Impact Factor ISSN: 1837-9664
J Cancer 2012; 3:58-66. doi:10.7150/jca.3872 This volume Cite
Case Report
Molecular Characterization of a Patient's Small Cell Carcinoma of the Ovary of the Hypercalcemic Type
1. Clinical Translational Research Division, Translational Genomics Research Institute, Phoenix, AZ, USA.
2. TGen Clinical Research Service, Translational Genomics Research Institute, Scottsdale, AZ, USA.
3. Scottsdale Healthcare, Scottsdale, AZ, USA.
4. Integrated Cancer Genomics Division, Translational Genomics Research Institute, Phoenix, AZ, USA.
Received 2011-11-28; Accepted 2012-1-10; Published 2012-1-26
Abstract
Small cell carcinoma of the ovary of the hypercalcemic type (SCCOHT) is a very rare tumor type that mainly affects young women. We report a 21-year old woman with SCCOHT. The patient initially presented with stage T3AN1MX disease and treated with surgery. The patient then received 8 cycles of multi-agent chemotherapy including cisplatin, bleomycin, cyclophosphamide, doxorubicin, and etoposide. Upon relapse, the patient underwent total abdominal hysterectomy, followed by chemotherapy with gemcitabine. The patient subsequently received radiation therapy and chemotherapy with bevacizumab, irinotecan and docetaxel. She passed away approximately 5 months after the second surgery and with her prior permission an immediate autopsy was performed. We examined the gene expression and copy number profiles of the tumor tissue samples obtained from the autopsy and compared them to normal ovary tissues. Our results indicated that although this tumor did not harbor chromosomal abnormalities nor gene copy number changes, there were significant gene expression changes in a number of genes/pathways. More than 5,000 genes showed significant differential expression in the tumor when compared to normal ovary tissue. Pathway enrichment analysis further identified several pathways/processes including the Vitamin D receptor signaling and the hedgehog signaling pathways to be significantly dysregulated. The gene expression profiling also suggests a number of agents such as pazopanib, bortezomib, 5-azacytidine, and PARP inhibitors as treatment options to possibly explore in future trials against this disease.
Keywords: Small cell carcinoma, hypercalcemic type, ovarian carcinoma
Introduction
Small cell carcinoma of the ovary of the hypercalcemic type (SCCOHT) is a rare highly aggressive neoplasm that almost exclusively affects young women. The tumor was first reported in 1979 by R. E. Scully as a distinct small cell ovarian tumor with unknown derivation [1]. Several cases have since been reported in the literature with the largest series consisting of 150 patients where it was observed that the mean age at diagnosis is 24 years and that approximately two thirds of patients present with hypercalcemia. [2]
Although SCCOHT is rare, it is the most common ovarian neoplasm of the many potential neoplasms composed of small round cells in women under 40. [3, 4] As it can be hard to distinguish between the types of uncommon small cell ovarian neoplasms, several reports and reviews describing the distinguishing clinical and histopathological features of these tumors have been written. [3-6]
Despite the number of treatment modalities investigated, longer term survival with SCCOHT still occurs infrequently, as any response to these treatments appears to be short lived. [7-11] A VPCBAE (vinblastine, cisplatin, cyclophosphamide, bleomycin, adriamycin, and etoposide) regimen appears to be the preferred treatment in most centers despite its high toxicity. [8, 12, 13] Clearly, better therapeutic options are needed to fight this disease.
There are an increasing number of examples of tumors where molecular profiling has been able to identify a context of vulnerability that has allowed a therapeutic chance where not too long ago they would have been considered not very treatable or untreatable. The most commonly cited example is the use of imatinib in targeting BCR-ABL in Chronic myelogenous leukemia. Another, more recent, example is the targeting of aberrant hedgehog signaling in basal cell carcinoma [14, 15]. To our knowledge, little work has been done to uncover the molecular determinants of SCCOHT.
Herein we present a study of a patient's SCCOHT. The possibility that a better molecular understanding of SCCOHT might lead to less toxic and more effective therapies, drove us to molecularly profile this tumor and make this information available to other translational investigators who have this same goal.
