Impact Factor ISSN: 1837-9664
J Cancer 2017; 8(6):940-949. doi:10.7150/jca.17496 This issue Cite
Research Paper
Mutual Regulation of MiR-199a-5p and HIF-1α Modulates the Warburg Effect in Hepatocellular Carcinoma
1. Department of Hepatobiliary and Pancreatic Surgery, Sun Yat-sen University Cancer Center, Guangzhou, China;
2. Department of Radiation Oncology, Sun Yat-sen University Cancer Center, Guangzhou, China;
3. State Key Laboratory of Oncology in South China, Guangzhou, China;
4. Collaborative Innovation Center for Cancer Medicine, Guangzhou, China.
Received 2016-9-6; Accepted 2016-12-23; Published 2017-3-12
Abstract
The Warburg effect is one of the major metabolic changes of cancer cells, which characterized by high level of glycolysis even in the presence of oxygen. However, the role of microRNAs (miRNAs) in regulating the glycolytic switch in cancer cells has not been well explored. In this study, we demonstrated that miR-199a-5p acts as a suppressor of the Warburg effect in hepatocellular carcinoma (HCC). MiR-199a-5p directly targets the 3′-untranslated region (UTR) of hypoxia-inducible factor-1α (HIF-1α), thereby suppressing glucose uptake, lactate production, cell growth, and expression of HIF-1α downstream glycolytic genes of HCC cells. Moreover, under hypoxic conditions, the expression of miR-199a-5p is suppressed by the up-regulation of HIF-1α. Thus, mutual regulation between miR-199a-5p and HIF-1α forms a positive feedback loop to promote glycolysis in HCC cells. Furthermore, miR-199a-5p is down-regulated in human HCC tissues and its low-level expression is associated with a worse survival of patients with HCC. Our findings suggest that miR-199a-5p/HIF-1α axis is critical in the regulation of the Warburg effect and also implicate miR-199a-5p as a potential therapeutic target for HCC.
Keywords: miR-199a-5p, the Warburg effect, hypoxia, HIF-1α, hepatocellular carcinoma.
Introduction
Altered energy metabolism is a typical hallmark of cancer [1]. A well-known metabolic phenotype in cancer cells is the “Warburg effect”, which characterized by high rates of glucose uptake and lactate production even in the presence of oxygen, leading to a state termed “aerobic glycolysis” [2]. Most cancer cells, including hepatocellular carcinoma (HCC) cells, display this fundamental change in their cellular metabolism to meet the increased energetic and anabolic demands required for rapid cell growth [2-4]. By using glucose as a carbon source to generate energy and metabolic intermediates, the metabolic shift toward the Warburg effect can provide non-oxidative ATP and biosynthetic advantages to proliferating cells [5]. However, the underlying mechanisms of the Warburg effect remain largely unknown.
MicroRNAs (miRNAs) are small noncoding RNAs that can regulate gene expression at the posttranscriptional level [6, 7]. The potential roles of microRNAs in energy metabolism have been recently studied [8-10]. These studies revealed that a few microRNAs are involved in regulation of the Warburg effect by targeting specific enzymes [11-13]. For example, miR-122 targets pyruvate kinase M2 (PKM2) and affects glycolysis in HCC [14].MiR-155/miR-143 cascade regulates glycolysis by targeting hexokinase 2 (HK2) in breast cancer cells [15]. MiR-199a-5p has been reported to be down-regulated and function as tumor-suppressor in HCC [16-18]. A recent study found that miR-199a-5palso participates in cellular metabolism by targeting HK2 in HCC [19]. However, given that miRNAs often exert their functions through multiple target genes [20], whether other molecules and pathways are involved in regulation of the Warburg effect by miR-199a-5p in HCC requires further investigation.
In this study, we demonstrate that HIF-1α is directly targeted by miR-199a-5p and miR-199a-5p/HIF-1α axis plays an important role in the regulation of the Warburg effect in HCC cells. We characterize a positive feedback loop between miR-199a-5p and HIF-1α to promote glycolysis. Our study suggests mutual regulation of miR-199a-5p and HIF-1α as a newly defined mechanism contributing to the Warburg effect and also implicates miR-199a-5p as a potential therapeutic target for HCC.
Methods
Cell lines
The cell lines used were HEK293T and the HCC cell lines, Huh-7 and HepG2.HEK293T, Huh-7, and HepG2 cells were obtained from American Type Culture Collection (ATCC). All cell lines were cultured in Dulbecco's Modified Eagles Medium (DMEM) (GIBCO, Carlsbad, CA) supplemented with 10% fetal bovine serum (FBS) at 37°C in 5% CO2. To create hypoxia conditions, cells were placed in a hypoxic chamber (Forma Scientific, Marietta, OH), where the cells were maintained at 37°C in an atmosphere containing 5% CO2, 1% O2 and 94% N2. Alternately, cells were treated for 4 hours with 100μM/L of the hypoxia-mimicking CoCl2 (Sigma-Aldrich, St Louis, MO).
Patients and specimens
Human HCC and their corresponding non-tumor liver tissue samples from 104 patients were collected at the time of surgical resection at Sun Yat-sen University Cancer Center, from 2003 to 2007. Fresh samples were immediately snap-frozen in liquid nitrogen and stored at -80°C until use. The pathological diagnoses were confirmed based on detailed examination of sections stained with H&E by the Department of Pathology. This study protocol conformed to the ethical guidelines of the Declaration of Helsinki and the Ethics Committee of Sun Yat-sen University Cancer Center approved use of human tissues. Written informed consent was obtained from each patient who participated in this study.
