancer associated fibroblast-related cellular neighborhoods
CAFs play multiple key functions in the TME, interacting with malignant cells, immune cells and other stromal cells through various mechanisms and have a significant impact on the occurrence, development, metastasis and therapy efficacy of tumors [89, 90]. Research found that CAFs promoted glycolysis in non-neuroendocrine (non-NE) small cell lung cancer cells (SCLC), thereby activating STING signaling in T cells and promoting chemokine expression. On the other hand, non-NE SCLC cells promoted the presence of antigen presenting CAFs, which might contribute to CD8+ T cell recruitment and Treg differentiation [91].
Through scRNA-seq and spatial transcriptome analysis of NSCLC tumor tissue, it was found that there was an increase in Th17 cells and inflammatory fibroblasts, and a decrease in Treg cells and stromal fibroblasts in effectively treated patients. SELENOP+ macrophages aggregated at tumor boundaries and lymphatic structures, synergistically recruiting T cells with antigen-presenting fibroblasts. After treatment, intercellular communication was enhanced, and cholesterol and other pathways were activated to promote immune response [88]. Another single-cell sequencing was performed on tumor samples of NSCLC patients before and after anti-PD-1 combined chemotherapy. Based on pathological evaluation, patients were divided into responders and non-responders. Research found that after combination therapy, the malignant epithelial cells in the tumor almost completely disappeared in responders, while non-responders had more monocytes/macrophages and dendritic cells, as well as a significantly increased proportion of COL11A1+ CAFs [82].
Additionally, scRNA-seq was performed on tumor, paired tumor adjacent, and normal samples from 16 NSCLC patients. It was found that POSTN+ CAFs and SPP1+macrophages had tight localization and might promote ECM remodeling and immunosuppressive TME formation [92]. Interestingly, the research team explored the spatial localization of fibroblasts relative to other cells in the lung TME and found that adjacent cells were cancer cells, macrophages, fibroblasts, and endothelial cells in order from near to far, indicating that fibroblasts and cancer cells could directly interact with each other [93]. These findings emphasize the controversial roles of CAFs and further analysis is required to figure out their contributions to LC progression.
2.2 Tumor cell-related cellular neighborhoods (Figure 3)
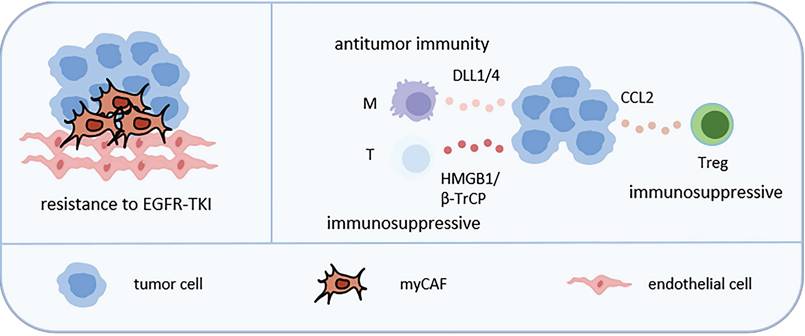
Tumor cells work together with neighboring cells to promote disease progression through interactions, signal transduction and physical contact. scRNA-seq and spatial transcriptome analysis revealed enrichment of CXCL14+ myofibroblasts CAF (myCAF) in advanced LUAD. This subgroup enhanced tumor invasiveness through epithelial to mesenchymal transition and angiogenesis and endowed LUAD cells with resistance to EGFR-TKI [94]. TKI treatment could lead to immunogenic cell death (ICD) and release of high mobility group box 1 protein (HMGB1) in LC cells. HMGB1 promoted macrophages to release cytokines and recruit T cells. At the same time, HMGB1 derived from tumor cells could also activate the NF-κB signaling pathway in T cells, leading to an increase in cytotoxic T lymphocyte-associated protein 4 (CTLA-4) expression, which weakened T cell function and created an immunosuppressive microenvironment [95].
Representative tumor cell-related CNs in LC. CAFs can interact with LC cells to promote their invasiveness and resistance to EGFR-TKI. Tumor cells can attract Treg by secreting CCL2, attract macrophages by secreting DLL1/4, attract T cells by secreting HMGB1 or β-TrCP, thereby reshaping TME. Abbreviations: myCAF, myofibroblasts cancer associated fibroblast; NE: neutrophil; M: macrophage; LUAD: lung adenocarcinoma; LUSC: lung squamous cell carcinoma; Treg, regulatory T cell.
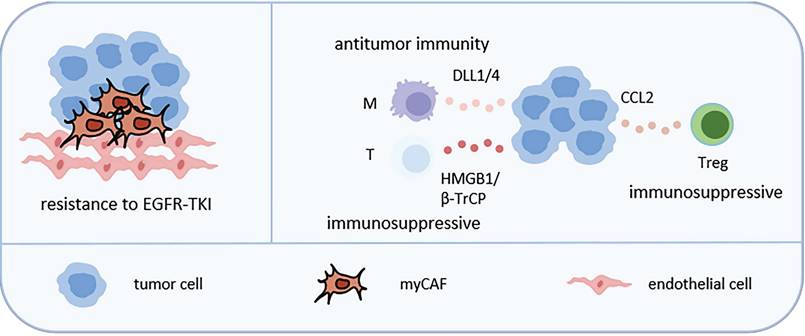
Other studies also showed the functions of some molecules. The high expression of JAG2 in NSCLC was negatively correlated with survival rate. Inhibiting the expression of JAG2 induced the expression of ligand DLL1/4 in NSCLC cells, activated the notch signaling pathway in macrophages, enhanced macrophage immune stimulatory function and triggered T-cell dependent anti-tumor immunity [96]. SOX2 promoted the secretion of C-C motif chemokine ligand 2 (CCL2) by NSCLC cells, leading to increased recruitment of Tregs and inhibition of CD8+ T cell infiltration [97]. NSCLC cells transferred extracellular vesicles containing β-TrCP to CD8+ T cells, promoting the ubiquitination of YAP1 protein and inhibiting the transcription and expression of downstream genes, leading to the inactivation of mTORC1 signaling pathway in CD8+ T cells and ultimately causing CD8+ T cell exhaustion [98].
