Impact Factor ISSN: 1837-9664
J Cancer 2019; 10(16):3632-3638. doi:10.7150/jca.33237 This issue Cite
Research Paper
Tumor Suppressor LKB1 inhibits both the mRNA Expression and the Amplification of hTERC by the Phosphorylation of YAP in Lung Cancer Cells
1. Department of Pathology, The First Affiliated Hospital and College of Basic Medical Sciences, China Medical University, Shenyang 110001, China
2. Department of Gynecology, The First Hospital of China Medical University, Shenyang 110001, China
3. Department of Pathology, Xuzhou City Hospital of TCM, Nanjing University of Chinese Medicine, Xuzhou 221000, China
Received 2019-1-17; Accepted 2019-5-3; Published 2019-6-9
Abstract
Liver kinase B1 (LKB1) is a critical tumor suppressor that is frequently mutated in human cancers. LKB1 has serine/threonine protein kinase activity, which regulates gene expression by phosphorylation of Yes-Associated protein (YAP). The phosphorylation-dependent YAP shuttling is critically important intracellular mechanism in the Hippo pathway. In our previous study, we found that the amplification of hTERC was significant higher in the bronchial brushing cells of patients with lung cancer, however, the underlying molecular mechanism is not clear. In this study, we showed that LKB1 overexpression could phosphorylate YAP and promoted its nuclear rejection. Silencing LKB1 could dephosphorylate YAP and promoted its entry into the nucleus. Here, we found that LKB1 inhibited the mRNA expression and the amplification of hTERC. YAP further up-regulated hTERC at mRNA and gene amplification levels. Therefore, we suggest that LKB1 may inhibit the expression and amplification of hTERC through the axis of LKB1-pYAP(YAP)-hTERC.
Keywords: liver kinase B1, yes-associated protein (YAP), human telomere RNA (hTERC), lung cancer
Introduction
Lung cancer is the most common malignant tumor in the respiratory system and the leading cause of cancer-related death worldwide [1].About 85% of lung cancer patients are diagnosed as non-small cell lung cancer (NSCLC). The incidence of NSCLC is on the rise worldwide, which is often diagnosed at advanced stage. Liver kinase B1 (LKB1) also known as serine/threonine kinase 11 (STK11), was first identified as the causal mutation in Peutz-Jeghers Syndrome (PJS) [2]. LKB1 has become recognised as a critical tumour-suppressor gene which has serine/ threonine protein kinase activity, which regulates gene expression by phosphorylation of substrate proteins or binding to target proteins [3]. LKB1 plays an important role in regulating cell growth, cell metabolism, cell proliferation, cell polarity and energy balance in mammalian tissues. Mutation inactivation of LKB1 can cause many human diseases. Functional deletion mutation of LKB1 gene is an important cause of tumorigenesis, and its inactivation is often observed in lung cancer [4], cervical cancer [5], breast cancer [6], and other tumors [3]. Our previous work found that the expression of LKB1 and hTERT was inversely proportional, and LKB1 could regulate hTERT [7]. In lung cancer, the molecular mechanism by which LKB1 affects the expression of hTERC remains to be elucidated.
Hippo signaling pathway, firstly discovered in Drosophila melanogaster [8], is a highly conserved pathway in mammals that maintains organ size and homeostasis in vivo by regulating cell proliferation and apoptosis [8,9]. Yes-Associated protein (YAP) is the downstream effector of the Hippo pathway, which is inactivated through its cytoplasmic retention by phosphorylation [10,11]. Inactivation of the Hippo pathway leads to YAP nuclear translocation and acts as a transcriptional co-activator to regulate the activity of target protein transcription factors. The phosphorylation-dependent YAP shuttling is critically important intracellular mechanism in the Hippo pathway. Phosphorylated YAP is sequestrated outside the nucleus, however, in the absence of phosphorylation, YAP transfers to the nucleus to activate transcription and promote the proliferation and metastasis of cancer cells. Studies [12] have shown that LKB1 can regulate cell growth and proliferation via the phosphorylation of YAP.