RNA oligo ribonucleotides, plasmids and cell transfections
All RNA oligo ribonucleotides including miRNA mimics, inhibitors and small interfering RNA against HIF-1α mRNA (siHIF-1α) were purchased from Gene Pharma (Shanghai, China). HIF-1α-ΔODD cDNA construct (synthesized by GenScript, Piscataway/NJ) that lacked the oxygen-dependent degradation domain (ODD401-603) was ligated into the KpnI site of pcDNA3.1 (+). To create luciferase reporter construct, a wild-type 3'-UTR fragment of human HIF-1αmRNA that contained the putative binding sites for miR-199a-5p was PCR-amplified and insert into the pGL3cm vector, which was generated previously. The mutant 3'-UTR, which carried the mutated sequence in the complementary site for the seed region of miR-199a-5p, was generated by site-specific mutagenesis based on the construct with wild-type 3'-UTR. Reverse transfections of RNA oligo ribonucleotides were performed using Lipofectamine-RNAiMAX (Invitrogen). A total of 50 nM of RNA duplex or 200nM of miRNA inhibitor were used for each transfection. Transfection of plasmid DNA or cotransfection of RNA duplex with plasmid DNA was performed using Lipofectamine 2000 (Invitrogen) according to the manufacturer's instructions.
RNA extraction, reverse transcription, and quantitative real-time RT-PCR
Total RNA from cancer cells or tissue samples was extracted by using the TRIzol reagent (Invitrogen, Carlsbad, CA, USA). For miRNA expression analysis, a stem-loop qRT-PCR assay was performed using Bulge-Loop™ miRNA qRT-PCR kits (Ribobio, Guangzhou, China) according to the manufacturer's instructions. U6 small nuclear RNA was used as an endogenous control. For mRNA expression analysis, the cDNA synthesis was performed using a Superscript First-Strand Synthesis System for RT-PCR (Invitrogen) according to the manufacturer's protocol. β-actin was used as an endogenous control. Quantitative real-time PCR reactions were conducted using Platinum SYBR Green qPCR SuperMix-UDG reagents (Invitrogen). The samples were analyzed using an ABI-Prism 7900HT system (Applied Biosystems, Foster City, CA) by standardized protocol. The relative levels for miRNA and mRNA expression were analyzed by the comparative 2-ΔΔCt method. Each sample was carried out in triplicate.
Glucose uptake assay
Cells were cultured in 6-well plate. After transfection with miR-199a-5p mimics or vector for 48 hours, cells were refreshed with serum-starved (0.1% FBS) and glucose-free DMEM. Sixteen hours later, cells were treated with 50mM 2-[N-(7-nitrobenz-2-oxa-1,3-diazol-4-yl) amino]-2-deoxy-D-glucose (2-NBDG) (Invitrogen) for 1h, and then glucose uptake was quantified using flow cytometry (BD, Franklin Lakes, NJ, USA).
Lactate production assay
Cells were cultured in 6-well plate. After transfection with miR-199a-5p mimics or vector for 48 hours, culture medium was removed from cells and lactate concentration was determined using lactate test strips and Accutrend Lactate analyzer (Accutrend Lactate, Roche). Results were normalized with cell number.
Measurement of LDH activity and extracellular pH
Measurement of LDH activity was performed using Lactate Dehydrogenase Activity Assay Kit (BioVision) according to the manufacturer's protocols. Absorbance was measured using GENios Plus Plate Reader (Tecan). Extracellular pH in the medium was measured with a pH meter (PB-11 Basic Meter; Sartorius). Measurements were made within 2 minutes of sample collection. All results were normalized with cell number.
Cell proliferation assay and colony formation assay
The cell proliferation was determined by way of WST-8 staining with Cell Counting Kit-8 (CCK-8, Dojindo, Japan) following the manufacturer's protocol. For colony formation assay, 1000 cells were plated onto six-well plates and incubated for 2 weeks. Cells were then stained with crystal violet, and the numbers of colonies per well were counted. All experiments were performed for 3 times, and the average of the results was calculated.
Luciferase reporter assay
HEK293T cells plated in a 48-well plate were co-transfected with 10 nM RNA duplex, 10 ng of firefly luciferase reporter containing the wild-type or mutant 3'-UTR of HIF-1α, and 2 ng of pRL-TK (Promega, Madison, WI). Cells were collected 48 hours after last transfection and analyzed using Dual-Luciferase Reporter Assay System (Promega). The pRL-TK vector expressing renilla luciferase was used as an internal control for normalization. Transfections were performed in duplicate and repeated in at least three independent experiments.
Western blot
For western blot analysis, cells were lysed on ice in RIPA Lysis Buffer (Pierce Biotechnology, IL, USA). The protein concentration was measured using BCA Protein Assay Kit (Pierce Biotechnology, IL, USA). Then 20 μg of total protein extract with loading buffer were heated at 100℃ for 5 minutes. After resolved on SDS-PAGE gels, the proteins were transferred onto PVDF Membrane (Millipore). Finally, the membranes were blotted with the appropriate primary antibody followed by secondary antibodies. The immunoblots were visualized using an enhanced chemiluminescence (ECL) kit. Anti-HIF-1α was purchased from Abcam (Cambridge, MA). β-actin (Abcam) was used as loading control.