Materials and Methods
Human tissues
With the patient's permission, fresh frozen small cell ovarian tumor tissue was obtained via an autopsy performed immediately after death. A portion of the tumor tissue was fixed with formalin and embedded in paraffin. Standard hematoxylin and eosin (H&E) staining was performed on the paraffin embedded tissue section using a BondMax autostainer (Leica Microsystems, Buffalo Grove, IL). Normal ovarian RNA was obtained through BioChain Institute, Inc (Hayward, CA).
RNA Isolation
Total RNA from the tumor tissues was extracted using TRIzol (Invitrogen, Carlsbad, CA), followed by isolation using RNeasy mini kit (Qiagen, Valencia, CA). Total RNA from cell pellets was isolated using the NucleoSpin® RNA II isolation kit (BD Biosciences, Palo Alta, CA). The RNA samples were then used in the microarray experiments.
Oligonucleotide Microarrays and Analysis
Total RNA (500 ng) was amplified and labeled with Cy3 or Cy5 dye using an Agilent (Palo Alto, CA) fluorescent linear amplification kit. Labeled amplified RNAs were fragmented and hybridized on Agilent Human 1A(V2) Oligo Microarray slides using an Agilent in situ hybridization kit. Slides were scanned using an Agilent G2505B scanner, and Agilent feature extraction software (v8.1) was used to calculate normalized signal intensity. Following feature extraction, files were opened up in excel and sorting was performed as follows: median normalized intensity values for each probe were calculated, intensity values less than 0.25 (median = 1) were changed to 0.25 to prevent transcripts expressed at low levels as being identified as differentially regulated, expression ratios were calculated by dividing the signal intensity of the tumor by the signal intensity of obtained from the normal ovarian RNA. Array analysis comparing the SCCOHT tumor to normal ovarian RNA was done in duplicates with dye-swab (For the first array RNA from the normal ovary was labeled with Cy3 and RNA from the tumor tissue was labeled with Cy5; for the second array RNA from the tumor tissue was labeled Cy3 and RNA from the normal ovary was labeled Cy5). Expression ratios from the 2 arrays were averaged for a final expression ratio.
Pathway Enrichment Analysis
To identify regulatory pathways/networks that differentiate the tumor from the normal ovary, we analyzed the differentially expressed genes between the tumor and normal ovary samples for relative enrichment of certain categories from several functional ontologies in MetaCore™ (Thomson Reuters v. 6.8), including GO (gene ontology), network processes, canonical pathway maps, and disease networks. The MetaCore™ database is a commercially available resource containing over 200,000 protein-protein and protein-small molecule interactions manually extracted from the literature by a group of experts [16]. To rank the results we calculated the p-values of each identified pathway/network using formula described by Nikolsky and colleagues [17]. The p-value is essentially the probability of a particular mapping arising by chance given the number of genes in the set relative to all genes on maps/processes, genes on a particular map/processes, and genes in the analyzed experiment [17]. A pathway or network with a p-value of 0.01 or lower was considered as significantly dysregulated in the tumor.