Space transcriptome technology provides a new perspective for understanding. For example, the interaction between B cells and cancer cells was more frequent in LUAD, while the interaction between neutrophils, cancer cells, and macrophages was more common in LUSC. By analyzing the CNs, the research team found that the TME of LUAD and LUSC had different spatial structures. The tumor boundary (CN1) was the most abundant in LUAD, while the tumor compartment (CN7) was the most abundant in LUSC, reflecting the immunological rejection characteristics of LUSC. The CN enriched with macrophages (CN3) in LUAD was associated with poor survival rates, indicating that the spatial dynamics of macrophages may play an important role in influencing patient prognosis [99].
3. Key signaling pathways in shaping cellular neighborhoods (Figure 4)
3.1 Metabolites
The characteristics of TME are local competition for nutrients, accumulation of metabolic waste, and unfavorable pH values. The common demand for similar nutrients among various cells within the TME leads to potential competitive TME and immune suppression. And metabolic pathways fundamentally participate in the determination of cell fate and cellular programs [100, 101]. Glucose metabolism, lipid metabolism, and amino acid metabolism are the main components of tumor cell metabolism.
Representative key signaling pathways in shaping CNs in LC. Key signaling pathways-related CNs, such as metabolite, hypoxia and angiogenesis, are shown. Lactic acid promotes TAM infiltration and inhibits CD8+ T cell infiltration. Lipids promote TAM and Treg aggregation and inhibits NK cells aggregation. Tumor cells and TAMs compete to utilize glutamine, inhibiting the nutritional metabolism of T cells. These three metabolisms lead to the formation of an immunosuppressive microenvironment. Hypoxia promotes the enrichment of Treg and MDSC, and inhibits the enrichment of NK cells, T cells and DC by regulating the content of ROS, leading to immunosuppression. In addition, hypoxia also promotes the interaction between neutrophils, macrophages, LC cells and endothelial cells, promoting the invasiveness of tumors. Angiogenesis promotes infiltration of TAM, TAN and CAF, resulting tumor invasion. Abbreviations: TAM, tumor associated macrophage; CAF, cancer associated fibroblast; MDSC, bone marrow-derived suppressor cell; NE, neutrophil; NK, natural killer; DC, dendritic cell; TAN, tumor associated neutrophil.
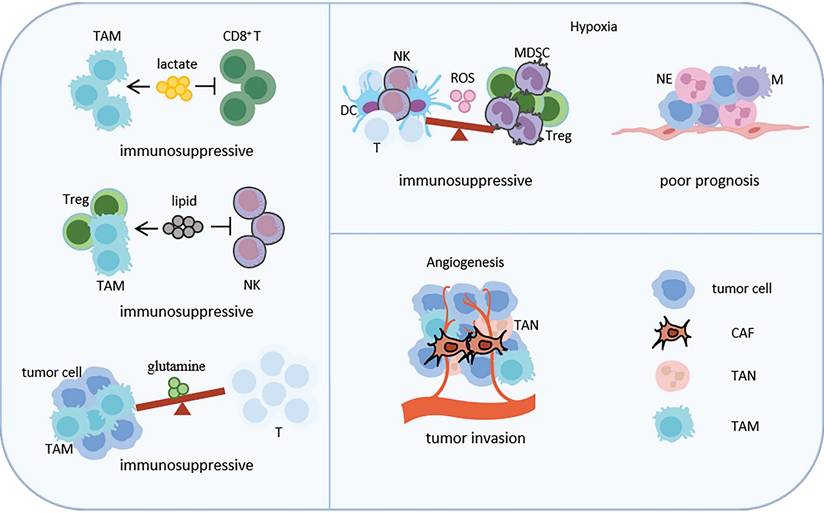
High expression of N-myc downstream-regulated gene 1 (NDRG1) in LUAD stabilized lactate dehydrogenase A (LDHA) by inhibiting its ubiquitination, thereby enhancing glycolysis and promoting lactate accumulation. This process promoted immune suppression by inducing M2 macrophage polarization and impairing the function of CD8+ T cells [102]. The inactivation of STK11 in LUAD affected the lactate transporter monocarboxylate transporter 4 (MCT4), leading to increased lactate production and efflux in the TME. Excessive lactate could polarize macrophages towards M2 type, thereby reducing T cell infiltration and inhibiting their function [103]. In addition, increased glycolytic activity in NSCLC patients after radiotherapy could lead to lactate accumulation and TME acidification [41].
Research also showed that TME could also provide lipids to cancer cells. The high lipid levels in TME altered the function of immune cells, with a large amount of free fatty acids leading to lipid accumulation in NK cells, thereby reducing their immune response. Immunosuppressed Tregs and TAMs also depend on fatty acid oxidation [104, 105]. LC cells secreted granulocyte-macrophage colony-stimulating factor (GM-CSF) to activate the PPAR-γ of TAM, promoting the release of fatty acids and cholesterol to support tumor metabolic needs [106]. Targeting lipid metabolism could inhibit the PI3K/AKT/mTOR signaling pathway, downregulating fatty acid synthesis and thereby inhibiting fatty acid oxidation in TAMs. On the other hand, it promoted the secretion of CXCL10 and CXCL11, thereby increasing the infiltration of CD8+T cells, ultimately inhibiting tumor progression [107].
In the immune microenvironment of LC, tumor cells compete with immune cells for the uptake of glutamine and glucose. Each cell type has a preference for the uptake of glucose and glutamine. Compared with T cells, LC cells are more likely to take up glutamine. Inhibiting TAM's uptake of glutamine can inhibit its transformation into M2-like macrophages [108].