Immunohistochemisty (IHC)
Immunohistochemical staining for HIF-1α was performed as follows: tissue sections were deparaffinized, rehydrated, and treated with 3% H2O2 for 15 minutes to inhibit endogenous peroxidase activity. Following heat-induced epitope retrieval in 10mM citrate buffer (pH 6.0) in a microwave for 30 minutes, the slides were incubated at 4°C overnight with a prediluted primary monoclonal anti- HIF-1α antibody (dilution 1:500; Abcam). After incubation with a rabbit anti-mouse secondary antibody, a reaction was performed using the EnVision plus detection system (DAKO, Carpinteria, CA). Slides were washed in distilled water, counterstained with hematoxylin, dehydrated with xylene, and coverslipped using cytoseal. External positive and negative controls were included in each batch of slides.
The results of IHC were evaluated by using a semi-quantitative scoring system including both intensity of staining and percentage of positive cells. The intensity was scored as follows: 0, no staining; 1, weak staining; 2, moderate or strong staining. The fraction was scored based on the proportion of positively stained cells (0-100%). The intensity and fraction scores were then multiplied to obtain a final score, which ranged 0-2.
Statistical analysis
Data were presented as mean ± SEM. The Student's t-test, Chi-square test or Fisher's exact test were used for analyzing the differences between groups. The survival curves were estimated by the Kaplan-Meier method, and differences between curves were compared by the log-rank test. Multivariate Cox regression analysis was used to identify independent prognostic factors for survival. All statistical analyses were performed with the SPSS (Statistical Package for the Social Sciences) 16.0 software (SPSS, Chicago, IL) and GraphPad Prism 5.0 software (GraphPad Software, San Diego, CA). Differences with P values < 0.05 were defined as statistically significant.
Results
MiR-199a-5p inhibits the Warburg effect in HCC cells and suppresses cell growth
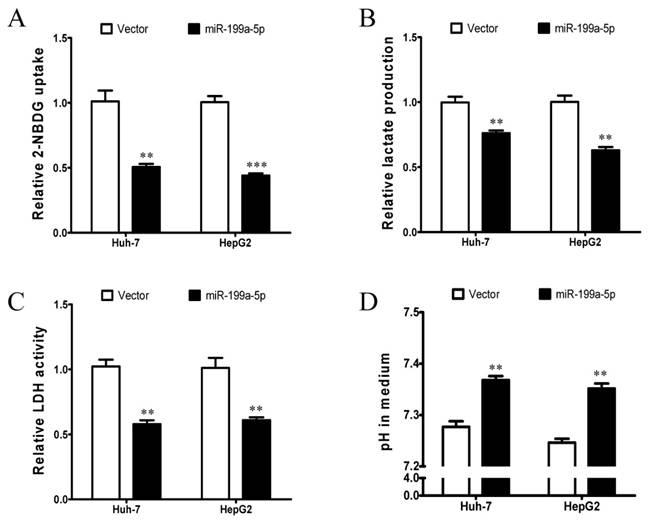
To investigate the potential role of miR-199a-5p in regulation of the Warburg effect in HCC, in vitro assays including glucose uptake, lactate production, LDH activity assays and pH value measurement were conducted in Huh-7 and HepG2 cells. Huh-7 and HepG2 cells were transiently transfected with miR-199a-5p mimics or vector controls. Successful re-expression of mature miR-199a-5p was confirmed by quantitative real-time PCR (qPCR). Remarkably, both Huh-7 and HepG2 cells with re-expression of miR-199a-5p displayed a significant decrease in glucose uptake, lactate production and LDH activity but higher pH value in the culture medium in comparison to cells transfected with vector controls (Figure 1A-D). These findings indicate that miR-199a-5p can inhibit the Warburg effect in HCC cells.
miR-199a-5p inhibits the Warburg effect in HCC cells. A, Glucose uptake; B, Lactate production; C, LDH activity; and D, culture medium pH were measured in Huh-7 and HepG2 cells transfected with miR-199a-5p or vector controls. **P< 0.01, ***P < 0.001.
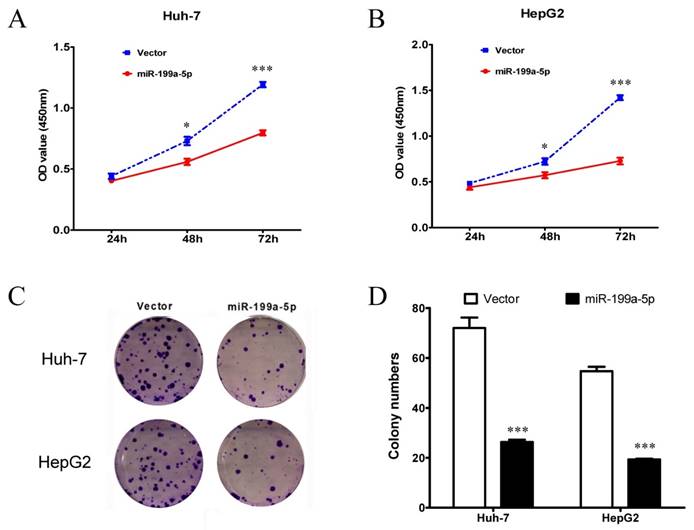
Since miR-199a-5p can regulate the Warburg effect in HCC cells, it suggested that miR-199a-5p could affect cell growth. Thus, MTT assay and colony formation assay were performed to examine the cell proliferation ability. Both Huh-7 and HepG2 cells transfected with miR-199a-5p mimics showed a significant inhibition of proliferation compared with those transfected with vector controls (Figure 2A-B). Moreover, cells transfected with miR-199a-5p mimics displayed much fewer and smaller colonies compared with controls (Figure 2C-D).
miR-199a-5p suppresses the growth of HCC cells. A, B, Cell proliferation assay results. MTT assay using CCK-8 was performed to test cell proliferation of Huh-7 and HepG2 cells transfected with miR-199a-5p or vector controls. C, D, Colony formation assay results. Representative results of colony formation, and colony numbers of Huh-7 and HepG2 cells transfected with miR-199a-5p or vector controls. *P< 0.05, ***P < 0.001.
