Impact Factor ISSN: 1837-9664
J Cancer 2017; 8(18):3764-3773. doi:10.7150/jca.20900 This issue Cite
Research Paper
ITGBL1 Predicts a Poor Prognosis and Correlates EMT Phenotype in Gastric Cancer
1. Department of Oncology, Huai'an First People's Hospital, Nanjing Medical University, 6 Beijing West Road, Huai'an 223300, PR China;
2. Department of General Surgery, Renji Hospital, School of Medicine, Shanghai Jiao Tong University, 1630 Dongfang Road, Shanghai 200127, PR China;
3. State Key Laboratory of Oncogenes and Related Genes, Shanghai Cancer Institute, Renji Hospital, School of Medicine, Shanghai Jiao Tong University, 800 Dongchuan Road, Shanghai 200240, PR China.
* These authors contributed equally to this work.
Received 2017-5-6; Accepted 2017-8-14; Published 2017-10-17
Abstract
Integrin, beta-like 1 (ITGBL1), a β-integrin-related extracellular matrix protein, was found more commonly up-regulated in gastric cancer (GC) by screening and analyzing Gene Expression Omnibus (GEO) and Oncomine databases, reminding us to explore its prognostic significance in GC. In our current study, we observed that ITGBL1 expression was significantly up-regulated in GC compared with normal controls in clinical specimens. In addition, elevated ITGBL1 expression was positively correlated with patients' tumor-node-metastasis (TNM) stage and distant metastasis. Kaplan-Meier analysis indicated that high ITGBL1 expression was significantly associated with shorter survival times in GC patients. Multivariate Cox regression analysis confirmed ITGBL1 expression as an independent prognostic factor in GC. Gene set enrichment analysis (GSEA) of multiple GEO datasets revealed a close relationship between ITGBL1 expression and the KRAS/epithelial-mesenchymal transition (EMT) signaling pathway. In conclusion, these data provide evidences that ITGBL1 is a potential predictor and may be involved in cancer cell invasion and metastasis via inducing EMT, and the ITGBL1-related pathways may represent a novel therapeutic strategy for treatment of GC.
Keywords: ITGBL1, prognosis, metastasis, EMT, gastric cancer.
Introduction
Although the incidence and mortality have been declining in recent decades, gastric cancer (GC) remains the fourth most common human malignancy worldwide [1,2]. In China, GC is the second most common cancer in men and third most common in women, and is second only to lung cancer as the cause of cancer-related mortality [3]. The only possibly curative therapy for GC is still surgical resection, however, approximately 60% of patients with GC are diagnosed with locally advanced and metastatic disease, resulting in poor prognosis for lack of early detection and loss of the opportunity of curative resection. Hence, the identification of novel diagnostic markers has become a major topic to the GC research.
Integrin, beta-like 1 (ITGBL1), which was first cloned from an osteoblast cDNA library, encodes a β-integrin-related extracellular matrix protein. The ITGBL1 protein comprises a typical signal peptide and a hydrophilic 471-amino-acid domain containing ten tandem integrin EGF-like repeats [4]. Although as a member of the EGF-like protein family, ITGBL1 contains neither a transmembrane domain nor an RGD (Arg-Gly-Asp)-binding domain, suggesting that ITGBL1 performs functions distinct from those of integrin [5]. Recently, few studies have reported the functions of ITGBL1 in human diseases. In breast cancer, ITGBL1 as a Runx2 transcriptional target promotes bone metastasis by activating the TGFβ signaling pathway [6]; in non-small cell lung cancer (NSCLC), epigenetic down-regulated ITGBL1 promotes cell invasion through the Wnt/PCP signaling pathway [7]; in ovarian cancer, ITGBL1 promotes cell migration and adhesion by activating the Wnt/PCP and FAK/SRC signaling pathways [8]; in patients with chronic hepatitis B, ITGBL1 is one of a six gene signature for identifying the risk of developing cirrhosis and identified as a key regulator of fibrogenesis [9,10]. In GC, however, the role of ITGBL1 remains to be uncharacterized.
In this study, we examined the expression pattern and prognostic value of ITGBL1 in GC. We found that ITGBL1 expression was significantly up-regulated in GC tissues compared to paired non-tumor tissues, and the expression level of ITGBL1 was correlated with the tumor-node-metastasis (TNM) stage, distant metastasis and poor prognosis of GC patients. Gene Set Enrichment Analysis (GSEA) of several Gene Expression Omnibus (GEO) datasets revealed that high expression of ITGBL1 was related to the KRAS/epithelial-mesenchymal transition (EMT) signaling pathway. Taken together, these data indicate that ITGBL1 may serve as a novel prognostic marker for GC and involve in gastric cancer metastasis via inducing EMT.
Materials and Methods
Ethics statement
This research was approved by the Ethics Committee of Renji Hospital, School of Medicine, Shanghai Jiao Tong University, and written informed consent was obtained from each patient involved in this study.
GC specimens and Tissue microarray (TMA) construction
All patient data were pseudonymized before study inclusion. The inclusion and exclusion criteria were as previously described [11]. According to the inclusion and exclusion criteria, a total of 231 eligible cases were identified from east and west surgery departments of Renji Hospital between January 2004 and December 2008. Tissue microarrays were constructed as previously described [11].