3.2 Hypoxia
Hypoxia combined with high levels of ROS causes a chronic inflammatory state and suppresses the infiltration of immune cells such as CD8+ T cells, CD4+ T cells, NK cells and DCs. In addition, this condition simultaneously increases the aggregation of immunosuppressive cells, such as Tregs and MDSCs, ultimately leading to an immunosuppressive environment [109]. The activation process of HIF-1α has stage specific characteristics. Under mild hypoxia, HIF-1α exhibits partial transcriptional activity and enhances primary adaptive responses such as glycolysis and immunosuppression. As oxygen tension further decreases to moderate hypoxia levels, HIF-1α is fully activated, significantly inducing stronger responses such as angiogenesis [110]. Under severe hypoxia, fully activated HIF-1α continues to accumulate to high levels, causing a significant accumulation of lactate and extracellular acidification, ultimately triggering pathological outcomes such as programmed necrosis. Under acute hypoxia, neutrophils promote degranulation and glycogen storage, and exacerbate inflammatory damage. Macrophages promote IL-1β production and their migration ability. DCs secrete more pro-inflammatory cytokines. Under chronic hypoxia, tumor cells promote PD-L1 expression and inhibit the function of T cells. The upregulation of apoptotic gene expression and decreased migration ability in DCs lead to tumor immune escape. TAMs enhance glycolysis and lipid uptake, supporting their immunosuppressive function. Chronic hypoxia also weakens neutrophils function through DNA hypermethylation [111]. Hypoxia induces NSCLC cells to secrete extracellular vesicles, promoting macrophages M2 polarization, and resulting in NSCLC metastasis [112]. Hypoxia also promotes the production of HIF-1α in NSCLC TME, which can activate immunosuppressive genes and pro-tumor genes [113]. It can also recruit Tregs and MDSCs, creating an environment conducive to tumor growth [114].
Single-cell sequencing analysis was performed on NSCLC. The results showed that compared with low-grade tissue subtypes, the interaction between neutrophils, endothelial cells and cancer cells was significantly enhanced in high-grade tumors. It was also observed that the proportion of HIF-1α+ neutrophils was negatively correlated with patient survival prognosis [81]. The study also found that in the hypoxic areas of NSCLC, especially in the necrotic areas, a large number of macrophages with high HIF-1α expression were found, which promoted the transcription of the angiogenesis-related gene VEGF. HIF-1α+ TAMs promoted tumor angiogenesis, increased the density of tumor blood vessels and facilitated local invasion of tumors [115].
3.3 Angiogenesis
Tumor associated vasculature (TAV) has characteristics such as structural distortion, strong leakage and disrupted blood flow, causing hypoxia and acidosis in the TME. These abnormalities not only hinder the infiltration of cytotoxic T lymphocytes (CTLs), but also inhibit their survival and function, thereby promoting tumor metastasis and becoming the main barrier for immunotherapy [116].
Spatial transcriptome analysis showed that although VEGFA and VEGFB were found to be expressed in both LUAD and LUSC, their receptors were more common in LUAD, especially in fibroblasts [117]. Single-cell sequencing results showed significant spatial heterogeneity in LUAD. CD4+ primitive T cells in the central region of the tumor had a stronger ability to promote angiogenesis. In contrast, CD4+ primitive T cells in the peripheral region of the tumor had stronger cytokine production and pro-inflammatory ability. The signaling pathways that promoted angiogenesis, fatty acid synthesis and hypoxia were significantly enriched in macrophages in the central region of the tumor compared to macrophages in the peripheral region [118].
Another study identified subpopulations of tissue-resident neutrophils (TRNs) in NSCLC TME. The results showed that tumor associated neutrophils (TANs) and adjacent normal tissue associated neutrophils (NANs) had different expression profiles and neutrophils were the main source of VEGFA expression in the TME, confirming the important pro-angiogenic role of TRNs [119]. To explore potential intercellular communication, researchers conducted single-cell transcriptomic analysis to obtain intercellular signaling network. In the VEGF signaling network, as LUSC progressed, plasmacytoid DCs and monocyte-derived macrophages (mo-macs) replaced non-classical monocytes. In addition, mo-macs and plasmacytoid DCs played a dominant role in the TGF-β and CXCL signaling networks, which were associated with EMT, angiogenesis and immune suppression [120].
4. Conclusion
The rapid developments of single-cell and spatial transcriptome techniques have revealed the tumor heterogeneity and immune microenvironment dynamics of LC, elucidated the communication network of CNs in the LC microenvironment, significantly enhancing our understanding of the pathogenesis of LC, and providing an important basis for precise classification and treatment strategies of LC.
Although there have been many studies focused on conquering LC, it remains one of the malignant tumors with the highest mortality rate. Given the complexity of the TME, interactions between different types of cells jointly regulate tumor progression. A deeper understanding of CNs may provide new treatment options. With the advancements of single-cell and spatial transcriptome technologies, we can also better understand the molecular mechanisms underlying LC formation.
With the increasing variety and precision of detection technologies, an increasing number of tumor biomarkers are being detected. However, there is still a lack of reliable biomarkers to predict patient prognosis and treatment response. The spatial heterogeneity of tumors can lead to different clinical outcomes. By mining local cell interaction patterns and relative spatial positions of cells, CNs can stratify patient prognosis and improve the accuracy of predicting treatment efficiency. Although CNs provide us with a more cutting-edge perspective for a deeper understanding of LC, further in vitro experiments and animal models are still needed to explore specific molecular mechanisms. Understanding the biological and spatial characteristics of LC can provide us with a deeper understanding of the tumor ecosystem and ultimately offer new strategies for spatially targeted therapy.
Future research can focus on potential molecular mechanisms related to CNs formation and higher-level organizational structure formation. The successful progress of these studies is expected to elucidate the function and clinical significance of CNs, provide a more solid theoretical basis for identifying early LC, detecting LC-related biomarkers and evaluating LC prognosis.