HIF-1α is a direct target of miR-199a-5p
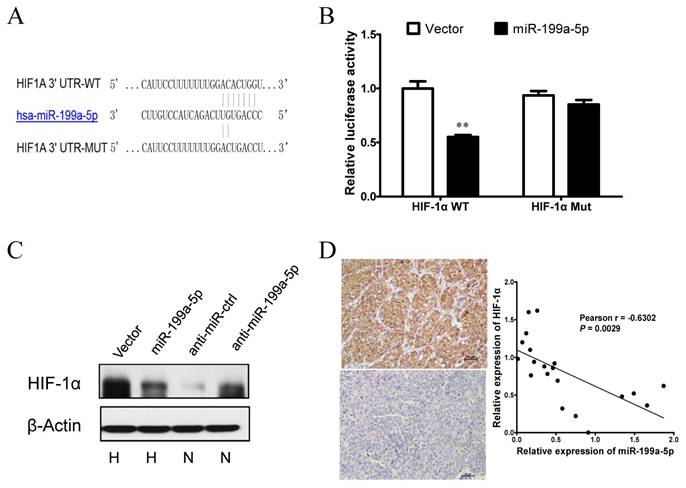
Next, we explored the molecular mechanisms responsible for the function of miR-199a-5p that was observed above. As HIF-1α was previously reported to be a target of miR-199a-5p in cardiomyocytes [21], we postulated that miR-199a-5p may exert the function above by direct targeting HIF-1α. To verify whether HIF-1α is a direct target of miR-199a-5p in HCC, a dual-luciferase reporter system was first employed. Co-transfection of miR-199a-5p significantly suppressed the firefly luciferase activity of the reporter with wild-type 3'-UTR of HIF-1α but not that of the mutant reporter (Figure 3A-B), indicating that miR-199a-5p can bind to the 3'-UTR of HIF-1α transcripts. Furthermore, ectopic expression of miR-199a-5p showed a clear attenuation of HIF-1α protein expression in HepG2 cells under hypoxia, whereas inhibition of miR-199a-5p showed an increase of HIF-1α protein expression under normoxia (Figure 3C). This finding supported that HIF-1α protein expression was inversely associated with miR-199a-5p expression in the HCC cell system. Correlation between the miR-199a-5p level and the expression of HIF-1α was further examined in HCC tissues. HIF-1α expression was analyzed by immunohistochemistry, and miR-199a-5p was determined by qPCR in a set of 20 HCC specimens. Markedly, miR-199a-5p level was inversely correlated with HIF-1α expression (Figure 3D). Taken together, these results indicate that HIF-1α is a direct target of miR-199a-5p in HCC.
HIF-1α is a direct target of miR-199a-5p in HCC. A, Diagram of the putative binding sites of miR-199a-5p on the 3′-UTRs of HIF-1α. The mutant sequences of 3′-UTRs of HIF-1α used in the luciferase reporters are indicated. B, Relative luciferase activity with HIF-1α wild type or mutant type 3′C.-UTR after co-transfection with miR-199a-5p or vector controls in HEK-293T cells. C, HIF-1α protein levels in HepG2 cells with miR-199a-5p overexpression under hypoxia or miR-199a-5p inhibition under normoxia. D, The inverse correlation between miR-199a-5p and HIF-1α levels in HCC tissues. (Left) Representative immunohistochemical staining of HIF-1α in HCC tissues. (Right) Inverse correlation between miR-199a-5p and HIF-1α levels in 20 HCC specimens. Pearson's correlation coefficient (r) and statistical significance are indicated. **P< 0.01.
MiR-199a-5p inhibits the Warburg effect and cell growth by direct targeting HIF-1α
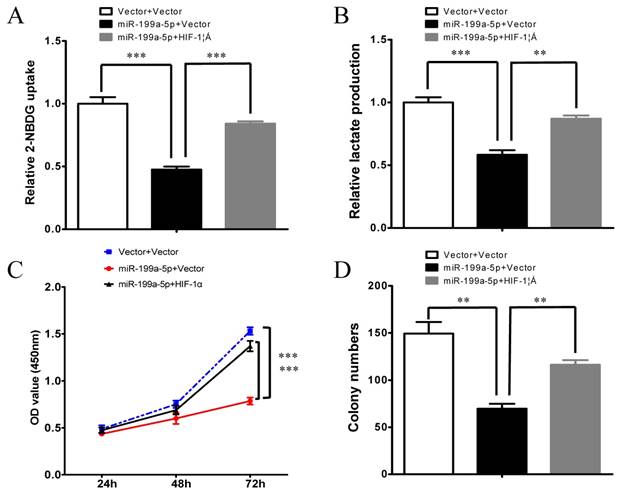
To further verify whether miR-199a-5p regulates the Warburg effect through HIF-1α in HCC, we subsequently investigated whether HIF-1α could counteract the function of miR-199a-5p. Notably, overexpression of HIF-1α attenuated miR-199a-5p-mediated reduction in glucose uptake and lactate production (Figure 4 A-B). Furthermore, overexpression of HIF-1α reversed the inhibition of cell proliferation ability in miR-199a-5p transfected cells (Figure 4C-D).
miR-199a-5p inhibits the Warburg effect and cell growth by direct targeting HIF-1α. A, Glucose uptake; B, Lactate production; C, Cell proliferation assay; and D, Colony formation assay were measured in HepG2 cells transfected with miR-199a-5p or vector controls. Ectopic expression of HIF-1α abolished the effects of miR-199a-5p on the Warburg effect and cell growth in HCC cells. **P< 0.01, ***P < 0.001.