Clinicopathological characteristics, including age, gender, Lauren type, tumor size, T classification, lymph node metastasis, TNM stage, early gastric cancer, and distant metastasis, were included in the records. Among them, tumor stage was determined by the American Joint Committee on Cancer (AJCC) staging system for gastric cancer; early gastric cancer was defined as “a carcinoma limited to the gastric mucosa and/or submucosa regardless of the lymph node status”.
Immunohistochemical (IHC) staining and scoring
IHC staining was performed using a two-step protocol. Briefly, after microwave antigen retrieval, tissue sections were incubated with the antibody for ITGBL1 (dilution 1:200, Abnova, US) at 4 ℃ overnight, labeled by HRP (rabbit) second antibody (Thermo Scientific, US) at room temperature for 60 min. Finally, sections were developed in DAB solution (Gene Tech, Shanghai) under microscopic observation and counterstained with hematoxylin.
All the sections were observed and photographed with a microscope (Carl Zeiss, Germany). Immunoreactivity of ITGBL1 was evaluated according to both the ratio of positive-staining cells and staining intensity. The extent of ITGBL1 staining was semiquantitatively scored as follows: 0-5% scored 0; 6-35% scored 1; 36-70% scored 2; more than 70% scored 3. Staining intensity was scored as follows: no staining scored 0, weakly staining scored 1, moderately staining scored 2 and strongly staining scored 3. The final score was designated using the extent of positive cell score × staining intensity score as follows: “-” for a score of 0-1, “+” for a score of 2-3, “++” for a score of 4-6 and “+++” for a score of > 6; low expression was defined as a total score < 4 and high expression with a total score ≥ 4. These scores were determined by two independent pathologists in a blinded manner, and mean percentage values of two scores were taken.
Western blotting
Total protein of GC tissue was extracted using a total protein extraction buffer (Beyotime, China) and the protein concentration was measured using a BCA Protein Assay Kit (Pierce Biotechnology, US), according to the manufacturer's instructions. Samples were separated by 10% SDS-PAGE gel electrophoresis and then transferred to nitrocellulose membrane (Millipore, US). Membranes were blocked with 5% (w/v) nonfat milk (BD, US) in Tris-buffered saline (TBS) and incubated with ITGBL1 antibody (dilution 1:500, Abnova, US) diluted in TBS containing 1% (w/v) bovine serum albumin at 4 ℃ overnight. Bound secondary antibodies were detected by Odessey Imaging System (LI-COR Biosciences, Lincoln, NE).
Gene Set Enrichment Analysis (GSEA)
Data used for GSEA were accessible from Gene Expression Omnibus (GEO, hhtp://www.ncbi.nlm.nih.gov/gds/) and analyzed using the software GSEA v2.2.2 (hhtp://www.broadinstitute.org/gsea). The high and low groups of clinical GC specimens were separated by the median ITGBL1 expression level. Statistical significance (false discovery rate, FDR) was set at 0.25.
Statistical analysis
All statistical analyses were done using SPSS 19.0 for windows (IBM Corporation) and GraphPad Prism 5 software (San Diego, CA). The chi-square test and Fisher's exact probability method were used to analyze the correlation between ITGBL1 expression and clinicopathological parameters in GC patients. Survival curves were evaluated by the Kaplan-Meier method and analyzed by the log-rank test. Univariate and multivariate analyses were performed to identify the factors that had a significant influence on survival based on the Cox proportional hazards regression model. P < 0.05 was considered statistically significant.
Results
The mRNA expression of ITGBL1 is increased in GC tissues
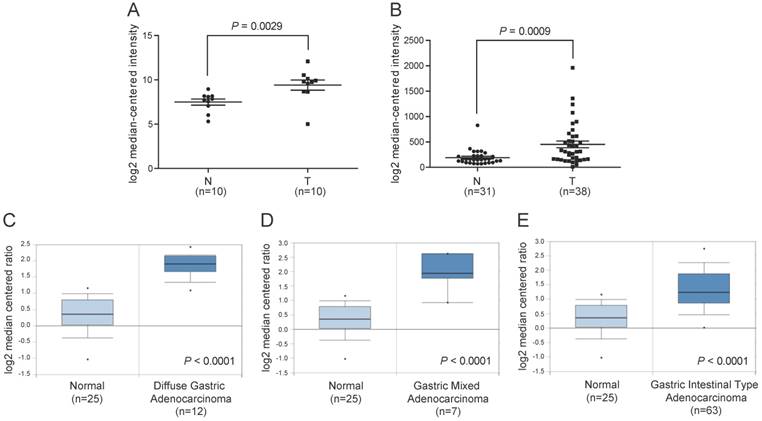
To illustrate the expression pattern of ITGBL1 in GC tissues, we first searched the mRNA expression level of ITGBL1 in two GEO datasets. The results showed that ITGBL1 expression was significantly increased in GC tissues comparing with normal adjacent gastric tissues revealed by GSE79973 [12] (Fig. 1A, P = 0.0029) and GSE13911 [13] datasets (Fig 1B, P = 0.0009). We then utilized microarray data available in the Oncomine database to analyze ITGBL1 mRNA expression in human clinical samples of GC and normal gastric tissues [14]. The results also revealed that ITGBL1 mRNA expression was markedly higher in diffuse, intestinal, and mixed gastric adenocarcinoma compared with gastric mucosa (Fig. 1C-E, all P < 0.0001).
