Impact Factor ISSN: 1837-9664
J Cancer 2018; 9(19):3626-3633. doi:10.7150/jca.26790 This issue Cite
Research Paper
Downregulation of Activin A Receptor Type 2A Is Associated with Metastatic Potential and Poor Prognosis of Colon Cancer
1. Department of Gastrointestinal Surgical Oncology, Fujian Cancer Hospital & Fujian Medical University Cancer Hospital, Fuzhou 350014, China
2. Department of Pathology, Fujian Cancer Hospital & Fujian Medical University Cancer Hospital, Fuzhou 350014, China
3. Departments of CyberKnife, Huashan Hospital, Fudan University, Shanghai 200032, China
4. Department of Clinical Laboratory, Fujian Cancer Hospital & Fujian Medical University Cancer Hospital, Fuzhou 350014, China
5. Department of Colorectal Surgery, Fudan University Shanghai Cancer Center; Department of Oncology, Shanghai Medical College, Fudan University, Shanghai 200032, China
6. Fujian Provincial Key Laboratory of Tumor Biotherapy, Fuzhou 350014, China
7. Fujian Provincial Key Laboratory of Translational Cancer Medicine, Fuzhou 350014, China
Received 2018-4-20; Accepted 2018-7-17; Published 2018-9-8
Abstract
Aims: Activin A receptor type 2A (ACVR2A) is a membrane receptor in the transforming growth factor- beta (TGF-β signaling pathway, which is involved in the regulation of cell proliferation, migration, and apoptosis. The aim of this study was to examine the expression profiles and biological functions of ACVR2A in colon cancer.
Methods: ACVR2A expression was investigated using the GSE39582 database and two validation cohorts. An in vitro study of cell proliferation and migration of human colon cell lines was also performed.
Results: In the GSE39582 database (n= 497), expression of ACVR2A mRNA was identified as a prognostic factor by linear regression analysis. In one validation cohort of 15 patients with stage IV cancer, the mRNA expression of ACVR2A was significantly reduced in metastatic lesions and primary tumors compared with adjacent normal controls (P = 0.001). In another validation cohort of tissue microarray (TMA) consisting of 193 cases, reduced ACVR2A protein expression correlated with advanced N stage (P = 0.001) and positive lymphovascular invasion (P = 0.005). Strong correlations between low ACVR2A mRNA or protein expression and worse survival were also observed in the GSE39582 database and the TMA validation cohort (all P < 0.05). Moreover, our in vitro studies showed a remarkable increase in cell migration in ACVR2A knockdown cells.
Conclusions: Our findings indicate that loss of ACVR2A has an important role in cancer progression and distant metastasis and may serve as a prognostic marker in patients with colon cancer.
Keywords: Activin A Receptor Type 2A (ACVR2A), Colon Cancer, Tissue Microarray, Liver Metastasis, Cell Proliferation, Cell Migration
Introduction
Colon cancer is a common malignancy and one of the leading causes of tumor-related death with an estimated 140,250 new cases and 50,630 deaths in the United States in 2018[1]. Modern surgery-based multidisciplinary team (MDT) strategies are now the standard of care for colon cancer and have improved the 5-year overall survival (OS) rate to approximately 60% [2]. Patient survival largely depends on disease stage at diagnosis. However, the challenges of considerable prognostic heterogeneity within each tumor stage need to be faced, as tumors at the same pathologic stage can produce considerably different clinical outcomes [3]. Therefore, in-depth understanding of the molecular mechanisms of tumorigenesis is needed to identify potential biomarkers that may serve as new treatment targets.
ACVR2A (activin A receptor type 2A) belongs to a receptor family that mediates the functions of activins, members of the transforming growth factor-beta (TGF-β) family of ligands with multiple biological functions [4-6]. The TGF-β signaling pathway is involved in the regulation of cell proliferation, differentiation, migration, and apoptosis of colon cancer [7-9]. The mutation rate of ACVR2A is approximately 60%, making it the most frequently mutated gene in hypermutated colon cancer [10]. However, the expression profiles of ACVR2A and its biological function in colon cancer tumorigenesis and progression are largely unknown.
In the present study, we first investigated the prognostic value of ACVR2A mRNA using the GSE39582 database, and then validated the results in a cohort of patients with metastatic colon cancer and in a tissue microarray (TMA) cohort. In addition, we examined the in vitro effects of ACVR2A expression on the proliferation and migration of human colon cell lines.