Results and Discussion
Case Report
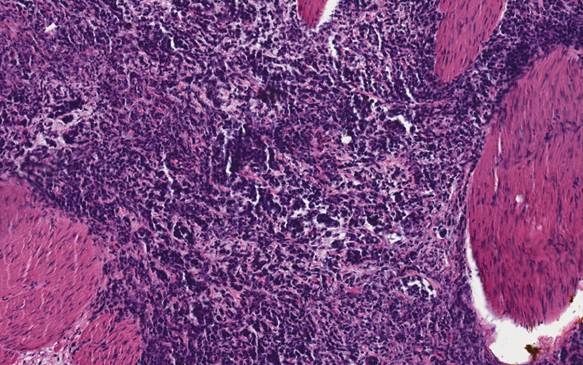
A 21-year-old- woman was in good health when presented with a one-month history of abdominal cramping and mild low back pain, fatigue and some abdominal bloating in July, 2006. This prompted work-up which included an ultrasound showing a cyst. CT scan confirmed that and in July of 2006 she underwent exploratory laparotomy with left salpingo-oophorectomy, omentectomy, radical tumor debulking and appendectomy. Initial pathology was consistent with a 12 cm left ovarian mass small cell sub-type (Figure 1). Vascular space invasion was present. Tumor involved the left pelvic peritoneum, the posterior cul-de-sac, the right bladder, peritoneum, the left external iliac node, left periaortic lymph node, left super renal lymph node. She was confirmed to be pathologic stage T3AN1MX. CAT scans at baseline showed pulmonary nodules of unclear significance. Initial treatment included multi-agent chemotherapy with cisplatin, bleomycin, cyclophosphamide, doxorubicin, and etoposide. The patient received a total of eight cycles in a period of five and one-half months. The patient had dramatic drop in her serum CA-125 level during this chemotherapy regimen from 243U/ml post-surgery to <30U/ml two months after treatment and to 7U/ml at the completion of chemotherapy. A routine PET/CT one month after the completion of therapy showed new uptake within the left external iliac lymph node as well as uptake within the right cul-de-sac as well as area near the rectum. As a result the patient underwent total abdominal hysterectomy with multiple lymph node samplings which were then sent for molecular profiling. Molecular profiling revealed multiple targets that suggest that single agent gemcitabine would be optimal. Patient was then initiated with gemcitabine as a single agent with the standard 1,000mg/m2 dose for three weeks in a row with one week off. After one month of therapy she began to experience pelvic discomfort. Imaging showed growth of mass in the cul-de-sac as well as growth in the left external iliac pelvic lymph nodes. Due to pain the patient was seen by a radiation oncologist and received radiation to both areas within the pelvis and the pelvic lymph node. Once the patient had completed the radiation and had a full hematologic recovery therapy was initiated with both irinotecan and bevacizumab based upon the previous molecular profiling results with the intent to repeat imaging one month after she started the therapy. Unfortunately, due to some hematuria, bevacizumab was unable to be initially added. As part of the work-up for hematuria patient was found to have bilateral hydronephrosis with renal insufficiency ultimately requiring bilateral nephrostomy tubes. Once the nephrostomy tubes had been added she was then re-started on bevacizumab in combination with irinotecan. After that initial treatment CT scans showed progression within the liver, retroperitoneum, and peritoneum. The patient then underwent palliative paracentesis which helped her symptomatically. Once again her profile was reviewed and docetaxel was felt to be an appropriate agent. The patient then went on to receive single agent docetaxel but after starting this for approximately three weeks, she once again developed intractable pain and was admitted to the hospital where she succumbed to her disease. The patient requested that an autopsy be performed after her death to try to help others and we followed her request.
H&E staining of the tumor shows the characteristics of a small cell carcinoma of the ovary.
Analysis of Gene Copy Number
Many malignant tumors and in particular poorly differentiated, highly aggressive ones are characterized in almost all instances by an unstable karyotype leading to large-scale chromosomal abnormalities. However, several reports using multiple methods have shown that SCCOHT tumors are essentially diploid [2, 18-20]. Chromosomal aberrations were investigated in one study by comparative genomic hybridization (CGH) for 19 SCCOHT tumors. All tumors had diploid DNA histograms, and repeatedly performed CGH analyses did not reveal any chromosomal aberration [21]. Likewise, the SCCOHT sample from our patient in this study was examined for an unstable karyotype by flow cytometry and by Agilent 244k aCGH (data not shown). In agreement with the previously reported findings, this tumor was diploid, and we were unable to detect any changes in chromosomal or gene copy number. Although aCGH may miss some minor changes it is intriguing that one of the most lethal gynecological malignancies does not appear to have significant chromosomal or gene copy abnormalities.
Analysis of Gene Expression
Global analysis of gene expression is a way to characterize tumors and find potentially disrupted pathways that might be able to be used in rationalizing better therapeutic targeting. Microarray analysis was used to compare the gene expression of this tumor to that of normal ovarian RNA. Based on the p-values calculated by the Agilent Feature Extraction software for the logarithm ratios of tumor vs. normal tissues (LogRatio Pvalue), we identified a total of 5,282 genes that showed significantly differential expression (LogRatio Pvalue <0.001) between the SCCOHT and normal ovary samples. Among them, 2,469 genes were up-regulated in the tumor (expression ratios of tumor vs. normal tissue ranging from 1.6 to 561) and 2,813 genes were down-regulated (expression ratios ranging from 0.6 to 0.004) (Additional file 1: Supplementary Table S1).