Increased ITGBL1 expression at mRNA level in gastric cancer tissues. A. The mRNA expression of ITGBL1 was up-regulated in GC tissues (T) compared with the normal non-tumor tissues (N) revealed using the GSE79973 dataset. B. ITGBL1 expression in the normal gastric and GC tissues revealed by the GSE13911 dataset. C-E. Box and whiskers plots of Oncomine data on ITGBL1 mRNA levels in the normal gastric and GC tissues.
The protein expression of ITGBL1 is up-regulated in GC tissues
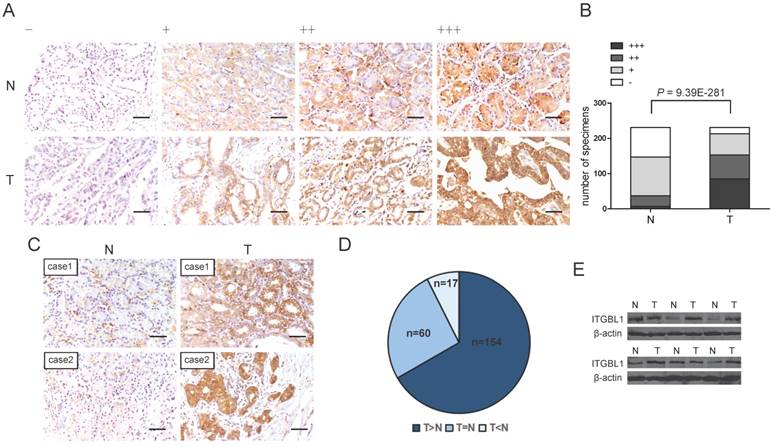
To further address the protein change of ITGBL1 in GC tissues, IHC analysis was performed in GC TMA. The representative staining of ITGBL1 expression in GC as well as paired non-tumor gastric tissues were shown in Fig. 2A, positive immunostaining for ITGBL1 was predominantly observed in the cytoplasm and extracellular matrix (ECM) of GC and normal gastric tissues. We found that ITGBL1 was significantly up-regulated in GC tissues compared with adjacent non-tumor tissues (Fig. 2B and 2C, P < 0.0001). Among 231 paired tissues, ITGBL1 expression was up-regulated in 154 cases compared with that of the paired non-tumor gastric tissues (Fig. 2D). Furthermore, we validated the protein expression of ITGBL1 in another six pairs of resected representative specimens (tumor tissues and paired adjacent non-tumor tissues) from GC patients using western blotting analysis. Consistently, an increase of ITGBL1 expression was observed in five GC tissues compared with the paired adjacent non-tumor tissues (Fig. 2E).
Increased ITGBL1 expression at protein level in gastric cancer tissues. A. Representative photographs of the ITGBL1 immunoreactivity in non-tumor gastric tissues (N) and GC tissues (T) (scale bar: 10 μm). B. Comparisons of ITGBL1 expression in TMA revealed by IHC analysis in paired non-tumor and GC tissues. C-D. ITGBL1expression was up-regulated in 154 GC tissues (T) compared with the paired adjacent non-tumor tissues (N) (scale bar: 10 μm). E. ITGBL1 expression in paired tissues from six GC patients analyzed by western blotting.
Correlation of ITGBL1 expression with clinicopathological parameters in patients with GC
To determine the clinical significance of ITGBL1 expression in GC, we used Chi-square test to assess the correlation between ITGBL1 protein expression and corresponding patients' clinicopathologic parameters shown in Table 1. The results showed that ITGBL1 expression in GC tissues was significantly correlated with TNM stage (P = 0.030), early gastric cancer (P = 0.007) and distant metastasis (P = 0.013). While there was no significant correlation with age, gender, Lauren type, tumor size, T classification and lymph node metastasis (Table 1). ITGBL1 expression was increased in advanced gastric cancer compared with that in early gastric cancer and was positively associated with TNM stage and metastasis, indicating that ITGBL1 may be implicated in the progression of GC.
Correlations between ITGBL1 expression and clinicopathologic parameters in patients with GC
| Clinicopathological parameter | Total 231 | Expression of ITGBL1 | p value | |
|---|---|---|---|---|
| Low (n=84, %) | High (n=147, %) | |||
| Age (years) | ||||
| < 60 | 100 | 38 (38.0) | 62 (62.0) | 0.680 |
| ≥ 60 | 131 | 46 (35.1) | 85 (64.9) | |
| Gender | ||||
| Male | 148 | 58 (39.2) | 90 (60.8) | 0.256 |
| Female | 83 | 26 (31.3) | 57 (68.7) | |
| Lauren type | ||||
| Intestinal type | 129 | 46 (35.7) | 83 (64.3) | 0.891 |
| Diffuse type | 102 | 38 (37.3) | 64 (62.7) | |
| Tumor size | ||||
| ≤ 5 cm | 127 | 53 (41.7) | 74 (58.3) | 0.074 |
| >5cm | 104 | 31 (29.8) | 73 (70.2) | |
| T classification | ||||
| T1, 2 | 84 | 37(44.0) | 47(56.0) | 0.239 |
| T3, 4 | 147 | 47 (32.0) | 100 (68.0) | |
| Lymph node metastasis | ||||
| Absent | 99 | 41 (41.4) | 58 (58.6) | 0.245 |
| Present | 132 | 43 (32.6) | 89 (67.4) | |
| TNM stage | ||||
| I | 63 | 29 (46.0) | 34 (54.0) | 0.030 |
| II | 55 | 18 (32.7) | 37 (67.3) | |
| III | 93 | 35 (37.6) | 58 (62.4) | |
| IV | 20 | 2 (10.0) | 18 (90.0) | |
| Early gastric cancer | ||||
| No | 183 | 58 (31.7) | 125 (68.3) | 0.007 |
| Yes | 48 | 26 (54.2) | 22 (45.8) | |
| Distant metastasis | ||||
| Absent | 211 | 82 (38.9) | 129 (61.1) | 0.013 |
| Present | 20 | 2 (10.0) | 18 (90.0) | |
aThe bold number represents the p-values with significant differences.