Acknowledgements
This study was supported by National Natural Science Foundation of China (82503907, 82203285), Scientific Research Program of Hubei Provincial Department of Education (B2023238), Scientific and Technological Project of Henan Province (242102310323).
Consent for publication
Both authors read and approved the final manuscript and agree to be accountable for all aspects of the work.
Author contributions
Rui Bai collected and reviewed literatures and wrote the manuscript. Wenjie Sun designed and illustrated figures. Both authors read and approved the final manuscript.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Malvezzi M, Santucci C, Boffetta P, Collatuzzo G, Levi F, La Vecchia C. et al. European cancer mortality predictions for the year 2023 with focus on lung cancer. Ann Oncol. 2023;34:410-9
2. Siegel RL, Miller KD, Wagle NS, Jemal A. Cancer statistics, 2023. CA Cancer J Clin. 2023;73:17-48
3. Liu Z, Yang Z, Wu J, Zhang W, Sun Y, Zhang C. et al. A single-cell atlas reveals immune heterogeneity in anti-PD-1-treated non-small cell lung cancer. Cell. 2025;188:3081-96.e19
4. Schürch CM, Bhate SS, Barlow GL, Phillips DJ, Noti L, Zlobec I. et al. Coordinated Cellular Neighborhoods Orchestrate Antitumoral Immunity at the Colorectal Cancer Invasive Front. Cell. 2020;182:1341-59.e19
5. Monkman J, Moradi A, Yunis J, Ivison G, Mayer A, Ladwa R. et al. Spatial insights into immunotherapy response in non-small cell lung cancer (NSCLC) by multiplexed tissue imaging. J Transl Med. 2024;22:239
6. Farah EN, Hu RK, Kern C, Zhang Q, Lu TY, Ma Q. et al. Spatially organized cellular communities form the developing human heart. Nature. 2024;627:854-64
7. Seidman JS, Troutman TD, Sakai M, Gola A, Spann NJ, Bennett H. et al. Niche-Specific Reprogramming of Epigenetic Landscapes Drives Myeloid Cell Diversity in Nonalcoholic Steatohepatitis. Immunity. 2020;52:1057-74.e7
8. Armingol E, Officer A, Harismendy O, Lewis NE. Deciphering cell-cell interactions and communication from gene expression. Nat Rev Genet. 2021;22:71-88
9. Arnol D, Schapiro D, Bodenmiller B, Saez-Rodriguez J, Stegle O. Modeling Cell-Cell Interactions from Spatial Molecular Data with Spatial Variance Component Analysis. Cell Rep. 2019;29:202-11.e6
10. Almet AA, Cang Z, Jin S, Nie Q. The landscape of cell-cell communication through single-cell transcriptomics. Curr Opin Syst Biol. 2021;26:12-23
11. Dominiak A, Chełstowska B, Olejarz W, Nowicka G. Communication in the Cancer Microenvironment as a Target for Therapeutic Interventions. Cancers (Basel). 2020 12
12. Bejarano L, Jordāo MJC, Joyce JA. Therapeutic Targeting of the Tumor Microenvironment. Cancer Discov. 2021;11:933-59
13. Wang Q, Gümüş ZH, Colarossi C, Memeo L, Wang X, Kong CY. et al. SCLC: Epidemiology, Risk Factors, Genetic Susceptibility, Molecular Pathology, Screening, and Early Detection. J Thorac Oncol. 2023;18:31-46
14. Zhang P, Chen PL, Li ZH, Zhang A, Zhang XR, Zhang YJ. et al. Association of smoking and polygenic risk with the incidence of lung cancer: a prospective cohort study. Br J Cancer. 2022;126:1637-46
15. WHO Guidelines Approved by the Guidelines Review Committee. WHO global air quality guidelines: Particulate matter (PM(25) and PM(10)), ozone, nitrogen dioxide, sulfur dioxide and carbon monoxide. Geneva: World Health Organization © World Health Organization 2021. 2021
16. Jarosz-Biej M, Smolarczyk R, Cichoń T, Kułach N. Tumor Microenvironment as A “Game Changer” in Cancer Radiotherapy. Int J Mol Sci. 2019 20
17. Wick MR. Pathologic features of smoking-related lung diseases, with emphasis on smoking-related interstitial fibrosis and a consideration of differential diagnoses. Semin Diagn Pathol. 2018;35:315-23
18. Arai K, Nishito Y, Mizuno H, Motoi N, Hiraoka N, Fuse M. et al. EML4-ALK rearrangement creates a distinctive myeloid cell-dominant immunosuppressive microenvironment in lung cancer. Cancer Immunol Res. 2025
19. Ding W, Mo J, Su Y, Zhang Q, Sun D, Yao X. et al. Metabolic reprogramming of tumor-associated macrophages via adenosine-A(2A)R signaling drives cross-resistance in non-small cell lung cancer. Drug Resist Updat. 2025;82:101272
20. Naoi Y, Inukai Y, Izumikawa T, Nagasaki J, Ishino T, Ueda Y. et al. Close Spatial Interactions between Cancer Cells and Cancer-Associated Fibroblasts Suppress Antitumor Immunity. Cancer Immunol Res. 2025
21. Che M, Wei W, Yang X, Liang J, Li Y, Ye Y. et al. Radiation-induced upregulation of itaconate in macrophages promotes the radioresistance of non-small cell lung cancer by stabilizing NRF2 protein and suppressing immune response. Redox Biol. 2025;85:103711
22. Runtsch MC, Angiari S, Kargl J. Targeting myeloid cell immunometabolism to improve current non-small cell lung cancer therapies. Pharmacol Ther. 2025;273:108893
23. Cornier MA, Dabelea D, Hernandez TL, Lindstrom RC, Steig AJ, Stob NR. et al. The metabolic syndrome. Endocr Rev. 2008;29:777-822