Methods
Patients in the GSE39582 database
Gene expression (RNA-Seq) data and corresponding clinical data of colon cancer patients were retrieved from the GSE39582 database (https://www.ncbi.nlm.nih.gov/geo/). Patients included in the present study were all pathologically diagnosed with colon adenocarcinoma, did not undergo chemotherapy or radiotherapy before surgery, and had intact survival information.
Tissue samples
Fifteen patients with stage IV colon cancer who underwent palliative surgery were included as one of the validation cohorts. Paired tissue fragments of the primary colon cancer, adjacent normal colonic tissue, and liver metastases were obtained after surgery, immediately stored in RNA-later, and kept at -80°C for further study of ACVR2A mRNA expression. Additionally, to validate the clinical significance of ACVR2A protein expression we obtained paired colon cancer specimens and corresponding adjacent normal colonic tissues from 193 patients. Tumor tissues were pathologically verified as adenocarcinoma and noncancerous tissues were confirmed as negative controls. These samples were constructed as a TMA for immunohistochemical (IHC) staining.
The study patients were treated at Fujian Cancer Hospital between January 2008 and December 2013. Use of clinical samples was approved by the Human Ethics Review Committee of the Fujian Cancer Hospital & Fujian Medical University Cancer Hospital, China. Written informed consent was obtained from all patients included in the study. All patients were restaged and pathologically graded according to the TNM staging system for colon cancer of the Union for International Cancer Control/American Joint Committee on Cancer (UICC/AJCC, 7th edition).
Colon cancer cell lines and culture
The human colon cancer cell lines (RKO and HCT116) were originally purchased from the American Type Culture Collection (Manassas, VA, USA), and cultured in DMEM medium supplemented with 10% FBS (Gibco, Life Technology, Austria) and 1% penicillin/streptomycin in a humidified 5% (v/v) atmosphere of CO2 at 37°C.
Quantitative reverse transcription polymerase chain reaction (qRT-PCR)
As described previously[11], total RNA was extracted from tissue or cell samples using TRIzol Reagent (Invitrogen, Life Technologies, USA), and cDNA was synthesized from 2 μg of RNA using the PrimeScript™ RT Master Mix Kit (RR036A, Takara, Japan) according to the manufacturers' instructions. qRT-PCR was carried out using Power SYBR® Green PCR Master Mix (Invitrogen, USA) on the 7900 real-time PCR system (ABI, Life Technology, USA). GAPDH was used as a loading control for each specific gene. The sequences for sense (S) and antisense (AS) primers were as follows: human-ACVR2A-S, 5′-GTTTGCCGTCTTTCTTATCTCCT-3′, human-ACVR2A-AS, 5′-GTCACCATAACACGGTTCAACA-3′, human-GAPDH-S, 5′-GCAAATTCCATGGCACCGT-3′, human-GAPDG-AS, 5′-TCGCCCCACTTGATTTTGG-3′. Relative quantitation (RQ) of the gene expression was applied to analyze the relative changes.
TMA construction and IHC analysis
TMA construction and IHC staining were performed as described previously [12]. Antibody specific for ACVR2A (anti-human goat antibody; #ab10595, Abcam, UK) was used at a dilution of 1:100 and was replaced with phosphate buffered saline (PBS) in a negative control. Every section was evaluated and scored independently by two pathologists using a semi-quantitative scoring system [12, 13]. The percentage of positively stained cells was scored as 0 (<5%), 1 (5%-25%), 2 (26%-50%), 3 (51%-75%), or 4 (>75%). Intensity was scored as 0 (negative), 1 (weak), 2 (moderate), or 3 (strong). We multiplied the percentage score by the staining intensity score to generate the immunoreactivity score (IS). High expression of ACVR2A was defined as detectable immunoreactivity with IS≥ 4.
Western blot analysis
For western blotting, proteins were isolated by lysing cells in ice-cold radio immunoprecipitation (RIPA) buffer containing protease and phosphatase inhibitors (Roche, USA). Protein was quantified using the BCA protein assay kit (Pierce, Thermo Scientific, USA) and 50 µg of total protein was resolved using 8%-12% Bis-Tris SDS-PAGE gels and transferred to PVDF membranes. Membranes were probed with primary antibodies. Following incubation with horseradish peroxidase-conjugated secondary species-specific antibodies, immunoreactive proteins were detected using ECL (Pierce, Thermo Scientific, USA) and a BioImaging System (UVP, LLC, USA).