Telomeres, special structures like caps at the chromosome tips in eukaryotic cells, can protect chromosomes tips from fusion, rearrangement, degeneration and translation [13]. With the proliferation of somatic cells, telomeres gradually shorten. When telomeres shrink to a certain extent, cells stop dividing and remain in a static state. Telomere length and stability determine cell life and are closely related to cell senescence and canceration. However, telomere shortening is a risk factor for many diseases [14]. Telomerase is a nucleoprotein complex with the function of synthesizing telomere sequences and balancing telomere consumption [15]. It includes human telomere RNA (hTERC), human telomere reverse transcriptase (hTERT) and human telomere binding protein (hTP1). Among them, hTERC is a DNA template for telomere elongation, and is one of the core components and necessary for the activity of telomere. It is very important for the structure and catalytic activity of telomere. Previous studies [16,7] have reported significantly higher percentages of cells with amplification of hTERC in pleural effusion of lung cancer, and the mRNAs of hTERT and hTERC were more likely to be expressed in the lung cancer group than in the benign group. Therefore, a comprehensive understanding of the regulation mechanism of telomerase activity in lung cancer cells can provide a new direction for the development of new targeted drugs for cancer and new ideas for clinical diagnosis and treatment.
Nguyen HB et al. reported that LKB1 can promote phosphorylation of YAP, and lead to YAP nuclear exclusion [12], and Yu P et al. indicated that YAP is one of the key factors to promote the expression of hTERT [17]. However, there is no research to explain the relationship between YAP and hTERC. We speculate that LKB1 can phosphorylate YAP to retain the phosphorylated YAP in the cytoplasm, thereby inhibiting the entry of YAP into the nucleus and exerting transcriptional co-activation, and then the expression and amplification of hTERC can be inhibited.
Materials and methods
Cell culture
Three human non-small-cell lung carcinomas (NSCLC) cell lines NCI-H460, A549, and LK2 were used in this study. NCI-H460 and A549 cells were purchased from the cell bank of Chinese Academy of Sciences (Shanghai, China) and grown in Hyclone RPMI 1640 medium supplemented with 10% fetal bovine serum (FBS, Cellmax, Beijing, China). The LK2 cell line was a gift from Dr. Hiroshi Kijima (Department of Pathology and Bioscience, Hirosaki University Graduate School of Medicine, Japan) and cultured in Hyclone Dulbecco's modified Eagle's medium (DMEM) supplemented with 10% FBS (Cellmax). Cells were cultured in a humidified incubator at 37°C with 5% CO2.
Plasmid construction and transfection
The plasmids for pcDNA3-LKB1-His and pcDNA3-His were gifts from Professor Xin Hou (College of Life Sciences, Inner Mongolia University, Huhhot, Inner Mongolia, China). The cells were incubated in appropriate antibiotic-free medium with 10% FBS (Cellmax), transferred to 6-well plates culture and incubated at 37°C, CO2 incubator to obtain 70-90% confluens. Thereafter, the cells were performed in 6-well plates using Lipofectamine 3000 (Invitrogen, Carlsbad, CA, USA) as per instructions from the maufacturer. Tansfection with empty vector and mock transfection were used as controls. 24 hours after transfected by quantitative real-time reverse transcriptase-polymerase chain reaction (qRT-PCR), the mRNA analysis was assessed. 48 hours after transfected by western blotting, the protein analysis was assessed.
Small-interfering RNA
Small-interfering RNA(siRNA) was performed to silence the expression of specific genes. siRNAs against LKB1, and YAP1 were purchased from RIBOBIO (Guangzhou, China). Scrambled siRNA (RIBOBIO, Guangzhou, China) was used as a nonspecific siRNA control.
The cells were incubated in appropriate antibiotic-free medium with 10% FBS (Cellmax), transferred to a 6-well plates culture and incubated at 37°C, CO2 incubator to obtain 40-60% confluens. Thereafter, the cells were performed in 6-well plates using Lipofectamine 3000 (Invitrogen, Carlsbad, CA, USA) as per instructions from the maufacturer. Transfection with Scrambled siRNA and mock transfection served as the control. The cells were incubated for 48h after transfection and various analysis to be conducted.
Western blot analysis
Western blot analysis was performed to quantify the amount of target protein. Cells were harvested on ice with ice-cold NP-40 lysis buffer (Beyotime Biotechnology, China) containing 1mM phenylmethylsulfonyl fluoride (PMSF, Beyotime Biotechnology, China). After the concentration of the protein was measured by the Bradford method, 60μg of protein was subjected to 10% SDS-PAGE, blotted onto PVDF membranes (Millipore, Bedford, MA, USA). The membranes were blocked with 5% bovine serum albumin (BSA) and then probed with appropriate primary antibodies separately overnight at 4°C: LKB1 (1:800; Bioss, Beijing, China), YAP (1:1000; Proteintech, Wuhan, China), pYAP (1:1000; Sangon Biotech, Shanghai, China), and GAPDH (1:15000, Proteintech, Wuhan, China). After incubated with appropriate horseradish peroxidase (HRP)-conjugated secondary antibody (1:5000; Proteintech, Wuhan, China), the immunosignal was detected by using ECL Western blot kit (Beyotime Biotechnology, China).The bands were analyzed with BioImaging systems (UVP Inc., Upland, CA).