bP value was calculated by χ2 test or Fisher's exact test.
Up-regulated ITGBL1 predicts a poor prognosis of GC patients
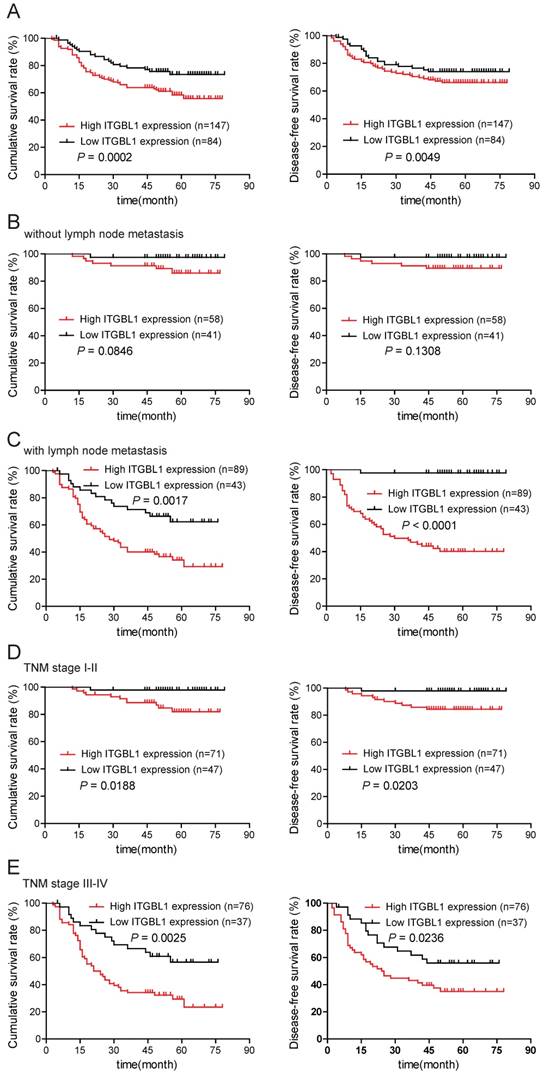
To evaluate the prognostic significance of ITGBL1 in GC patients, we used Kaplan-Meier analysis and log-rank test to analyze the correlation between ITGBL1 expression and corresponding clinical follow-up information. As shown in Fig. 3A, high ITGBL1 expression was remarkably associated with decreased overall survival (OS) (P = 0.0002) and disease-free survival (DFS) (P = 0.0049). In addition, we determined the correlation between ITGBL1 expression and OS or DFS in GC patients in the present or absent of lymphatic metastasis and in early or advanced TNM stage. Kaplan-Meier analyses showed that both OS and DFS were shorter in GC patients with higher ITGBL1 expression in GC patients with lymphatic metastasis (Fig. 3C, all P < 0.05), but neither in GC patients without lymphatic metastasis (Fig. 3B, all P > 0.05); both OS and DFS were shorter in GC patients with higher ITGBL1 expression regardless of TNM stage (Fig. 3D and 3E, all P < 0.05). Furthermore, univariate and multivariate analyses were conducted to identify the risk factors correlated with the prognosis of GC patients. Univariate analysis showed that ITGBL1 expression, Lauren type, tumor size, T classification, lymph node metastasis, TNM stage, and distant metastasis were significantly associated with overall survival (Table 2). Meanwhile, multivariate analysis using the Cox proportional hazards model revealed that ITGBL1 expression was a significant independent prognostic factor for GC patients (P = 0.011) in addition to distant metastasis (P < 0.001) (Table 2). Taken together, these data above suggest that up-regulated ITGBL1 predicts a poor prognosis, and might contribute to tumor progression in GC.
ITGBL1 expression is correlated with survival rate. A. Overall survival and disease-free survival analyses of GC patients with different ITGBL1 protein expression. B-C. Comparison of overall and disease-free survival in patients without (B) or with (C) lymph node metastasis was conducted based on ITGBL1 expression. D-E. Comparison of overall survival and disease-free survival between lower ITGBL1 expression group and higher ITGBL1 expression group in early TNM stage cohort (I-II) (D) and in advanced TNM stage cohort (III-IV) (E). P value was calculated by log-rank test.