Stable transfection of the cell lines
Biologically active short hairpin RNAs (shRNAs) were generated using the lentiviral expression vector pLKO.1-puro. The shRNA target sequence for human ACVR2A was 5'-GGTGTTGGAGGGTGCTATAAA-3'. PLKO.1-scramble shRNA with limited homology to any known sequences in the human was used as a negative control. HCT116 and RKO cells were transfected with the pLKO.1-shACVR2A expression vector or pLKO.1-scramble. Stably transfected cells were isolated using Puromycin selection to obtain stable ACVR2A knockdown cells.
Cell proliferation and migration assays
Cell proliferation reagent kit (CCK-8, Dojindo, Japan) was used to assess cell proliferation. Transfected cells were plated in each well of a 96-well plate and assessed every 24 h according to the manufacturer's instructions. The cell viability of different groups at each measurement time point was compared.
The migration ability of HCT116 and RKO cells after transfection was measured by Transwell assay. Approximately 105 cells were seeded on the upper chamber of the Transwell (8.0 μm Pore Polycarbonate Membrane Insert) with 200 μl serum-free growth medium. Complete medium containing 10% FBS was added to the lower chamber as a chemoattractant. After incubation at 37°C for 48 h, non-migratory cells on the upper surface of the upper chamber were lightly removed using cotton swabs, and cells that migrated to the bottom of the membrane were fixed and stained. The number of invaded cells was counted under a light microscope. To minimize the bias, cells in five randomly selected fields with 200× magnification were counted and the average number was calculated.
Statistical analysis
Patients were followed up regularly according to the National Comprehensive Cancer Network (NCCN) Guidelines for Colon Cancer. The last follow-up was set on June 30, 2017. Patients who were alive at the last follow-up were censored for analysis. Death from colon cancer or any other cause at last follow-up was defined as the clinical endpoint. The overall survival (OS) was defined as the time from treatment to death from any cause, and the relapse-free survival (RFS) was defined as the time from treatment to the first recurrence or death. Patients who died within 1 month after diagnosis were excluded from this study. Each experiment was performed three times and each sample was tested in triplicate. Two-tailed χ2 test was used to evaluate differences in ACVR2A expression according to the patients' clinicopathological features. The survival curves were estimated by Kaplan-Meier plots, and P values were calculated by log-rank test. Multivariate Cox regressions analysis was applied to estimate the individual hazard ratio (HR) for OS and RFS. All P values were two-sided, and P < 0.05 was considered statistically significant. All statistical calculations were performed using SPSS 22.0 (IBM Corp., USA).
Results
Clinicopathological characteristics
The median follow-up time for all patients included in this study was 68 months, ranging from 42 to 115 months. A total of 497 eligible patients with colon cancer from GSE39582 were included in the study. All patients were diagnosed as adenocarcinoma and underwent surgical resection. The study population contained 36 (7.2%), 260 (52.3%), and 201 (40.4%) patients diagnosed with stage I, II, and III disease, respectively. During the period of follow-up, 140 (28.2%) patients were confirmed with tumor relapse.
The validation cohort of 15 stage IV patients consisted of 11 men and 4 women. Four patients had only one metastasis in the liver and 11 had more than one metastasis in the liver or extrahepatic metastases. All patients underwent palliative resection of the primary colon cancer with concurrent hepatic metastasectomy (5 cases) or biopsy (10 cases). Five patients were alive at the date of the last follow-up.
In the validation TMA cohort, the median age of the 193 colon cancer patients was 66 years (ranging from 22 to 75 years); 87 (45.1%) patients were men and 106 (54.9%) were women. At the last follow-up, 73 (37.8%) patients were diagnosed with tumor relapse and 59 (30.6%) patients had died. The characteristics of the patients in the TMA validation cohort are shown in Table 1.