One group of transcripts identified as up-regulated in SCCOHT are cancer/testis (CT) antigens. These include MAGEA4, CSAG2, PRAME and DDX43. Adult ovarian tissue is usually reported to be CT antigen negative, but CT antigen expression has been reported in fetal ovary [22]. CT antigens provide attractive targets for cancer-specific immunotherapy, as CT genes are expressed in some normal tissues, such as the testis and in some cases placenta. These immunologically protected sites lack MHC I expression and as such, do not present 'self' antigens to T cells. CSAG2, also known as Taxol-resistance-associated gene (TRAG) is a cancer/testis antigen commonly overexpressed in melanoma cell lines and malignant melanoma. CSAG2 expression has been found to be predictive for clinical outcome in ovarian carcinoma patients.[23]
To further characterize the gene expression patterns of this tumor, we used MetaCore pathway/network enrichment analysis tool to identify pathways and processes that are most significantly altered in the tumor compared to normal ovarian cells. We conducted enrichment analysis of the differential expressed genes across four functional ontologies: canonical pathway maps, GO processes, GeneGo process networks, and disease networks, using the hypergeo-metric distribution [24]. The top 10 pathways and top 10 process networks were identified as significantly (P value <0.005) altered in the tumor tissue were listed in Tables 1 and 2, respectively.
Pathways significantly dysregulated in tumor.
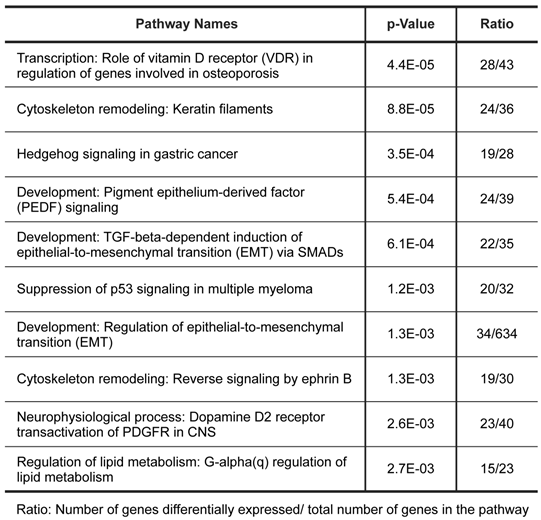
Biological process networks significantly dysregulated in tumor.
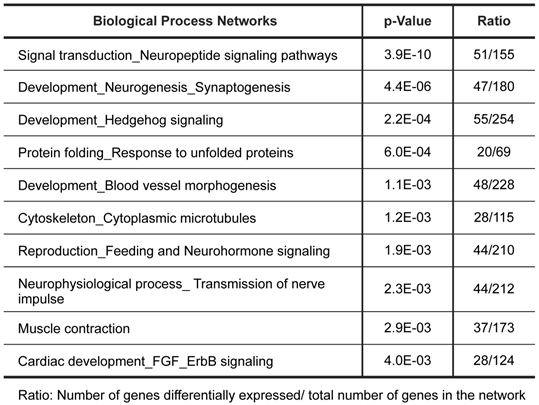
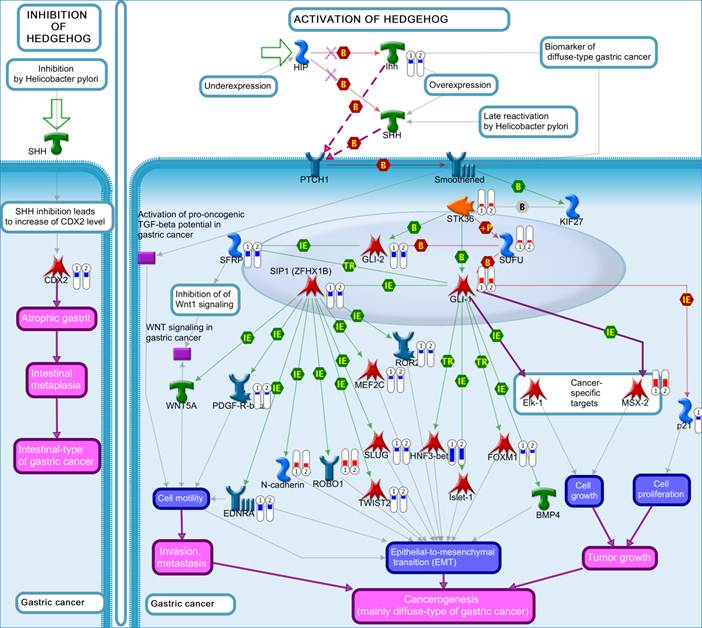