Univariate and multivariate analysis of prognostic parameters for survival in patients with GC
| Prognostic parameter | Univariate analysis | Multivariate analysis | |||
|---|---|---|---|---|---|
| HR (95% CI) | p value | HR (95% CI) | p value | ||
| Expression of ITGBL1 (low vs. high) | 2.728 (1.576-4.724) | <0.001 | 2.094(1.184-3.703) | 0.011 | |
| Age (< 60 vs. ≥ 60) | 1.013 (0.998-1.028) | 0.083 | - | - | |
| Gender (male vs. female) | 0.918 (0.649-1.299) | 0.631 | - | - | |
| Lauren type (intestinal vs. diffuse) | 2.257 (1.625-3.134) | <0.001 | 1.340 (0.850-2.112) | 0.208 | |
| Tumor size (≤ 5 cm vs. > 5 cm) | 3.665 (2.576-5.215) | <0.001 | 1.314 0.796-2.168) | 0.295 | |
| T classification (T1, 2 vs. T3, 4) | 8.497 (4.703-15.353) | <0.001 | 1.379 (0.395-4.819) | 0.614 | |
| Lymph node metastasis (absent vs. present) | 7.771 (4.737-12.750) | <0.001 | 3.323 (0.916-12.056) | 0.068 | |
| TNM stage (I - II vs. III - IV) | 9.135 (5.879-14.193) | <0.001 | 1.562 (0.371-6.575) | 0.543 | |
| Early gastric cancer (no vs. yes) | 14.973 (4.771-46.992) | <0.001 | 4.491 (0.510-39.517) | 0.176 | |
| Distant metastasis (absent vs. present) | 9.667 (6.331-14.761) | <0.001 | 5.229 (2.861-9.557) | <0.001 |
aHR: Hazard ratio; CI: Confidence interval
bThe bold number represents the p value with significant differences
Correlation between the mRNA expression of ITGBL1 and the KRSA/EMT signaling pathway in human GC specimens
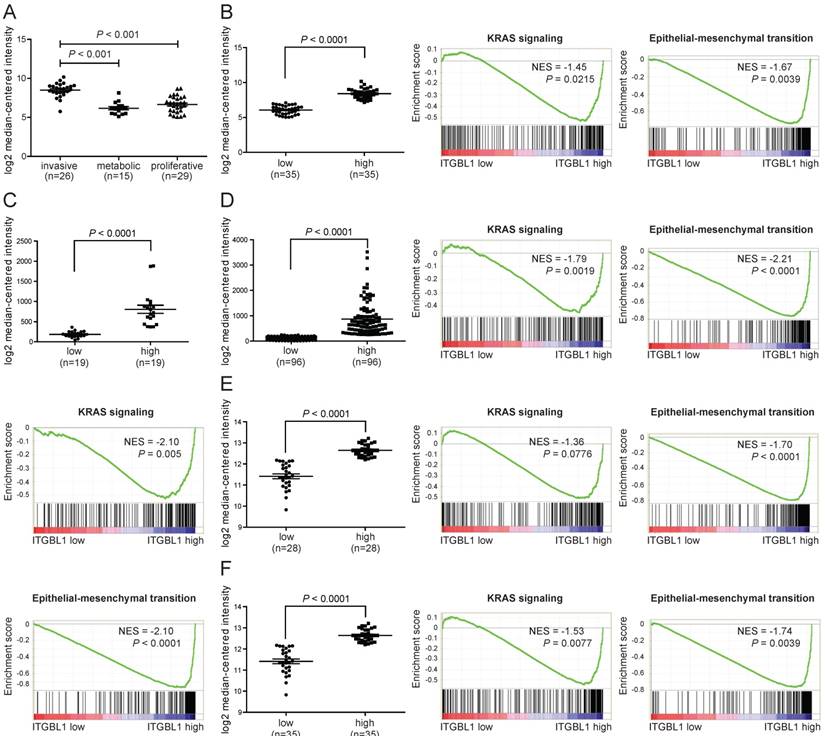
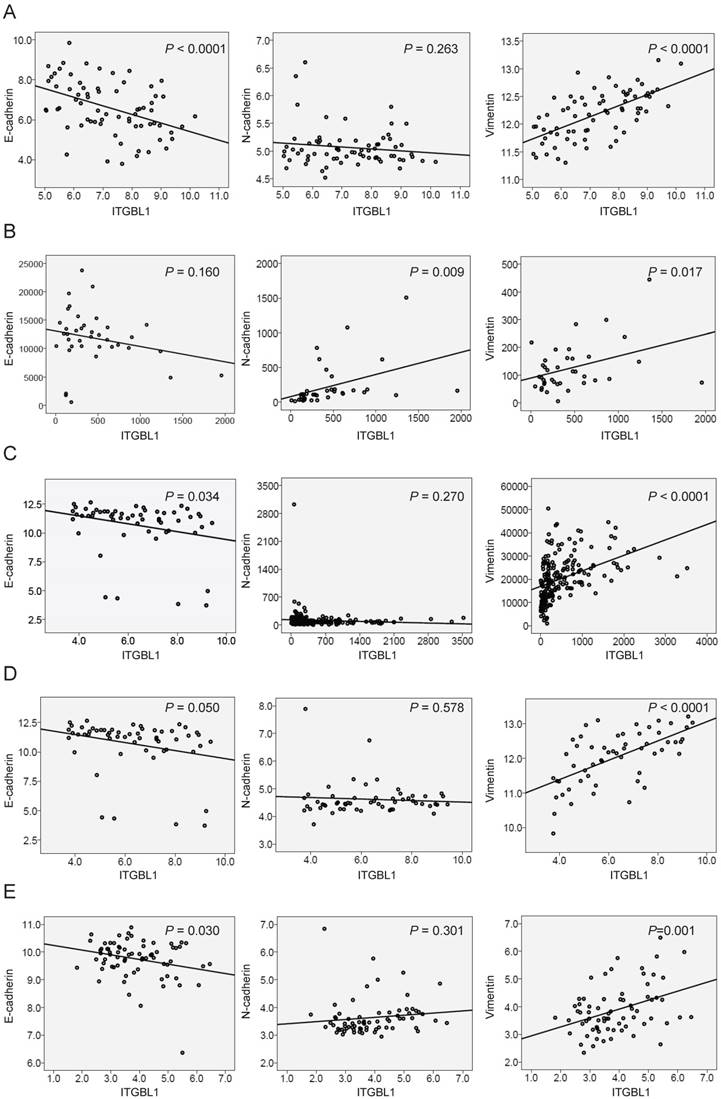
When searching the mRNA expression level of ITGBL1 in GSE35809 dataset [15], we found that ITGBL1 expression was significantly increased in GC of invasive subtype compared with that in GC of metabolic and proliferative subtypes (Fig. 4A, P < 0.001), suggesting that ITGBL1 may participate in GC invasion. Moreover, GSEA comparing GC tumors with high versus low expression of ITGBL1 using five independent GEO datasets showed a strong correlation between ITGBL1 expression and the KRAS/EMT signaling pathway (Fig. 4B-F) revealed using the GSE35809, GSE13911, GSE15459 [16], GSE34942 [16], and GSE57303 [17] datasets. Finally, based on the results of GSEA, we analyzed the correlation between ITGBL1 expression and EMT markers' expression. Consistent with the results of GSEA, ITGBL1 expression was negatively associated with the expression of epithelial marker E-cadherin, and positively with the expression of mesenchymal marker vimentin rather than N-cadherin (Fig. 5A-E), as the expression of EMT markers often depends on the cell type and initiating signaling pathway. Taken together, these data suggested that ITGBL1 may promote GC invasion and metastasis via activating KRAS signaling to induce EMT progression.
ITGBL1 expression is positively correlated with a KRAS/EMT gene program in GC. A. The mRNA expression of ITGBL1 was increased in invasive subtype of GC compared with the metabolic and proliferative subtypes of GC revealed using the GSE35809 dataset. B-F. Performance of GSEA based on GEO datasets. Patients were divided into low and high groups by the median ITGBL1 expression. High expression of ITGBL1 correlated with the KRAS/EMT signaling pathway revealed using the GSE35809 (B), and GSE13911 (C), GSE15459 (D), GSE34942 (E) and GSE57303 (F) datasets. NES, normalized enrichment score.