Associations between ACVR2A expression and the clinicopathological variables in patients with colon cancer in the TMA validation cohort (n= 193).
| Variables | n | ACVR2A Expression* | χ2 | P | |
|---|---|---|---|---|---|
| Low (n=79) | High (n=114) | ||||
| Gender | 0.225 | 0.635 | |||
| Male | 87 | 34 | 53 | ||
| Female | 106 | 45 | 61 | ||
| Age (years) | 0.110 | 0.740 | |||
| ≦ 60 | 76 | 30 | 46 | ||
| > 60 | 117 | 49 | 68 | ||
| T category | 0.477 | 0.788 | |||
| T1/2 | 31 | 11 | 20 | ||
| T3 | 75 | 31 | 44 | ||
| T4 | 87 | 37 | 50 | ||
| N stage | 13.070 | 0.001 | |||
| N0 | 107 | 32 | 75 | ||
| N1 | 57 | 29 | 28 | ||
| N2 | 29 | 18 | 11 | ||
| TNM stage | 11.684 | 0.003 | |||
| I | 25 | 9 | 16 | ||
| II | 81 | 23 | 58 | ||
| III | 87 | 47 | 40 | ||
| Pathological grade | 1.039 | 0.308 | |||
| High/ Moderate | 123 | 47 | 76 | ||
| Poor/ Undifferentiation | 70 | 32 | 38 | ||
| Lymphovascular invasion | 8.038 | 0.005 | |||
| Negative | 175 | 66 | 109 | ||
| Positive | 18 | 13 | 5 | ||
| Perineural invasion | 3.015 | 0.081 | |||
| Negative | 160 | 61 | 99 | ||
| Positive | 33 | 18 | 15 | ||
* High and low expression of ACVR2A protein was defined according to the cut-off value of 4 for immunoreactivity score (IS).
Abbreviation: TMA: tissue microarray; T category: depth of tumor infiltration; N stage: lymph node status. Two-tailed χ2 test was used to evaluate the differences between the groups. P value <0.05 was considered to be statistically significant.
Positive role of ACVR2A expression in the GSE39582 database
The expression level of ACVR2A mRNA was first treated as a continuous variable. The normality tests showed that the distribution of expression was normally distributed. The differences in ACVR2A mRNA expression among the subgroups with different TNM stage were not statistically significant (P = 0.107, data not shown). Linear regression analysis verified that the expression level of ACVR2A mRNA was a predictor for RFS in patients with colon cancer in the GSE39582 database (hazard ratio [HR]: 0.519, 95% confidence interval [CI]: 0.387-0.696, P < 0.001).
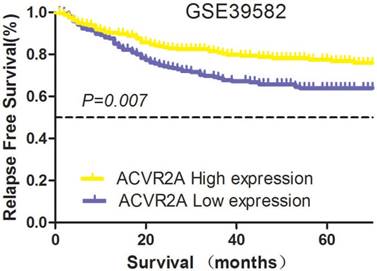
The patient cohort was divided into high and low expression groups according to the median RQ level of ACVR2A mRNA (6.4). For ACVR2A high and low expression groups, 5-year RFS rates were 76.9% and 63.7%, respectively (χ2 = 7.239, P = 0.007) (Fig. 1). The data from this large-sized population suggested that ACVR2A might have a positive role in the survival of patients with colon cancer.
Prognostic significance of ACVR2A in the GSE39582 database. The expression level of ACVR2A mRNA was a predictor for relapse-free survival (RFS) in patients with colon cancer. The patient cohort was divided into high and low expression groups according to the median relative quantitation (RQ) level of ACVR2A mRNA (6.4). The 5-year RFS rates of the high and low expression subgroups were 76.9% and 63.7%, respectively (χ2 = 7.239, P = 0.007).
Reduced ACVR2A expression correlated with colon cancer progression in two validation cohorts
To investigate the expression level of ACVR2A mRNA in patients with metastatic colon cancer, qRT-PCR assays were performed on 15 paired liver metastases, primary colon cancer, and adjacent normal colon tissues. mRNA expression of ACVR2A was the lowest in liver metastases followed by tumor tissues and was highest in adjacent normal controls, with significant difference between each pair of groups (P < 0.001, Fig. 2a). The result indicated that loss of ACVR2A might affect the initiation, progression, and metastasis of colon cancer.
Reduced ACVR2A expression correlates with colon cancer progression. (a) ACVR2A mRNA expression was significantly different among paired tissue samples from 15 stage IV colon cancer patients (P = 0.036 for primary lesions vs. adjacent normal controls; and P = 0.001 for liver metastases vs. primary lesions). The most significant difference was seen between adjacent normal controls and liver metastases (P < 0.001). (b) Representative images of ACVR2A protein expression by immunohistochemical staining of colon cancer lesions in the tissue microarray (TMA) cohort with low expression (i) and high expression (ii), visualized at 200× and 400× (lower right insets) magnification. (c) ACVR2A expression gradually decreased from N0 stage to N2 stage in the TMA cohort (N0 vs. N2, P = 0.001). (d) ACVR2A expression was inversely related to the presence of lymphovascular invasion (LVI) (P = 0.005).