The top scored pathway maps is the vitamin D receptor (VDR) signaling pathway (Figure 2). VDR is modestly down-regulated whereas a number of genes that are transcriptionally regulated by VDR are highly up- or down-regulated. The expression of parathyroid hormone and parathyroid hormone-related proteins was not significantly changed. It is therefore not clear whether or not the dysregulation of this particular pathway is related to the hypercalcemia of the patient. Another top scored pathway map is the hedgehog signaling pathway (Figure 3). The hedgehog pathway has been reported to be activated in a number of cancers including basal cell carcinoma, medulloblastoma, colon, pancreatic, and ovarian cancer. What is interesting in this case is that it is the hedgehog regulated transcription factors (Gli1 and Gli2) and their downstream effectors are dysregulated while no significant change in the expression of the hedgehog ligands were observed. Further studies are needed to confirm and validate the dysregulation of these pathways/process networks and their roles in SCCOHT tumorigenesis and progression.
The vitamin D receptor pathway. Genes that were differentially expressed in the tumor in both replicates (Array #1 and Array #2 in Additional File 1: Supplementary Table S1) were mapped on pathway. The relative expression of the genes was visualized by thermometer-like symbols next to the protein icons (Red indicates upregulation, blue indicates downregualtion and the height of the bar indicates the level of change relative to normal control). Detailed legends are depicted in Additional file 2: Supplementary Figure S1.
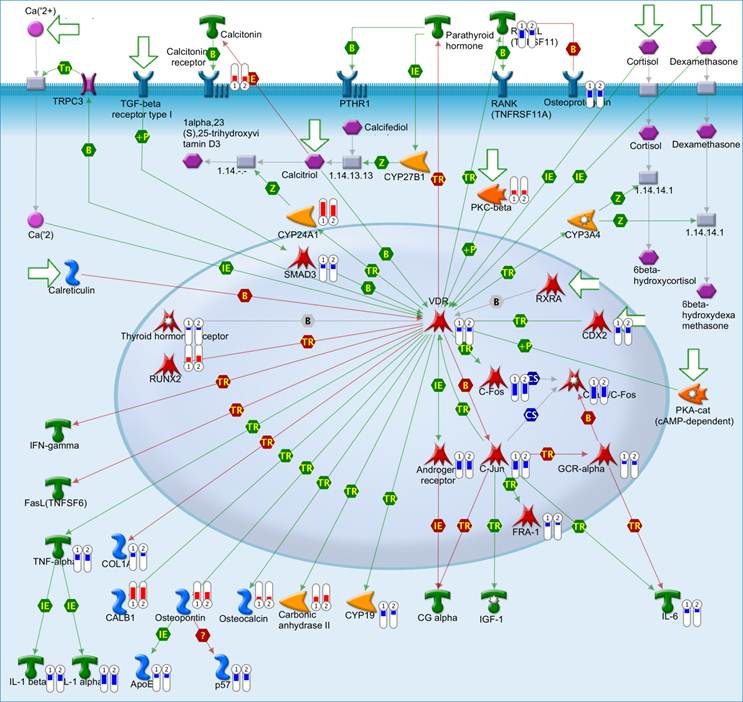
The hedgehog pathway. Genes that were differentially expressed in the tumor in both replicates (Array #1 and Array #2 in Additional File 1: Supplementary Table S1) were mapped on pathway. The gene expression data were represented in the map as described in Figure 2.