24. Dobrowolski P, Prejbisz A, Kuryłowicz A, Baska A, Burchardt P, Chlebus K. et al. Metabolic syndrome - a new definition and management guidelines: A joint position paper by the Polish Society of Hypertension, Polish Society for the Treatment of Obesity, Polish Lipid Association, Polish Association for Study of Liver, Polish Society of Family Medicine, Polish Society of Lifestyle Medicine, Division of Prevention and Epidemiology Polish Cardiac Society, “Club 30” Polish Cardiac Society, and Division of Metabolic and Bariatric Surgery Society of Polish Surgeons. Arch Med Sci. 2022;18:1133-56
25. Russo A, Autelitano M, Bisanti L. Metabolic syndrome and cancer risk. European journal of cancer (Oxford, England: 1990). 2008;44:293-7
26. Esposito K, Chiodini P, Colao A, Lenzi A, Giugliano D. Metabolic syndrome and risk of cancer: a systematic review and meta-analysis. Diabetes Care. 2012;35:2402-11
27. Viola A, Munari F, Sánchez-Rodríguez R, Scolaro T, Castegna A. The Metabolic Signature of Macrophage Responses. Front Immunol. 2019;10:1462
28. Noy R, Pollard JW. Tumor-associated macrophages: from mechanisms to therapy. Immunity. 2014;41:49-61
29. DeNardo DG, Ruffell B. Macrophages as regulators of tumour immunity and immunotherapy. Nat Rev Immunol. 2019;19:369-82
30. Cheng C, Wang P, Yang Y, Du X, Xia H, Liu J. et al. Smoking-Induced M2-TAMs, via circEML4 in EVs, Promote the Progression of NSCLC through ALKBH5-Regulated m6A Modification of SOCS2 in NSCLC Cells. Adv Sci (Weinh). 2023;10:e2300953
31. Luo W, Zeng Z, Jin Y, Yang L, Fan T, Wang Z. et al. Distinct immune microenvironment of lung adenocarcinoma in never-smokers from smokers. Cell Rep Med. 2023;4:101078
32. Krishna C, Tervi A, Saffern M, Wilson EA, Yoo SK, Mars N. et al. An immunogenetic basis for lung cancer risk. Science. 2024;383:eadi3808
33. Long E, Yin J, Shin JH, Li Y, Li B, Kane A. et al. Context-aware single-cell multiomics approach identifies cell-type-specific lung cancer susceptibility genes. Nat Commun. 2024;15:7995
34. Gómez-López S, Alhendi ASN, Przybilla MJ, Bordeu I, Whiteman ZE, Butler T. et al. Aberrant basal cell clonal dynamics shape early lung carcinogenesis. Science. 2025;388:eads9145
35. Ge X, Song C, Zhu C, Ding Y, Liu M, Yang J, et al. Association of ambient fine particulate matter (PM(2.5)) and its constituents with risk of pulmonary nodules in a lung cancer screening project. Environ Res. 2025: 122409
36. Chen S, Ye P, Wei J, Cheng H, Zhang Y, Ma Y. et al. Causal links between long-term exposure to the chemical constituents of PM(2.5) and lung cancer mortality in southern China. J Hazard Mater. 2025;495:139096
37. DIESEL AND GASOLINE ENGINE EXHAUSTS AND SOME NITROARENES. IARC MONOGRAPHS ON THE EVALUATION OF CARCINOGENIC RISKS TO HUMANS. IARC Monogr Eval Carcinog Risks Hum. 2014;105:9-699
38. Outdoor Air Pollution. IARC Monogr Eval Carcinog Risks Hum. 2016; 109: 9-444.
39. Díaz-Gay M, Zhang T, Hoang PH, Leduc C, Baine MK, Travis WD. et al. The mutagenic forces shaping the genomes of lung cancer in never smokers. Nature. 2025
40. Hill W, Lim EL, Weeden CE, Lee C, Augustine M, Chen K. et al. Lung adenocarcinoma promotion by air pollutants. Nature. 2023;616:159-67
41. Mittal A, Nenwani M, Sarangi I, Achreja A, Lawrence TS, Nagrath D. Radiotherapy-induced metabolic hallmarks in the tumor microenvironment. Trends Cancer. 2022;8:855-69
42. Yamaga S, Aziz M, Murao A, Brenner M, Wang P. DAMPs and radiation injury. Front Immunol. 2024;15:1353990
43. Wei J, Montalvo-Ortiz W, Yu L, Krasco A, Ebstein S, Cortez C. et al. Sequence of αPD-1 relative to local tumor irradiation determines the induction of abscopal antitumor immune responses. Sci Immunol. 2021 6
44. Hellevik T, Berzaghi R, Lode K, Islam A, Martinez-Zubiaurre I. Immunobiology of cancer-associated fibroblasts in the context of radiotherapy. J Transl Med. 2021;19:437
45. Zhai D, An D, Wan C, Yang K. Radiotherapy: Brightness and darkness in the era of immunotherapy. Transl Oncol. 2022;19:101366
46. Wang Q, Li S, Qiao S, Zheng Z, Duan X, Zhu X. Changes in T Lymphocyte Subsets in Different Tumors Before and After Radiotherapy: A Meta-analysis. Front Immunol. 2021;12:648652
47. Shirasawa M, Yoshida T, Matsumoto Y, Shinno Y, Okuma Y, Goto Y. et al. Impact of chemoradiotherapy on the immune-related tumour microenvironment and efficacy of anti-PD-(L)1 therapy for recurrences after chemoradiotherapy in patients with unresectable locally advanced non-small cell lung cancer. European journal of cancer (Oxford, England: 1990). 2020;140:28-36
48. Han L, Shi H, Luo Y, Sun W, Li S, Zhang N. et al. Gene signature based on B cell predicts clinical outcome of radiotherapy and immunotherapy for patients with lung adenocarcinoma. Cancer Med. 2020;9:9581-94
49. Van Dieren L, Quisenaerts T, Licata M, Beddok A, Lellouch AG, Ysebaert D. et al. Combined Radiotherapy and Hyperthermia: A Systematic Review of Immunological Synergies for Amplifying Radiation-Induced Abscopal Effects. Cancers (Basel). 2024 16
50. Shi Z, Hu C, Zheng X, Sun C, Li Q. Feedback loop between hypoxia and energy metabolic reprogramming aggravates the radioresistance of cancer cells. Exp Hematol Oncol. 2024;13:55