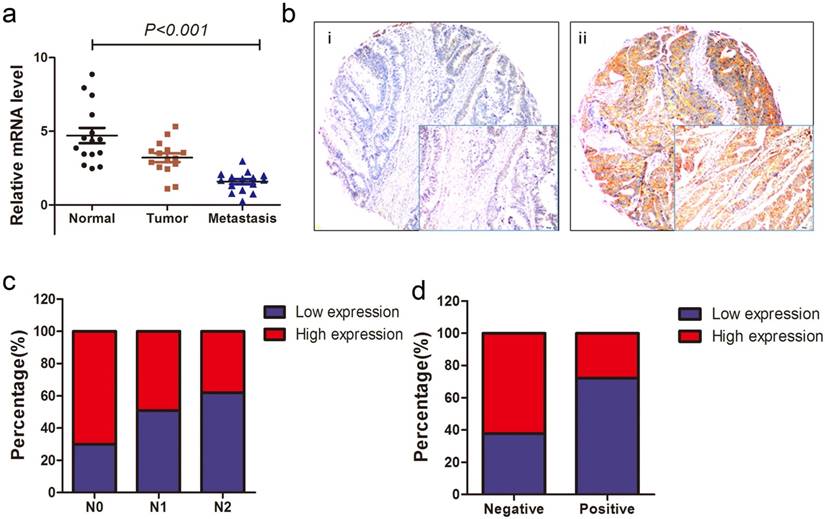
We also examined the correlation between ACVR2A protein expression and patients' clinicopathological characteristics for the TMA cohort. Representative images of ACVR2A IHC staining are shown in Fig. 2b. There were significant correlations between reduced ACVR2A expression and advanced N stage (P = 0.001) or positive lymphovascular invasion (LVI; P = 0.005) (Figs. 2c, 2d, and Table 1), suggesting that ACVR2A may have a key role in colon cancer progression.
Higher ACVR2A expression correlated with survival benefit in colon cancer patients in the TMA cohort
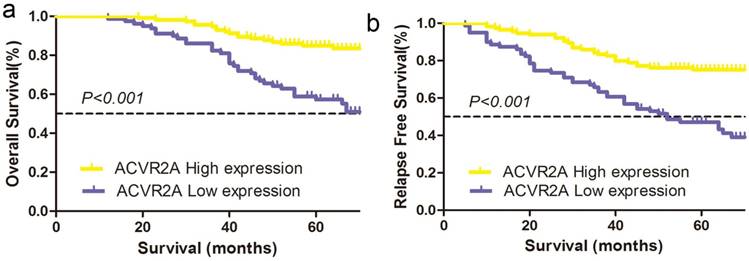
We further evaluated the correlations between the expression of nuclear ACVR2A protein and 5-year OS or RFS of patients in the TMA cohort. Our data revealed a better OS in patients with higher ACVR2A expression (high vs. low expression: 84.9% vs. 57.3%, χ2 =20.427, P < 0.001) (Fig. 3a). Similarly, a better RFS was also observed in patients with higher ACVR2A expression (high vs. low expression: 71.0% vs. 51.4%, χ2 =16.749, P < 0.001) (Fig. 3b).
Higher ACVR2A expression by immunohistochemical staining correlates with better survival of patients with colon cancer in the TMA cohort. (a) The 5-year overall survival (OS) rates in the high and low ACVR2A expression groups were 84.9% and 57.3% respectively (χ2 = 20.427, P < 0.001). (b) The 5-year relapse-free survival (RFS) rates in the high and low ACVR2A expression groups were 71.0% and 51.4%, respectively (χ2 =16.749, P < 0.001).
Furthermore, multivariate Cox regression analysis showed that ACVR2A expression was an independent factor predicting either OS (HR: 0.498, 95% CI: 0.281-0.883, P = 0.017) or RFS (HR: 0.475, 95% CI: 0.287- 0.786, P = 0.004) (Table 2). Our data indicated that ACVR2A might serve as a potential prognostic marker in clinical practice.