Correlation of ITGBL1 expression with the level of EMT markers, including E-cadherin, N-cadherin and vimentin (expressed as the log2 median-centered intensity) revealed using the GSE35809 (A), and GSE13911 (B), GSE15459 (C), GSE34942 (D) and GSE57303 (E) datasets.
Discussion
To date, there are few conflicting reports on ITGBL1 roles in the field of tumor biology. It has been reported that ITGBL1 promotes bone metastasis by activating the TGFβ signaling pathway in breast cancer [6] and is up-regulated and promotes cell migration and adhesion through the Wnt/PCP and FAK/SRC signaling pathways in ovarian cancer [8]. However, another study reported that ITGBL1 is more commonly down-regulated in NSCLC tissues, and epigenetic down-regulated ITGBL1 promotes cancer cell invasion through the Wnt/PCP signaling pathway [7]. In current study, ITGBL1 expression and its association with clinicopathological features were investigated across a cohort of Chinese GC patients.
We demonstrated that ITGBL1 expression was commonly up-regulated in GC patients at protein level. These findings were supported at the mRNA level by non-overlapping data from GEO and Oncomine databases, which highlighted the same trends in GC. In our study, however, ITGBL1 expression was also observed in normal gastric tissues, which might be explained by that these normal gastric tissues were mainly non-tumor tissues derived from patients with GC and the tumor microenvironment of GC was likely to turn on the expression of ITGBL1 in adjacent normal tissues. In addition, up-regulated ITGBL1 expression was significantly correlated with TNM stage, early gastric cancer and distant metastasis of GC. The findings remind us that ITGBL1 may play important roles in the progression of GC.
Furthermore, survival analyses in our study revealed that GC patients with high ITGBL1 expression levels had a poorer prognosis than those with low expression levels. Multivariate analysis demonstrated that increased ITGBL1 expression and distant metastasis were independent risk factors in the prognosis of GC patients. These results suggested that ITGBL1 may represent a novel prognostic marker for GC patients.
Previous studies have indicated that ITGBL1 is involved in cancer metastasis [6-8]. When screening the GSE35809 database, we found that ITGBL1 expression was significantly increased in invasive subtype of GC than that of metabolic and proliferative subtypes, suggesting that ITGBL1 might play an important role in gastric cancer invasion and metastasis. Invasion and metastasis, the major causes of GC-related relapse and death, greatly impede the treatment efficiency [18]. The precise molecular mechanisms through which ITGBL1 impacts on GC invasion and metastasis have yet to be elucidated, GSEA of multiple independent datasets revealed that human GC specimens with high expression of ITGBL1 had elevated expression of EMT genes.