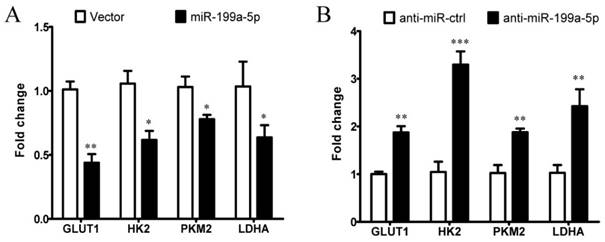
We next examined whether miR-199a-5p affects the expression of HIF-1α downstream target genes involved in glycolytic regulation. As assessed by qPCR, we found that overexpression of miR-199a-5p reduced the expression levels of glucose transporter 1 (GLUT1), HK2, PKM2, and lactate dehydrogenase A (LDHA) in Huh-7 cells (Figure 5A), whereas inhibition of the endogenous miR-199a-5p increased the expression levels of GLUT1, HK2, PKM2, and LDHA in cultured cells (Figure 5B). These results indicate that miR-199a-5p can affect the expression of multiple glucose-related genes through targeting HIF-1α in HCC, causing enhanced glycolysis. Collectively, our data indicate that miR-199a-5p regulates the Warburg effect via targeting HIF-1α in HCC.
miR-199a-5p affects the expression of HIF-1α downstream target genes involved in glycolytic regulation. A, Expression of HIF-1α target genes in Huh-7 cells transfected with miR-199a-5p or vector controls, using qPCR. B, Expression of HIF-1αtarget genes in Huh-7 cells transfected with anti-miR-199a-5p or anti-miR-ctrl, using qPCR. *P< 0.05, **P< 0.01, ***P < 0.001.
Expression of miR-199a-5p is regulated by hypoxia and HIF-1α in HCC cells
Given that hypoxia and HIF-1α are closely associated with the Warburg effect, our results above promoted us to determine the effect of hypoxia and HIF-1α on miR-199a-5p levels in HCC cells. We observed that miR-199a-5p expression is significantly decreased in HCC cells under hypoxia (Figure 6A) or following CoCl2 treatment (Figure 6B), which mimics hypoxic conditions. These results indicate that miR-199a-5p expression in HCC cells is down-regulated in response to hypoxia, which is a common microenvironment in solid tumor.
Hypoxia and HIF-1α inhibits miR-199a-5p expression in HCC cells. A, miR-199a-5p expression levels in Huh-7 and HepG2 cells cultured in both normoxic and hypoxic conditions (1% O2). B, miR-199a-5p expression levels in Huh-7 and HepG2 cells treated with or without CoCl2. C, miR-199a-5p expression levels in Huh-7 cells with or without HIF-1αoverexpression.Huh-7 cells were transfected with HIF-1α-ΔODD plasmids or vector controls for 24 h. D, miR-199a-5p expression levels in Huh-7 cells with or without HIF-1αknockdown. Huh-7 cells were transfected with si-HIF-1αfor 48 h and then treated with 100 μM CoCl2 for 24 h. **P< 0.01, ***P < 0.001.
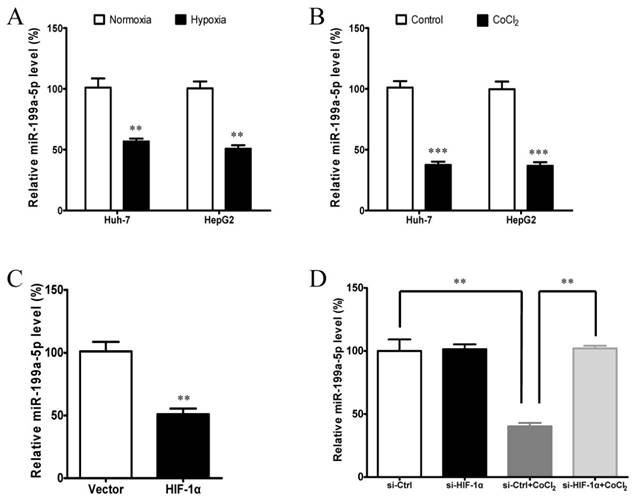
We then reasoned that the suppression of miR-199a-5p under hypoxia might be due to the results of HIF-1α induction. To test this hypothesis, we examined whether HIF-1α can regulate the expression of miR-199a-5p. The results showed that ectopic expression of HIF-1α by transfection of plasmid HIF-1α-ΔODD could greatly suppress the expression of miR-199a-5p (Figure 6C). Conversely, knockdown of HIF-1α by siRNA could remarkably increase the expression of miR-199a-5p under hypoxic condition (Figure 6D). These results indicate that the induction of HIF-1α is responsible for the decrease of miR-199a-5p levels in HCC cells under hypoxia.