51. Liu L, Yu J, Liu Y, Xie L, Hu F, Liu H. Hypoxia-driven angiogenesis and metabolic reprogramming in vascular tumors. Front Cell Dev Biol. 2025;13:1572909
52. Zhang Y, Hu J, Ji K, Jiang S, Dong Y, Sun L. et al. CD39 inhibition and VISTA blockade may overcome radiotherapy resistance by targeting exhausted CD8+ T cells and immunosuppressive myeloid cells. Cell Rep Med. 2023;4:101151
53. Raaijmakers K, Adema GJ, Bussink J, Ansems M. Cancer-associated fibroblasts, tumor and radiotherapy: interactions in the tumor micro-environment. J Exp Clin Cancer Res. 2024;43:323
54. Zhi Y, Chen H, Huang Y, Li L, Chen Z, Huang H. et al. Novel NLRP3 inhibitors mitigate acute radiation-induced lung injury by suppressing pyroptosis in alveolar epithelial cells. Toxicol Appl Pharmacol. 2025;502:117458
55. Xiao Z, Puré E. The fibroinflammatory response in cancer. Nat Rev Cancer. 2025;25:399-425
56. Drakopanagiotakis F, Krauss E, Michailidou I, Drosos V, Anevlavis S, Günther A. et al. Lung Cancer and Interstitial Lung Diseases. Cancers (Basel). 2024 16
57. Guo JL, Griffin M, Yoon JK, Lopez DM, Zhu Y, Lu JM. et al. Histological signatures map anti-fibrotic factors in mouse and human lungs. Nature. 2025;641:993-1004
58. Franzén L, Olsson Lindvall M, Hühn M, Ptasinski V, Setyo L, Keith BP. et al. Mapping spatially resolved transcriptomes in human and mouse pulmonary fibrosis. Nat Genet. 2024;56:1725-36
59. Mayr CH, Santacruz D, Jarosch S, Bleck M, Dalton J, McNabola A. et al. Spatial transcriptomic characterization of pathologic niches in IPF. Sci Adv. 2024;10:eadl5473
60. Vannan A, Lyu R, Williams AL, Negretti NM, Mee ED, Hirsh J. et al. Spatial transcriptomics identifies molecular niche dysregulation associated with distal lung remodeling in pulmonary fibrosis. Nat Genet. 2025;57:647-58
61. Baslan T, Hicks J. Unravelling biology and shifting paradigms in cancer with single-cell sequencing. Nat Rev Cancer. 2017;17:557-69
62. Goveia J, Rohlenova K, Taverna F, Treps L, Conradi LC, Pircher A. et al. An Integrated Gene Expression Landscape Profiling Approach to Identify Lung Tumor Endothelial Cell Heterogeneity and Angiogenic Candidates. Cancer Cell. 2020;37:21-36.e13
63. Leader AM, Grout JA, Maier BB, Nabet BY, Park MD, Tabachnikova A. et al. Single-cell analysis of human non-small cell lung cancer lesions refines tumor classification and patient stratification. Cancer Cell. 2021;39:1594-609.e12
64. Marjanovic ND, Hofree M, Chan JE, Canner D, Wu K, Trakala M. et al. Emergence of a High-Plasticity Cell State during Lung Cancer Evolution. Cancer Cell. 2020;38:229-46.e13
65. Liu B, Hu X, Feng K, Gao R, Xue Z, Zhang S. et al. Temporal single-cell tracing reveals clonal revival and expansion of precursor exhausted T cells during anti-PD-1 therapy in lung cancer. Nat Cancer. 2022;3:108-21
66. Yang J, Davidoff AM, Murphy AJ. From preneoplastic lesion to heterogenous tumor: recent insights into hepatoblastoma biology and therapeutic opportunities. Mol Cancer. 2025;24:198
67. Maddipati R. Metastatic heterogeneity in pancreatic cancer: mechanisms and opportunities for targeted intervention. J Clin Invest. 2025 135
68. Cheng X, Peng T, Chu T, Yang Y, Liu J, Gao Q. et al. Application of single-cell and spatial omics in deciphering cellular hallmarks of cancer drug response and resistance. J Hematol Oncol. 2025;18:70
69. Dakal TC, Bhushan R, Xu C, Gadi BR, Cameotra SS, Yadav V. et al. Intricate relationship between cancer stemness, metastasis, and drug resistance. MedComm (2020). 2024;5:e710
70. Chen Y, Wang D, Li Y, Qi L, Si W, Bo Y. et al. Spatiotemporal single-cell analysis decodes cellular dynamics underlying different responses to immunotherapy in colorectal cancer. Cancer Cell. 2024;42:1268-85.e7
71. St Paul M, Ohashi PS. The Roles of CD8(+) T Cell Subsets in Antitumor Immunity. Trends Cell Biol. 2020;30:695-704
72. Qu J, Sun W, Li Y, Cai Z, Wu B, Shen Q. et al. Immune profiling identifies CD161(+)CD127(+)CD8(+) T cells as a predictive biomarker for Anti-PD-L1 therapy response in the SCLC-I subtype. J Transl Med. 2025;23:711
73. Lopez de Rodas M, Villalba-Esparza M, Sanmamed MF, Chen L, Rimm DL, Schalper KA. Biological and clinical significance of tumour-infiltrating lymphocytes in the era of immunotherapy: a multidimensional approach. Nat Rev Clin Oncol. 2025;22:163-81
74. Srivastava S, Furlan SN, Jaeger-Ruckstuhl CA, Sarvothama M, Berger C, Smythe KS. et al. Immunogenic Chemotherapy Enhances Recruitment of CAR-T Cells to Lung Tumors and Improves Antitumor Efficacy when Combined with Checkpoint Blockade. Cancer Cell. 2021;39:193-208.e10
75. Kratzmeier C, Taheri M, Mei Z, Lim I, Khalil MA, Carter-Cooper B. et al. Lung adenocarcinoma-derived IFN-γ promotes growth by modulating CD8+ T cell production of CCR5 chemokines. J Clin Invest. 2025
76. Fu R, Zhang C, Song MM, Gao X, Li F, Cai M. et al. A single-cell map of patients with non-small cell lung cancer harboring rare-driver mutations after anti-PD-1 treatment. Cancer Lett. 2025;616:217595
77. Huang Z, Xiao Z, Yu L, Liu J, Yang Y, Ouyang W. Tumor-associated macrophages in non-small-cell lung cancer: From treatment resistance mechanisms to therapeutic targets. Crit Rev Oncol Hematol. 2024;196:104284