Multivariate Cox proportional hazards analysis of ACVR2A expression and overall survival and relapse-free survival for patients with colon cancer in validation cohort (n= 193).
| Variables | Overall survival | Relapse-free survival | ||
|---|---|---|---|---|
| HR (95% CI) | P | HR (95% CI) | P | |
| Gender | 1.095 (0.640-1.873) | 0.740 | 1.095 (0.682-1.758) | 0.707 |
| Age | 1.058 (0.603-1.854) | 0.845 | 0.920 (0.562-1.506) | 0.740 |
| T category | 1.288 (0.901-1.842) | 0.165 | 1.351 (0.984-1.855) | 0.063 |
| N stage | 3.007 (2.068-4.372) | <0.001 | 2.462 (1.762-3.439) | <0.001 |
| Pathological grade | 1.690 (0.969-2.947) | 0.065 | 1.074 (0.647-1.781) | 0.783 |
| Lymphovascular invasion | 1.279 (0.641-2.551) | 0.485 | 1.316 (0.681-2.542) | 0.414 |
| Perineural invasion | 1.747 (0.979-3.119) | 0.059 | 1.457 (0.857-2.478) | 0.165 |
| ACVR2A expression | 0.498 (0.281-0.883) | 0.017 | 0.475 (0.287-0.786) | 0.004 |
Abbreviation: TMA: tissue microarray; CI, confidence interval; HR, hazard ratio; T category: depth of tumor infiltration; N stage: lymph node status. Univariate Cox analysis was first used to identify the risk factors; Cox regression model was then used to perform multivariate analysis. P value <0.05 was considered to be statistically significant.
ACVR2A inhibits the migration of human colon cancer cells
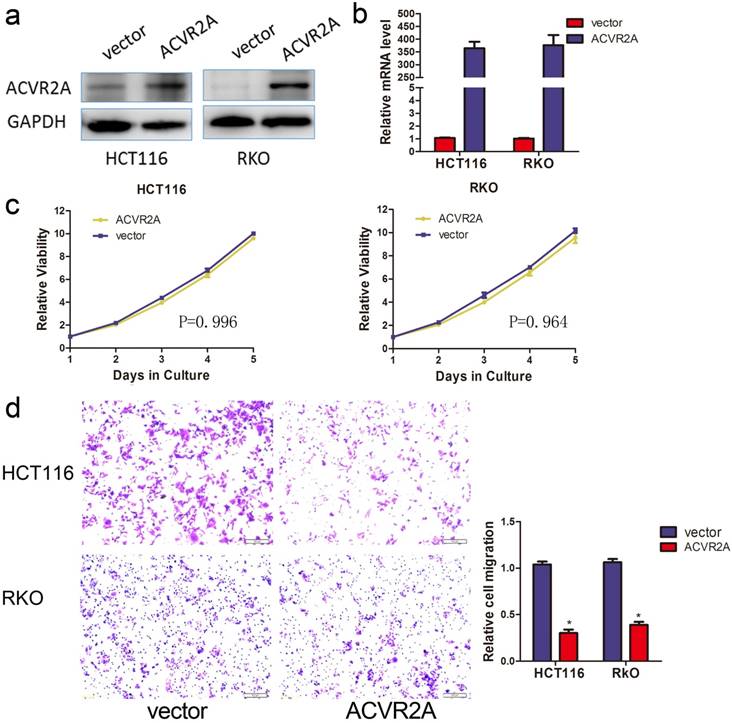
To assess the biological function of ACVR2A in colon cancer, we established stable transfection models of ACVR2A silencing in RKO and HCT116 cell lines. Knockdown efficiencies were verified by western blotting (Fig. 4a) and qRT-PCR (Fig. 4b). Our data indicated that silencing of ACVR2A did not influence the proliferation of colon cancer cells (Fig. 4c). We next investigated whether downregulation of ACVR2A affected the migration of colon cancer cells by Transwell assay. Silencing of ACVR2A resulted in a 2- to 3-fold increase in the number of migrated cells compared with the control group (Fig. 4d).
Silencing of ACVR2A expression promotes the migration of human colon cancer cells. Knockdown of ACVR2A in HCT116 and RKO cells after stable transfection of shRNA against ACVR2A or scramble sequence was analyzed by western blotting (a) and RT-PCR (b). GAPDH was used as the loading control. (c) Silencing of ACVR2A expression did not affect the proliferation of human colon cancer cell lines by CCK8 assay. The cell viability of different groups at each time point of measurement was compared with their relative optical density (OD) value. (d) Knockdown of ACVR2A expression increased the migration of human colon cancer cells by approximately 2- to 3-fold as demonstrated taking from 48 hours after cell seeding (*P < 0.05).