Down-regulation of miR-199a-5p is a frequent event in human HCC and is associated with poor prognosis
To further investigate the clinic pathological and prognostic significance of miR-199a-5p in patients with HCC, the levels of miR-199a-5p in a cohort of 104 paired HCC and adjacent non-tumor tissues were examined by qPCR. As shown in Figure 7A, after normalization to U6 expression levels, the expression level of miR-199a-5p in HCC tissues was significantly lower than that in adjacent non-tumor tissues (median fold change of T/ NT = 0.65, P< 0.001). MiR-199a-5p was down-regulated in 66 of 104 patients (63.5%) totally (Figure 7B).
miR-199a-5p is down-regulated in HCC and associated with a poor prognosis. A, The expression levels of miR-199a-5p in HCC tissues and adjacent non-tumor tissues (n =104). ***P < 0.001. B, Down-regulation of miR-199a-5p was observed in 66/104 (63.5%) HCCs. C, Kaplan-Meier analysis of 104 HCC patients according to miR-199a-5p level. Overall survival rate in patients with low miR-199a-5p expression level was significantly worse than that in patients with high miR-199a-5p expression level (log-rank P = 0.024).
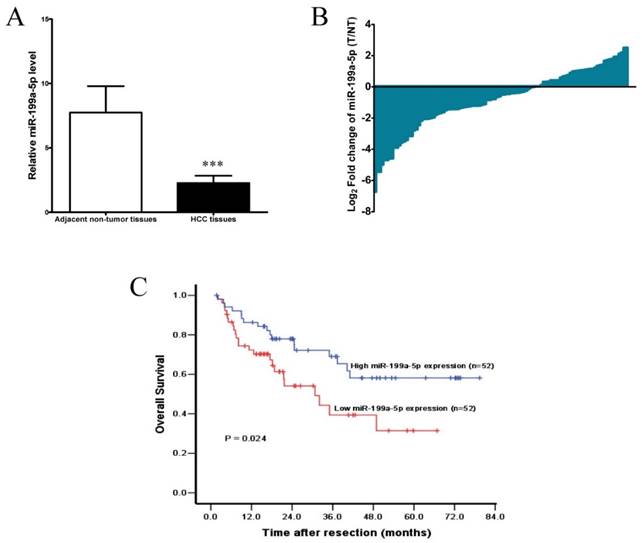
Accordingly, the median fold change (0.65) of miR-199a-5p was used as a cut-off value, thus 52/104 (50.0%) HCCs had low-level expression of miR-199a-5p, while 52/104 (50.0%) HCCs had high-level expression of miR-199a-5p.Kaplan-Meier analysis revealed that low-level expression of miR-199a-5p was associated with short overall survival of patients with HCC (Figure 7C). The 1, 3, and 5-year overall survival rate in the low expression group was 72.3%, 39.4%, and 31.5%, respectively, compared with 86.3%, 69.0%, and 58.1%, respectively, in the high expression group (log-rank test, P = 0.024). Further multivariate Cox regression analysis indicated that low-level expression of miR-199a-5p is an independent prognostic factor for poor survival of patients with HCC after adjustment for other significant variables (P =0.046, Table 1).
Univariate and multivariate Cox regression analyses for overall survival
| Variable | Univariate analysis | Multivariate analysis | ||||||
|---|---|---|---|---|---|---|---|---|
| HR (95% CI) | P | HR (95% CI) | P | |||||
| miR-199a-5p (low versus high) | 2.030 (1.082-3.809) | 0.024 | 1.943 (1.011-3.734) | 0.046 | ||||
| Age (≥50 versus <50 years) | 1.324 (0.707-2.481) | 0.381 | - | - | ||||
| Gender (M versus F) | 0.961 (0.296-3.117) | 0.947 | - | - | ||||
| HBsAg (positive versus negative) | 1.994 (0.781-5.093) | 0.149 | - | - | ||||
| Child-Pugh (B versus A) | 2.062 (0.803-5.295) | 0.133 | - | - | ||||
| AFP (≥200 versus <200 μg/L) | 2.085 (1.111-3.912) | 0.022 | 1.400 (0.712-2.751) | 0.330 | ||||
| Tumor size (≥7 cm versus <7 cm) | 3.098 (1.578-6.082) | 0.001 | 2.189 (1.025-4.675) | 0.043 | ||||
| Tumor number (multiple versus solitary) | 1.472 (0.794-2.729) | 0.219 | - | - | ||||
| Vascular invasion (present versus absent) | 3.450 (1.831-6.500) | <0.001 | 1.907 (0.880-4.133) | 0.102 | ||||
| Edmondson-Steiner (III-IV versus I-II) | 2.105 (0.954-4.641) | 0.065 | - | - | ||||
| TNM stage (II-III versus I ) | 2.353(1.199-4.619) | 0.013 | 1.303(0.564-3.010) | 0.536 | ||||
Statistically significant values are given in bold
Discussion
Although there has been considerable progress in understanding the functional significance and probable mechanisms of the Warburg effect, the potential role of miRNAs in cancer cell metabolism, particularly in liver cancer, is still needed to be further explored. In this study, we provided evidence that miR-199a-5p is an important regulator of the Warburg effect in HCC cells. More importantly, we identified HIF-1α as a direct target of miR-199a-5p, and induction of the metabolic shift by miR-199a-5p is dependent on HIF-1α. Moreover, we found that mutual regulation of miR-199a-5p and HIF-1α forms a positive feedback loop to promote glycolysis. Our findings indicate that mutual regulation of miR-199a-5p and HIF-1α is crucial for regulating the biological processes of glucose metabolism, and has an important role in tumor progression of HCC.