78. Chi Y, Jiang H, Yin Y, Zhou X, Shao Y, Li Y. et al. Macrophage Signaling Pathways in Health and Disease: From Bench to Bedside Applications. MedComm (2020). 2025;6:e70256
79. Li Y, You J, Zou Z, Sun G, Shi Y, Sun Y. et al. Decoding the Tumor Microenvironment: Exosome-Mediated Macrophage Polarization and Therapeutic Frontiers. Int J Biol Sci. 2025;21:4187-214
80. Yang Y, Li S, To KKW, Zhu S, Wang F, Fu L. Tumor-associated macrophages remodel the suppressive tumor immune microenvironment and targeted therapy for immunotherapy. J Exp Clin Cancer Res. 2025;44:145
81. Sorin M, Rezanejad M, Karimi E, Fiset B, Desharnais L, Perus LJM. et al. Single-cell spatial landscapes of the lung tumour immune microenvironment. Nature. 2023;614:548-54
82. Yan Y, Sun D, Hu J, Chen Y, Sun L, Yu H. et al. Multi-omic profiling highlights factors associated with resistance to immuno-chemotherapy in non-small-cell lung cancer. Nat Genet. 2025;57:126-39
83. Yang L, Li A, Yu W, Wang H, Zhang L, Wang D. et al. Blockade of purine metabolism reverses macrophage immunosuppression and enhances anti-tumor immunity in non-small cell lung cancer. Drug Resist Updat. 2025;78:101175
84. Fan Z, Yi Z, Li S, He J. Parabacteroides distasonis promotes CXCL9 secretion of tumor-associated macrophages and enhances CD8(+)T cell activity to trigger anti-tumor immunity against anti-PD-1 treatment in non-small cell lung cancer mice. BMC Biotechnol. 2025;25:30
85. Sautès-Fridman C, Petitprez F, Calderaro J, Fridman WH. Tertiary lymphoid structures in the era of cancer immunotherapy. Nat Rev Cancer. 2019;19:307-25
86. Petitprez F, de Reyniès A, Keung EZ, Chen TW, Sun CM, Calderaro J. et al. B cells are associated with survival and immunotherapy response in sarcoma. Nature. 2020;577:556-60
87. Vanhersecke L, Brunet M, Guégan JP, Rey C, Bougouin A, Cousin S. et al. Mature tertiary lymphoid structures predict immune checkpoint inhibitor efficacy in solid tumors independently of PD-L1 expression. Nat Cancer. 2021;2:794-802
88. Cui X, Liu S, Song H, Xu J, Sun Y. Single-cell and spatial transcriptomic analyses revealing tumor microenvironment remodeling after neoadjuvant chemoimmunotherapy in non-small cell lung cancer. Mol Cancer. 2025;24:111
89. Masuda H. Cancer-associated fibroblasts in cancer drug resistance and cancer progression: a review. Cell Death Discov. 2025;11:341
90. Li Y, Liu Q, Jing X, Wang Y, Jia X, Yang X. et al. Cancer-Associated Fibroblasts: Heterogeneity, Cancer Pathogenesis, and Therapeutic Targets. MedComm (2020). 2025;6:e70292
91. Lu Y, Li H, Zhao P, Wang X, Shao W, Liu Y. et al. Crosstalk between cancer-associated fibroblasts and non-neuroendocrine tumor cells in small cell lung cancer involves in glycolysis and antigen-presenting features. Mol Med. 2024;30:274
92. Chen C, Guo Q, Liu Y, Hou Q, Liao M, Guo Y. et al. Single-cell and spatial transcriptomics reveal POSTN(+) cancer-associated fibroblasts correlated with immune suppression and tumour progression in non-small cell lung cancer. Clin Transl Med. 2023;13:e1515
93. Weigert A, Zheng X, Nenzel A, Turkowski K, Günther S, Strack E. et al. Fibrocytes boost tumor-supportive phenotypic switches in the lung cancer niche via the endothelin system. Nat Commun. 2022;13:6078
94. Xu W, Yang H, Xu K, Zhu A, Hall SRR, Jia Y. et al. Transitional CXCL14(+) cancer-associated fibroblasts enhance tumour metastasis and confer resistance to EGFR-TKIs, revealing therapeutic vulnerability to filgotinib in lung adenocarcinoma. Clin Transl Med. 2025;15:e70281
95. Fu M, Zhao J, Zhang L, Sheng Z, Li X, Qiu F. et al. Overcoming tyrosine kinase inhibitor resistance in lung cancer brain metastasis with CTLA4 blockade. Cancer Cell. 2024;42:1882-97.e7
96. Mandula JK, Sierra-Mondragon RA, Jimenez RV, Chang D, Mohamed E, Chang S. et al. Jagged2 targeting in lung cancer activates anti-tumor immunity via Notch-induced functional reprogramming of tumor-associated macrophages. Immunity. 2024;57:1124-40.e9
97. Torres-Mejia E, Weng S, Whittaker CA, Nguyen KB, Duong E, Yim L. et al. Lung Cancer-Intrinsic SOX2 Expression Mediates Resistance to Checkpoint Blockade Therapy by Inducing Treg-Dependent CD8+ T-cell Exclusion. Cancer Immunol Res. 2025;13:496-516
98. Li G, Wen Z, Xiong S. Microenvironmental β-TrCP negates amino acid transport to trigger CD8(+) T cell exhaustion in human non-small cell lung cancer. Cell Rep. 2025;44:115128
99. Desharnais L, Sorin M, Rezanejad M, Liu B, Karimi E, Atallah A. et al. Spatially mapping the tumour immune microenvironments of non-small cell lung cancer. Nat Commun. 2025;16:1345
100. Kay EJ, Zanivan S. The tumor microenvironment is an ecosystem sustained by metabolic interactions. Cell Rep. 2025;44:115432
101. Mao Y, Xia Z, Xia W, Jiang P. Metabolic reprogramming, sensing, and cancer therapy. Cell Rep. 2024;43:115064
102. Wu G, Cheng H, Yin J, Zheng Y, Shi H, Pan B. et al. NDRG1-Driven Lactate Accumulation Promotes Lung Adenocarcinoma Progression Through the Induction of an Immunosuppressive Microenvironment. Adv Sci (Weinh). 2025: e01238.