Abundant evidences have revealed the importance of EMT, during which epithelial cells lose their junctions and cellular polarity, reorganize their cytoskeleton, finally acquire migratory and invasive capabilities [19]. EMT is critical for multiple pathophysiological processes, especially tumorigenesis and progression [20]. Aberrant EMT activation provides cancer cells with enhanced migratory and invasive capabilities and promotes tumor progression [21]. In tumor cells, an essential EMT step is the down-regulation of E-cadherin to reinforce the destabilization of cell junctions [22]. The E- to N-cadherin switch has critical functions in cancer progression for being essential for enhanced motility and migration [23]. Vimentin expression cause EMT phenotype changes in human GC cells [24]. To further confirm the results of GSEA, correlation of ITGBL1 expression with EMT markers' expression was analyzed. ITGBL1 expression was found to be closely associated the expression of EMT markers E-cadherin and vimentin.
Recent evidence indicates that EMT is a key GC progression driver, and plays a fundamental role during early steps of GC invasion, metastasis and relapse [25]. Many studies have shown that a variety of signaling pathways are involved in the EMT process in tumor cells. Recently, it has been reported that the activation of the PI3K/AKT axis leads to EMT through various mechanisms and has emerged as a central feature of EMT in GC [26-29]. In addition, MEK/ERK pathway has been shown to induce EMT to promote GC migration, invasion and metastasis [29-32]. Above all, PI3K/AKT and MEK/ERK signaling pathways can promote GC metastasis by inducing EMT. Among KRAS-related downstream pathways are mainly PI3K-Akt and MAPK/ERK signaling pathways, the results of GSEA analysis revealed that ITGBL1 expression had a positive correlation with KRAS signaling pathway in human GC specimens, which plays vital roles in EMT process of GC, suggesting that ITGBL1 may induce EMT by activating KRAS-related signaling pathways.
Investigating possible mechanisms modulating malignant cell EMT and discovering novel EMT regulators will further elucidate GC biology, and may provide novel biomarkers and efficient therapeutic targets for GC [24]. EMT is induced by diverse extracellular stimuli [32], and in GC stroma, there are various cues which can influence EMT. We hypothesize that ITGBL1, as an extracellular matrix protein, may interact with cell membrane receptors, thus activating KRAS-related pathways, such as PI3K-Akt and MAPK/ERK signaling pathways, which affect the activity and abundance of transcription factors that regulate the expression of EMT markers, finally resulting in EMT process to promote GC invasion and metastasis. The hypothesis needs to be further confirmed by in vitro and in vivo experiments; however, these data provide a clue for studying the mechanism of GC metastasis.
In conclusion, for the first time to our knowledge, we showed that ITGBL1 was up-regulated in GC by immunohistochemistry, and described ITGBL1 as a crucial factor in the clinical outcomes during human GC. Furthermore, based on bioinformatic analysis, we found that ITGBL1 may induce EMT via activating KRSA-related signaling pathways to promote cell invasion and metastasis of GC. These data provide a further insight into the pathogenesis of GC and provide substantial new evidence that ITGBL1 may serve as a novel prognostic marker and therapeutic target for GC.
Abbreviations
DFS: disease-free survival; ECM: extracellular matrix; EMT: epithelial-mesenchymal transition; GC: gastric cancer; GEO: Gene Expression Omnibus; GSEA: gene set enrichment analysis; IHC: immunohistochemical; ITGBL1: integrin, beta-like 1; NSCLC: non-small cell lung cancer; OS: overall survival; TBS: Tris-buffered saline; TMA: tissue microarray; TNM: tumor-node-metastasis.
Acknowledgements
We would like to thank the stuff of the tumor microenvironment research group of State Key Laboratory of Oncogenes and Related Genes for assistance with this study.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Siegel R, Naishadham D, Jemal A. Cancer statistics, 2012. CA Cancer J Clin. 2012;62:10-29
2. Jemal A, Bray F, Center MM. et al. Global cancer statistics. CA Cancer J Clin. 2011;61:69-90
3. Chen W, Zheng R, Baade PD. et al. Cancer Statistics in China, 2015. CA Cancer J Clin. 2016;66(2):115-32
4. Berg RW, Leung E, Gough S. et al. Cloning and characterization of a novel beta integrin-related cDNA coding for the protein TIED ("ten beta integrin EGF-like repeat domains") that maps to chromosome band 13q33: A divergent stand-alone integrin stalk structure. Genomics. 1999;56(2):169-78
5. Takagi J, Beglova N, Yalamanchili P. et al. Definition of EGF-like, closely interacting modules that bear activation epitopes in integrin beta subunits. Proc Natl Acad Sci USA. 2001;98:11175-80