There is growing evidence supporting that miRNAs play critical roles in regulation of cellular metabolism [8-13]. Recent studies revealed that a few miRNAs, such as miR-23a [22, 23], miR-34a [24], miR-122 [14, 25], and miR-155 [15], are involved in glycolysis, gluconeogenesis, and the pentose phosphate pathway (PPP). In a previous study, miR-199a-5p has been found to have important metabolic implications in cardiomyocytes [21]. In the meantime of our present study, another group recently also reported that miR-199a-5p could regulate glycolysis and lactate production by targeting HK2 in HCC [19]. Similar to their results, our data demonstrated that miR-199a-5p can inhibit aerobic glycolysis and cell growth in HCC cells. Thus, our study together with other studies confirms that miR-199a-5p is an important miRNA linking to cancer cell metabolism.
The underlying mechanism of altered metabolism of cancer cells might be complicated due to multiple genes and pathways involved [26]. Of note, HK2 has been recently reported to be a target of miR-199a-5p in HCC cells, which is involved in glycolysis [19]. As miRNAs often exert their functions via multiple target genes, in the present study, we focused on the potential role of miR-199a-5p/HIF-1α axis contributing to the metabolic change of HCC cells. Our data demonstrated that HIF-1α is a direct target of miR-199a-5p. And most importantly, we discovered that the inhibition of the Warburg effect by miR-199a-5p is dependent on HIF-1α, as re-expression of HIF-1α ablates the effects of miR-199a-5p on aerobic glycolysis. As HIF-1α is a master regulator gene of glycolysis [27, 28], we also demonstrate that expressions of HIF-1α downstream target genes, including GLUT1, HK2, PKM2, and LDHA, were all affected by miR-199a-5p, also suggesting that miR-199a-5p links to metabolic control through its downstream regulation of HIF-1α. Our work identifies HIF-1α as a key mediator of the metabolic effects triggered by miR-199a-5p in HCC.
Hypoxia in microenvironment is one of the most common features of solid tumors [29, 30]. Cancer cells adjust their transcription and translation to maintain cellular homeostasis and enable an adaptive survival under hypoxic stress conditions [31, 32]. A number of miRNAs are regulated by hypoxia and thus termed hypoxia-responsive miRNA (HRM) [33-36]. Consistent with previous studies, we found a reduction of miR-199a-5p expression in HCC cells exposed to hypoxia or a hypoxic condition reproduced by the use of cobalt chloride. Moreover, in this study, we found that miR-199a-5p is also HIF-1α-responsive, ectopic expression of HIF-1α can greatly suppress the expression of miR-199a-5p. Down-regulation of miR-199a-5p can in turn lead to further up-regulation of its target HIF-1α, thus forming a positive feedback loop to ensure hypoxia-induced HIF-1α expression. This positive feedback loop reinforces a critical role of miR-199a-5p/HIF-1α in mediating the Warburg effect and tumor progression in HCC.
Although some studies have reported that miR-199a-5p is down-regulated in HCC and acts as a tumor suppressor [16-18], the clinical prognostic value of miR-199a-5p remains elusive. Importantly, our results demonstrated that miR-199a-5p expression was significantly associated with overall survival of HCC patients. Kaplan-Meier analysis showed that patients with low-level miR-199a-5p expression had a significant worse overall survival rate than those with high-level miR-199a-5p expression. Furthermore, multivariate Cox analysis identified that miR-199a-5p was an independent prognostic indicator for HCC patients. Thus, miR-199a-5p could be used as a potential biomarker to classify a subtype of HCC patients who have higher risk of death, thus, might receive more frequent monitoring and effective adjuvant treatment.
In summary, we demonstrated that mutual regulation of miR-199a-5p and HIF-1α is critical for controlling the Warburg effect in HCC cells. Under hypoxic conditions or within solid tumor microenvironments, miR-199a-5p is down-regulated by HIF-1α, then leads to further up-regulation of its target HIF-1α and HIF-1α downstream genes including glycolytic enzymes, thus forming a positive feedback loop to promote glycolysis. Our current study suggests a newly defined mechanism contributing to the Warburg effect and also implicates miR-199a-5p as a potential therapeutic target for HCC.
Acknowledgements
This study was supported by grants from the National Natural Science Foundation of China (81101863 and 81372571), Guangdong Department of Science & Technology Translational Medicine Center (No.2011A080300002), Sun Yat-sen University Translational Medicine 985 Project (No.90003-3283404), and Sun Yat-sen University Clinical Research 5010 Program (No. 2012010).
Competing Interests
The authors have declared that no competing interest exists.