103. Qian Y, Galan-Cobo A, Guijarro I, Dang M, Molkentine D, Poteete A. et al. MCT4-dependent lactate secretion suppresses antitumor immunity in LKB1-deficient lung adenocarcinoma. Cancer Cell. 2023;41:1363-80.e7
104. Yu W, Lei Q, Yang L, Qin G, Liu S, Wang D. et al. Contradictory roles of lipid metabolism in immune response within the tumor microenvironment. J Hematol Oncol. 2021;14:187
105. Prendeville H, Lynch L. Diet, lipids, and antitumor immunity. Cell Mol Immunol. 2022;19:432-44
106. Kuhlmann-Hogan A, Cordes T, Xu Z, Kuna RS, Traina KA, Robles-Oteíza C. et al. EGFR-Driven Lung Adenocarcinomas Co-opt Alveolar Macrophage Metabolism and Function to Support EGFR Signaling and Growth. Cancer Discov. 2024: Of1-of22.
107. Chen Y, Zhou Y, Ren R, Chen Y, Lei J, Li Y. Harnessing lipid metabolism modulation for improved immunotherapy outcomes in lung adenocarcinoma. J Immunother Cancer. 2024 12
108. Li J, Xiang L, Wang S, Zhang Y, Zhao S, Zhang D. et al. Targeting the crosstalk between glutamine metabolism and tumor immune microenvironment for lung cancer immunotherapy. Interdisciplinary Medicine. 2025 3
109. Chen Z, Han F, Du Y, Shi H, Zhou W. Hypoxic microenvironment in cancer: molecular mechanisms and therapeutic interventions. Signal Transduct Target Ther. 2023;8:70
110. Wang P, Zhang XP, Liu F, Wang W. Progressive Deactivation of Hydroxylases Controls Hypoxia-Inducible Factor-1α-Coordinated Cellular Adaptation to Graded Hypoxia. Research (Wash D C). 2025;8:0651
111. Low JT, Kim Y, Matsushita M, Ho PC. Role of oxygen sensing and hypoxia-inducible factors in orchestrating innate immune responses. Nat Immunol. 2025;26:2138-47
112. Wang D, Wang S, Jin M, Zuo Y, Wang J, Niu Y. et al. Hypoxic Exosomal circPLEKHM1-Mediated Crosstalk between Tumor Cells and Macrophages Drives Lung Cancer Metastasis. Adv Sci (Weinh). 2024;11:e2309857
113. Salem A, Asselin MC, Reymen B, Jackson A, Lambin P, West CML. et al. Targeting Hypoxia to Improve Non-Small Cell Lung Cancer Outcome. J Natl Cancer Inst. 2018 110
114. Robles-Oteíza C, Hastings K, Choi J, Sirois I, Ravi A, Expósito F. et al. Hypoxia is linked to acquired resistance to immune checkpoint inhibitors in lung cancer. J Exp Med. 2025 222
115. Yang L, Zhang Y. Tumor-associated macrophages: from basic research to clinical application. J Hematol Oncol. 2017;10:58
116. Lanitis E, Irving M, Coukos G. Tumour-associated vasculature in T cell homing and immunity: opportunities for cancer therapy. Nat Rev Immunol. 2025
117. De Zuani M, Xue H, Park JS, Dentro SC, Seferbekova Z, Tessier J. et al. Single-cell and spatial transcriptomics analysis of non-small cell lung cancer. Nat Commun. 2024;15:4388
118. Wang Y, Li X, Peng S, Hu H, Wang Y, Shao M. et al. Single-Cell Analysis Reveals Spatial Heterogeneity of Immune Cells in Lung Adenocarcinoma. Front Cell Dev Biol. 2021;9:638374
119. Salcher S, Sturm G, Horvath L, Untergasser G, Kuempers C, Fotakis G. et al. High-resolution single-cell atlas reveals diversity and plasticity of tissue-resident neutrophils in non-small cell lung cancer. Cancer Cell. 2022;40:1503-20.e8
120. Luo B, Luo Y, Chen S, Yang T, Liu B, Liao X. et al. Real-Time Evolutionary Landscape of the Bronchial Epithelium and Corresponding Dynamic Immune Cell Alterations in Lung Squamous Cell Carcinogenesis. Adv Sci (Weinh). 2025: e13256.
Author contact
Corresponding author: Wenjie Sun (Email: wenjie1026edu.cn).

 Global reach, higher impact
Global reach, higher impact