Discussion
In this study, we sought to determine the prognostic value and biological role of ACVR2A in patients with colon cancer. Analysis of the GSE39582 database demonstrated that there was a tendency of downregulated expression of ACVR2A mRNA with more advanced TNM stage. Importantly, downregulated ACVR2A expression as a negative prognostic factor for RFS in patients with colon cancer. Validation cohorts of stage I-III colon cancer patients and TMA identified correlations between reduced ACVR2A expression and colon cancer progression as well as poor survival of patients. Our in vitro studies also showed a remarkable increase in cell migration in ACVR2A knockdown cells. Together, our findings indicate that ACVR2A has an important role in cancer progression and metastasis.
Activin A and TGF-β have been shown to stimulate migration and invasion of various cell types through activation of PI3K/AKT and the MEK/MAPK signaling pathway [6, 8]. Analysis of 15 paired liver metastases, primary tumors, and adjacent normal controls revealed that the mRNA expression of ACVR2A was significantly reduced in metastatic lesions and primary colon cancer tumors compared with adjacent normal controls. Importantly, we also found that reduced ACVR2A expression was significantly correlated with advanced N stages and positive LVI, indicating that ACVR2A may function in the disease progression and metastasis of colon cancer. Our data imply that loss of ACVR2A is correlated with colon cancer progression.
Members of the TGF-β superfamily are involved in regulating diverse biological processes including cell apoptosis, proliferation, and tumor progression [9]. Interestingly, our analysis showed that reduced ACVR2A expression was significantly correlated with a poor 5-year survival for colon cancer patients. These results were obtained from both the publicly available GSE39582 database at mRNA levels and from a validation database at protein levels, therefore our results are both reliable and convincing. Moreover, multivariate Cox repression analysis revealed that ACVR2A is an independent factor for predicting both OS and RFS.
Functional study indicated that silencing of ACVR2A by shRNA significantly decreased colon cancer cell migration in vitro by Transwell assay, but had no impact on cell proliferation. These data further confirmed that ACVR2A is involved in tumor progression and metastasis in colon cancer. Previous studies demonstrated that Activin A inhibits bone morphogenetic protein (BMP) signaling by binding ACVR2A [9], and that BMP signaling could promote colon cancer cell metastasis and invasion via activation of various tumor oncogenes [14-16]. Therefore, it is not surprising that loss of ACVR2A promotes colon cancer migration and progression.
It must be noted that our current study had certain limitations. First, we obtained the conclusions from an in vitro study and tissue samples of human colon cancer from three independent cohorts; however, in vivo experiments on animal models are yet to be performed. Second, we mainly focused on the clinical significance of ACVR2A in colon cancer, and further studies are warranted to explore the mechanism of ACVR2A in the progression and metastases of colon cancer.
In summary, our study provides compelling evidence for an important role of ACVR2A in colon cancer progression. The clinical results showed that downregulation of ACVR2A expression correlates with poor prognostic factors and is associated with worse survival in patients with colon cancer. In vitro studies highlighted the tumorigenic behavior of ACVR2A in human colon cancer cells. Taken together, our findings suggest that ACVR2A may serve as a prognostic marker and potential therapeutic target for the treatment of colon cancer.
Abbreviations
ACVR2A: Activin A receptor type 2A; TGF-β: transforming growth factor-beta; TMA: tissue microarray; IHC: immunohistochemistry; LVI: lymphovascular invasion; PNI: perineural invasion; qRT-PCR: quantitative reverse transcription polymerase chain reaction; IS: immunoreactivity score.
Acknowledgements
This study was accepted and presented as a poster presentation at the 2018 American Society of Clinical Oncology Gastrointestinal Cancers Symposium (ASCO GI 2018) in San Francisco, California, USA (Jan 18-20, 2018).
Funding
This research was sponsored by the grant from the National Key R&D Program of China (No. 2017YFC0908200 to Yang C.), Fujian Provincial Department of Science & Technology (No. 2015Y0009 to Yang C.), and the Medical Innovation Program of Fujian Province (No. 2015-CXB-6 to Zhuo C.).