References
1. Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144:646-674
2. Warburg O. On the origin of cancer cells. Science. 1956;123:309-314
3. Vander Heiden MG, Cantley LC, Thompson CB. Understanding the Warburg effect: the metabolic requirements of cell proliferation. Science. 2009;324:1029-1033
4. Vander Heiden MG. Targeting cancer metabolism: a therapeutic window opens. Nat Rev Drug Discov. 2011;10:671-684
5. Deberardinis RJ, Sayed N, Ditsworth D. et al. Brick by brick: metabolism and tumor cell growth. Curr Opin Genet Dev. 2008;18:54-61
6. Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116:281-297
7. He L, Hannon GJ. MicroRNAs: small RNAs with a big role in gene regulation. Nat Rev Genet. 2004;5:522-531
8. Rottiers V, Naar AM. MicroRNAs in metabolism and metabolic disorders. Nat Rev Mol Cell Biol. 2012;13:239-250
9. Gao P, Sun L, He X. et al. MicroRNAs and the Warburg Effect: new players in an old arena. Curr Gene Ther. 2012;12:285-291
10. Chan B, Manley J, Lee J. et al. The emerging roles of microRNAs in cancer metabolism. Cancer Lett. 2015;356:301-308
11. Singh PK, Mehla K, Hollingsworth MA. et al. Regulation of Aerobic Glycolysis by microRNAs in Cancer. Mol Cell Pharmacol. 2011;3:125-134
12. Chen B, Li H, Zeng X. et al. Roles of microRNA on cancer cell metabolism. J Transl Med. 2012;10:228
13. Tomasetti M, Santarelli L, Neuzil J. et al. MicroRNA regulation of cancer metabolism: role in tumour suppression. Mitochondrion. 2014;19(Pt A):29-38
14. Liu AM, Xu Z, Shek FH. et al. miR-122 targets pyruvate kinase M2 and affects metabolism of hepatocellular carcinoma. PLoS One. 2014;9:e86872
15. Jiang S, Zhang LF, Zhang HW. et al. A novel miR-155/miR-143 cascade controls glycolysis by regulating hexokinase 2 in breast cancer cells. EMBO J. 2012;31:1985-1998
16. Shen Q, Cicinnati VR, Zhang X. et al. Role of microRNA-199a-5p and discoidin domain receptor 1 in human hepatocellular carcinoma invasion. Mol Cancer. 2010;9:227
17. Duan Q, Wang X, Gong W. et al. ER stress negatively modulates the expression of the miR-199a/214 cluster to regulates tumor survival and progression in human hepatocellular cancer. PLoS One. 2012;7:e31518
18. Xu N, Zhang J, Shen C. et al. Cisplatin-induced downregulation of miR-199a-5p increases drug resistance by activating autophagy in HCC cell. Biochem Biophys Res Commun. 2012;423:826-831
19. Guo W, Qiu Z, Wang Z. et al. MiR-199a-5p is negatively associated with malignancies and regulates glycolysis and lactate production by targeting hexokinase 2 in liver cancer. Hepatology. 2015;62:1132-1144
20. Kwak PB, Iwasaki S, Tomari Y. The microRNA pathway and cancer. Cancer Sci. 2010;101:2309-2315
21. Rane S, He M, Sayed D. et al. Downregulation of miR-199a derepresses hypoxia-inducible factor-1alpha and Sirtuin 1 and recapitulates hypoxia preconditioning in cardiac myocytes. Circ Res. 2009;104:879-886
22. Wang B, Hsu SH, Frankel W. et al. Stat3-mediated activation of microRNA-23a suppresses gluconeogenesis in hepatocellular carcinoma by down-regulating glucose-6-phosphatase and peroxisome proliferator-activated receptor gamma, coactivator 1 alpha. Hepatology. 2012;56:186-197
23. Gao P, Tchernyshyov I, Chang TC. et al. c-Myc suppression of miR-23a/b enhances mitochondrial glutaminase expression and glutamine metabolism. Nature. 2009;458:762-765
24. Kim HR, Roe JS, Lee JE. et al. p53 regulates glucose metabolism by miR-34a. Biochem Biophys Res Commun. 2013;437:225-231
25. Fong MY, Zhou W, Liu L. et al. Breast-cancer-secreted miR-122 reprograms glucose metabolism in premetastatic niche to promote metastasis. Nat Cell Biol. 2015;17:183-194
26. Kroemer G, Pouyssegur J. Tumor cell metabolism: cancer's Achilles' heel. Cancer Cell. 2008;13:472-482
27. Courtnay R, Ngo DC, Malik N. et al. Cancer metabolism and the Warburg effect: the role of HIF-1 and PI3K. Mol Biol Rep. 2015;42:841-851
28. Semenza GL. HIF-1: upstream and downstream of cancer metabolism. Curr Opin Genet. Dev. 2010;20:51-56
29. Wilson WR, Hay MP. Targeting hypoxia in cancer therapy. Nat Rev Cancer. 2011;11:393-410
30. Vaupel P. Hypoxia in neoplastic tissue. Microvasc Res. 1977;13:399-408
31. Bensaad K, Harris AL. Hypoxia and metabolism in cancer. Adv Exp Med Biol. 2014;772:1-39
32. Kenneth NS, Rocha S. Regulation of gene expression by hypoxia. Biochem J. 2008;414:19-29
33. Du R, Sun W, Xia L. et al. Hypoxia-induced down-regulation of microRNA-34a promotes EMT by targeting the Notch signaling pathway in tubular epithelial cells. PLoS One. 2012;7:e30771
34. Ying Q, Liang L, Guo W. et al. Hypoxia-inducible microRNA-210 augments the metastatic potential of tumor cells by targeting vacuole membrane protein 1 in hepatocellular carcinoma. Hepatology. 2011;54:2064-2075
35. Fasanaro P, D'Alessandra Y, Di Stefano V. et al. MicroRNA-210 modulates endothelial cell response to hypoxia and inhibits the receptor tyrosine kinase ligand Ephrin-A3. J Biol Chem. 2008;283:15878-15883
36. Kulshreshtha R, Ferracin M, Wojcik SE. et al. A microRNA signature of hypoxia. Mol Cell Biol. 2007;27:1859-1867
Author contact
Corresponding authors: Dr. Binkui Li, Department of Hepatobiliary Oncology, Sun Yat-sen University Cancer Center, 651 Dongfeng Road East, Guangzhou 510060, China; E-Mail: libkorg.cn; Tel & Fax: (86) 20-87343951. Or Dr. Yunfei Yuan, Department of Hepatobiliary Oncology, Sun Yat-sen University Cancer Center, 651 Dongfeng Road East, Guangzhou 510060, China; email: yuanyfsysu.edu.cn; Tel & fax: (86) 20-87343118.

 Global reach, higher impact
Global reach, higher impact