6. Li XQ, Du X, Li DM. et al. ITGBL1 Is a Runx2 Transcriptional Target and Promotes Breast Cancer Bone Metastasis by Activating the TGFβ Signaling Pathway. Cancer Res. 2015;75(16):3302-13
7. Gan X, Liu Z, Tong B. et al. Epigenetic downregulated ITGBL1 promotes non-small cell lung cancer cell invasion through Wnt/PCP signaling. Tumor Biol. 2016;37(2):1663-9
8. Sun L, Wang D, Li X. et al. Extracellular matrix protein ITGBL1 promotes ovarian cancer cell migration and adhesion through Wnt/PCP signaling and FAK/SRC pathway. Biomed Pharmacother. 2016;81:145-51
9. Xu MY, Qu Y, Li Z. et al. A 6 gene signature identifies the risk of developing cirrhosis in patients with chronic hepatitis B. Front Biosci (Landmark Ed). 2016;21:479-86
10. Wang M, Gong Q, Zhang J. et al. Characterization of gene expression profiles in HBV-related liver fibrosis patients and identification of ITGBL1 as a key regulator of fibrogenesis. Sci Rep. 2017;7:43446
11. Li RK, Zhao WY, Fang F. et al. Lysyl oxidase-like 4 (LOXL4) promotes proliferation and metastasis of gastric cancer via FAK/Src pathway. J Cancer Res Clin Oncol. 2015;141(2):269-81
12. He J, Jin Y, Chen Y. et al. Downregulation of ALDOB is associated with poor prognosis of patients with gastric cancer. Onco Targets Ther. 2016;9:6099-109
13. D'Errico M, de Rinaldis E, Blasi MF. et al. Genome-wide expression profile of sporadic gastric cancers with microsatellite instability. Eur J Cancer. 2009;45(3):461-9
14. Chen X, Leung SY, Yuen ST. et al. Variation in gene expression patterns in human gastric cancers. Mol Biol Cell. 2003;14(8):3208-15
15. Lei Z, Tan IB, Das K. et al. Identification of molecular subtypes of gastric cancer with different responses to PI3-kinase inhibitors and 5-fluorouracil. Gastroenterology. 2013;145(3):554-65
16. Ooi CH, Ivanova T, Wu J. et al. Oncogenic pathway combinations predict clinical prognosis in gastric cancer. PLoS Genet. 2009;5(10):e1000676
17. Qian Z, Zhu G, Tang L. et al. Whole genome gene copy number profiling of gastric cancer identifies PAK1 and KRAS gene amplification as therapy targets. Genes Chromosomes Cancer. 2014;53(11):883-94
18. Chaffer CL, Weinberg RA. A perspective on cancer cell metastasis. Science. 2011;331:1559-64
19. Thiery JP, Sleeman JP. Complex networks orchestrate epithelial-mesenchymal transitions. Nat Rev Mol Cell Biol. 2006;7:131-42
20. De Craene B, Berx G. Regulatory networks defining EMT during cancer initiation and progression. Nat Rev Cancer. 2013;13:97-110
21. Harris TJ, Tepass U. Adherens junctions: from molecules to morphogenesis. Nat Rev Mol Cell Biol. 2010;11:502-14
22. Christofori G. New signals from the invasive front. Nature. 2006;441:444-50
23. Watanabe T, Takahashi A, Suzuki K. et al. Epithelial-mesenchymal transition in human gastric cancer cell lines induced by TNF alpha-inducing protein of Helicobacter pylori. Int J Cancer. 2014;134:2373-82
24. Lei Huang, Ruo-Lin Wu, A-Man Xu. Epithelial-mesenchymal transition in gastric cancer. Am J Transl Res. 2015;7(11):2141-58
25. Matsuoka T, Yashiro M, Nishioka N. et al. PI3K/Akt signalling is required for the attachment and spreading, and growth in vivo of metastatic scirrhous gastric carcinoma. Br J Cancer. 2012;106:1535-42
26. Cantley LC. The phosphoinositide 3-kinase pathway. Science. 2002;296:1655-57
27. Luo J, Manning BD, Cantley LC. Targeting the PI3K-Akt pathway in human cancer: rationale and promise. Cancer Cell. 2003;4:257-62
28. Cui Y, Wang Y, Li H. et al. Asparaginyl endopeptidase promotes the invasion and metastasis of gastric cancer through modulating epithelial-to-mesenchymal transition and analysis of their phosphorylation signaling pathways. Oncotarget. 2016;7(23):34356-70
29. Liao A, Wang W, Sun D. et al. Bone morphogenetic protein 2 mediates epithelial-mesenchymal transition via AKT and ERK signaling pathways in gastric cancer. Tumor Biol. 2015;36:2773-8
30. Lee WS, Kim N, Park YR. et al. Myeloid cell leukemia-1 promotes epithelial-mesenchymal transition of human gastric cancer cells. Oncol Rep. 2015;34(2):1011-6
31. Liang Z, Wu R, Xie W. et al. Curcumin Suppresses MAPK Pathways to Reverse Tobacco Smoke-induced Gastric Epithelial-Mesenchymal Transition in Mice. Phytother Res. 2015;29(10):1665-71
32. Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144:646-74
Author contact
Corresponding authors: Xiaofei Chen, M.D. Department of Oncology, Huai'an First People's Hospital, Nanjing Medical University, Huai'an 223300, PR China. Phone: +86-517-84922412; Fax: +86-517-84922412; email: xf_chen1962com Or: Zhigang Zhang, Ph.D. State Key Laboratory of Oncogenes and Related Genes, Shanghai Cancer Institute, Renji Hospital, Shanghai Jiao Tong University School of Medicine, 800 Dongchuan Road, Shanghai 200240, PR China. Phone: +86-21-34206763; Fax: +86-21-34206022; E-mail: zhg_zhang1973com

 Global reach, higher impact
Global reach, higher impact