Potential Therapeutic Targets Based on Gene Expression Data
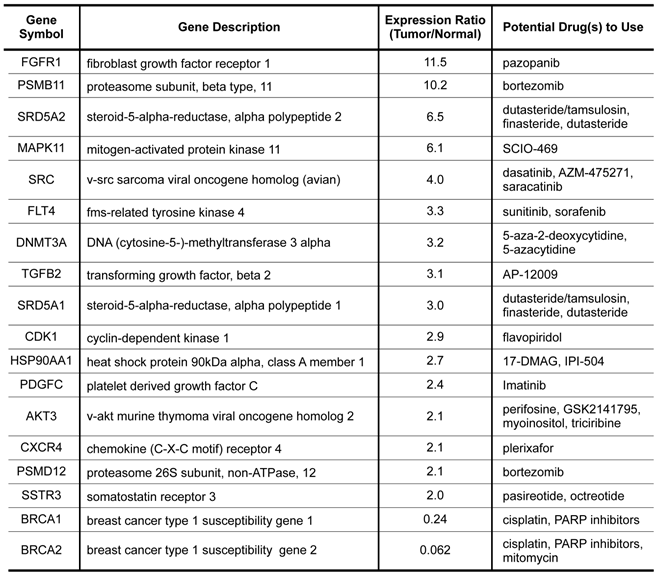
To further explore the possibility of using gene expression profiling to select potential therapeutics for this dreadful disease, we did a drug targets enrichment analysis in which we mapped the differentially expressed genes onto a drug-target database that contain a relationship table of drugs and their known targets. As shown in Table 3, a total of 18 genes that significantly overexpressed or underexpressed in the tumor have approved or experimental targeted therapeutic agents that could be potentially explored against the disease. Among the agents, cisplatin was used in the treatment of the patient. Other agents such as pazopanib, bortezomib, 5-azacytidine, and PARP inhibitors could be explored in future trials against this disease.
Molecular targets with available targeted agents identified from expression profiling.
Conclusions
A case study of a patient with SCCOHT is reported. Due to the low incidence of this disease, as well as a lack of information on the molecular profile of the disease, we molecularly profiled this patient's tumor sample. Specifically, we saw no chromosomal abnormalities by examination with aCGH. However, a number of genes highly up and down regulated compared to normal ovarian RNA were discovered by oligonucleotide microarray. Pathway analyses further identified several biological processes and signaling pathways that were significantly enriched for genes of which expressions were altered in the patient's tumor. Our results combined with previously published as well as future molecular characterization of this tumor type could help in the identification and development of new targeted therapeutics for the treatment of patients with SCCOHT.
Supplementary Material
Additional File 1Supplementary Table S1
Supplementary Fig. S1.
Acknowledgements
We would like to dedicate this work to the loving memory of Ms. Taryn Ritchey whose courage inspired us all.
Conflict of Interest
The authors have declared that no conflict of interest exists.