Contributors
Changhua Zhuo, Dan Hu and Jing Li: Acquisition of data, drafting of the article; Changhua Zhuo, Hui Yu, Qingguo Li, Jing Li, Ying Chen, and Hui Yu: Analysis and interpretation of the data, and statistical expertise; Changhua Zhuo, Xiandong Lin, Xiongwei Zheng and Chunkang Yang: Conception and design of the study. Changhua Zhuo and Chunkang Yang: obtaining of funding; Changhua Zhuo, Jing Li, Hui Yu, Xiongwei Zheng and Qingguo Li: Administrative, technical, or logistic support; Changhua Zhuo, Hui Yu, Yong Zhuang and Ying Chen: Provision of study materials; All of authors: final approval of the article.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Siegel RL, Miller KD, Jemal A. Cancer statistics, 2018. CA: a cancer journal for clinicians. 2018;68:7-30
2. Fahy BN, D'Angelica M, DeMatteo RP, Blumgart LH, Weiser MR, Ostrovnaya I. et al. Synchronous hepatic metastases from colon cancer: changing treatment strategies and results of surgical intervention. Annals of surgical oncology. 2009;16:361-70
3. Edge SB, Compton CC. The American Joint Committee on Cancer: the 7th edition of the AJCC cancer staging manual and the future of TNM. Annals of surgical oncology. 2010;17:1471-4
4. Bauer J, Ozden O, Akagi N, Carroll T, Principe DR, Staudacher JJ. et al. Activin and TGFbeta use diverging mitogenic signaling in advanced colon cancer. Molecular cancer. 2015;14:182
5. Satheesh P, Bhattacharya TK, Kumar P, Chatterjee RN, Dhara SK, Paswan C. et al. Gene expression and silencing of activin receptor type 2A (ACVR2A) in myoblast cells of chicken. British poultry science. 2016;57:763-70
6. Dean M, Davis DA, Burdette JE. Activin A stimulates migration of the fallopian tube epithelium, an origin of high-grade serous ovarian cancer, through non-canonical signaling. Cancer letters. 2017;391:114-24
7. Massague J. TGFbeta signalling in context. Nature reviews Molecular cell biology. 2012;13:616-30
8. Loomans HA, Andl CD. Intertwining of Activin A and TGFbeta Signaling: Dual Roles in Cancer Progression and Cancer Cell Invasion. Cancers. 2014;7:70-91
9. Olsen OE, Wader KF, Hella H, Mylin AK, Turesson I, Nesthus I. et al. Activin A inhibits BMP-signaling by binding ACVR2A and ACVR2B. Cell communication and signaling: CCS. 2015;13:27
10. Network CGA. Comprehensive molecular characterization of human colon and rectal cancer. Nature. 2012;487:330-7
11. Zhuo C, Li Z, Xu Y, Wang Y, Li Q, Peng J. et al. Higher FOXP3-TSDR demethylation rates in adjacent normal tissues in patients with colon cancer were associated with worse survival. Molecular cancer. 2014;13:153
12. Li Q, Wu J, Wei P, Xu Y, Zhuo C, Wang Y. et al. Overexpression of forkhead Box C2 promotes tumor metastasis and indicates poor prognosis in colon cancer via regulating epithelial-mesenchymal transition. American journal of cancer research. 2015;5:2022-34
13. Sinicrope FA, Ruan SB, Cleary KR, Stephens LC, Lee JJ, Levin B. bcl-2 and p53 oncoprotein expression during colorectal tumorigenesis. Cancer research. 1995;55:237-41
14. Bertrand FE, Angus CW, Partis WJ, Sigounas G. Developmental pathways in colon cancer: crosstalk between WNT, BMP, Hedgehog and Notch. Cell cycle (Georgetown, Tex). 2012;11:4344-51
15. Voorneveld PW, Kodach LL, Jacobs RJ, Liv N, Zonnevylle AC, Hoogenboom JP. et al. Loss of SMAD4 alters BMP signaling to promote colorectal cancer cell metastasis via activation of Rho and ROCK. Gastroenterology. 2014;147:196-208.e13
16. Kim BR, Oh SC, Lee DH, Kim JL, Lee SY, Kang MH. et al. BMP-2 induces motility and invasiveness by promoting colon cancer stemness through STAT3 activation. Tumour biology: the journal of the International Society for Oncodevelopmental Biology and Medicine. 2015;36:9475-86
Author contact
Corresponding author: Prof. Chunkang Yang, MD, PhD, Department of Gastrointestinal Surgical Oncology, Fujian Cancer Hospital, No. 420 Fu-ma Road, Fuzhou 350014, China. E-mail address: chuck330com.

 Global reach, higher impact
Global reach, higher impact