References
1. Scully RE. Tumors of the ovary and maldeveloped gonads. Washington, DC: Armed Forces Institute of Pathology. 1979
2. Young RH, Oliva E, Scully RE. Small cell carcinoma of the ovary, hypercalcemic type. A clinicopathological analysis of 150 cases. Am J Surg Pathol. 1994;18:1102-16
3. McCluggage WG. Ovarian neoplasms composed of small round cells: a review. Adv Anat Pathol. 2004;11:288-96
4. Clement PB. Selected miscellaneous ovarian lesions: small cell carcinomas, mesothelial lesions, mesenchymal and mixed neoplasms, and non-neoplastic lesions. Mod Pathol. 2005;18(Suppl 2):S113-S129
5. McCluggage WG. The Value of Immunohistochemistry as a Diagnostic Aid in Ovarian Neoplasia. Pathol Case Rev. 2006;11:31-7
6. Young RH, Goodman A, Penson RT. et al. Case records of the Massachusetts General Hospital. Case 8-2010. A 22-year-old woman with hypercalcemia and a pelvic mass. N Engl J Med. 2010;362:1031-40
7. Harrison ML, Hoskins P, du Bois A. et al. Small cell of the ovary, hypercalcemic type -- analysis of combined experience and recommendation for management. A GCIG study. Gynecol Oncol. 2006;100:233-8
8. Hamilton S, Beattie GJ, Williams AR. Small cell carcinoma of the ovary: a report of three cases and review of the literature. J Obstet Gynaecol. 2004;24:169-72
9. McCormick TC, Muffly T, Lu G, Shoup B. Aggressive small cell carcinoma of the ovary, hypercalcemic type with hypercalcemia in pregnancy, treated with conservative surgery and chemotherapy. Int J Gynecol Cancer. 2009;19:1339-41
10. Barondeau J, Rodgers M, Braun L. et al. Small cell ovarian carcinoma: a rare, aggressive tumor masquerading as constipation in a teenager with a fatal outcome. J Pediatr Hematol Oncol. 2010;32:e139-141
11. Dykgraaf RH, de Jong D, van Veen M. et al. Clinical management of ovarian small-cell carcinoma of the hypercalcemic type: a proposal for conservative surgery in an advanced stage of disease. Int J Gynecol Cancer. 2009;19:348-53
12. Nelsen LL, Muirhead DM, Bell MC. Ovarian small cell carcinoma, hypercalcemic type exhibiting a response to high-dose chemotherapy. S D Med. 2010;63:375-7
13. Martinez-Borges AR, Petty JK, Hurt G. et al. Familial small cell carcinoma of the ovary. Pediatr Blood Cancer. 2009;53:1334-6
14. Epstein EH. Basal cell carcinomas: attack of the hedgehog. Nat Rev Cancer. 2008;8:743-54
15. Von Hoff DD, LoRusso PM, Rudin CM. et al. Inhibition of the hedgehog pathway in advanced basal-cell carcinoma. N Engl J Med. 2009;361:1164-72
16. Dezso Z, Nikolsky Y, Nikolskaya T. et al. Identifying disease-specific genes based on their topological significance in protein networks. BMC Syst Biol. 2009;3:36
17. Nikolsky Y, Sviridov E, Yao J. et al. Genome-wide functional synergy between amplified and mutated genes in human breast cancer. Cancer Res. 2008;68:9532-40
18. Forster C, Ostertag H, Schmitt J, Roessner A. Small cell carcinoma of the ovary, hypercalcemic type. A case report with immunohistochemical, ultrastructural and cytophotometric analysis and review of the literature. Gen Diagn Pathol. 1997;142:365-70
19. Lamovec J, Bracko M, Cerar O. Familial occurrence of small-cell carcinoma of the ovary. Arch Pathol Lab Med. 1995;119:523-7
20. Eichhorn JH, Bell DA, Young RH. et al. DNA content and proliferative activity in ovarian small cell carcinomas of the hypercalcemic type. Implications for diagnosis, prognosis, and histogenesis. Am J Clin Pathol. 1992;98:579-86
21. Hauptmann S, Schmidt H, Bilkenroth U. et al. Small Cell Carcinoma of the Ovary of the Hypercalcemic Type: A Highly Aggressive Neoplasm without CGH Aberrations. Atlanta, Georgia: US and Canadian Acad Pathol 2006 Ann Meeting. 2006 Abstr836
22. Nelson PT, Zhang PJ, Spagnoli GC. et al. Cancer/testis (CT) antigens are expressed in fetal ovary. Cancer Immun. 2007;7:1
23. Materna V, Surowiak P, Kaplenko I. et al. Taxol-resistance-associated gene-3 (TRAG-3/CSAG2) expression is predictive for clinical outcome in ovarian carcinoma patients. Virchows Arch. 2007;450:187-94
24. Dezso Z, Nikolsky Y, Sviridov E. et al. A comprehensive functional analysis of tissue specificity of human gene expression. BMC Biol. 2008;6:49
Author contact
Corresponding author: Daniel D. Von Hoff, M.D., Division of Clinical Translational Res. The Translational Genomics Research Institute, 13208 E Shea Blvd, Scottsdale, AZ 85259. Phone: 602-358-8327 Email: dvhorg.

 Global reach, higher impact
Global reach, higher impact