Quantitative real-time PCR
Total RNA was extracted from cells with Trizol solution (TaKaRa, Dalian, China) according to the manufacturer's instructions. A total of 1μg RNA was subjected to reverse transcription reaction to obtain cDNAs by using a Prime Script TM RT reagent Kit (TaKaRa, Dalian, China). qRT-PCR was carried out on 7900HT Fast Real-Time PCR System (Applied Biosystems) using SYBR® Premix Ex Taq II (TaKaRa, Dalian, China). Non-template controls were carried out every time for each primer pair to detect nonspecific amplification. Glyceraldehyde-3-phosphate dehydrogenase (GAPDH) was amplified as the internal control. All reactions were run in triplicate. The fold change of mRNA expression was calculated using the 2-ΔΔct method. The detailed information of the primers is given in Table 1.
Sequences and features of primers used for qRT-PCR
| Gene | Forward/Reverse | Sequence | Size (bp) | mRNA | |
|---|---|---|---|---|---|
| LKB1 | 223 | AGGGCCGTCAAGATCCTCAA | 187 | KU178339 | |
| 409 | GCATGCCACACACGCAGTA | ||||
| YAP | 875 | TGGCAGCAGTACCAATGGC | 177 | NM_006106 | |
| 1051 | CCAGGTAGTCCTGTCAGAACTT | ||||
| hTERC | 5045 | TCTAACCCTAACTGAGAAGGGCGTAG | 125 | NG_016363.1 | |
| 5170 | GTTTGCTCTAGAATGAACGGTGGAAG | ||||
| GAPDH | 108 | GGAGCGAGATCCCTCCAAAAT | 197 | NM_001256799 | |
| 304 | GGCTGTTGTCATACTTCTCATGG | ||||
mRNA: messenger RNA; qRT-PCR: quantitative real-time reverse transcriptase- polymerase chain reaction
Immunofluorescence
Cells of each group were seeded onto glass slides and keep it for 24 hours in 24-well plates. The cells were washed with PBS and fixed with 4% formaldehyde (Solarbio, Beijing, China) for about 20 minutes after the medium was removed. Then the cells were permeabilized with 0.1% Triton X‐100 (Solarbio, Beijing, China) for 15 min and blocked with 3% bovine serum albumin for 1 h at 37°C. Subsequently, YAP1 (1:50; Proteintech, Wuhan, China) antibody (1:50) was added and incubated overnight at 4 °C. The slides were incubated with the secondary antibody (1:50; Proteintech, Wuhan, China) for 1.5 h at 37°C and with DAPI (Beyotime Biotechnology, China) for 15min. Images were obtained using a fluorescent microscopy (BX-51, Olympus Corporation, Tokyo, Japan).
Fluorescence in situ hybridization (FISH)
FISH was performed according to the manufacturer's protocols (F01007-00, China Medical Technologies, China). The Red-labeled GLP TERC probe which we need to calculate and Green-labeled control probe CSP 3 probe were obtained from 100 independent nuclei. When the TERC signal value was more than 2, and the CSP 3 signal value equaled to 2, the amplified cells are half-segment.
Nuclear and cytoplasmic protein extraction
The cells were harvested on ice with ice-cold Cytoplasmic protein extraction reagent A (Beyotime Biotechnology, China) containing 1mM PMSF (Beyotime Biotechnology, China) and incubated in a 0°C ice bath for 15 mins after putting in vigorous vortex turbulence for 5 seconds. Cytoplasmic protein extraction reagent A (Beyotime Biotechnology, China) was added to the suspension and mixed adequately for 1 min at 0°C in an ice bath, and then centrifuged for 5 mins at 12 000g. The supernatant was absorbed into a pre-cooled plastic tube. The nucleoprotein extraction reagent (Beyotime Biotechnology, China) containing 1mM PMSF was added to the precipitation. The precipitation was suspended and dispersed after 30-second vigorous vortex turbulence, and then intense vortex 30 seconds every 1 minutes for 30 minutes at 0°C in an ice bath. They were centrifuged for 10 mins at 12 000g. The supernatant was absorbed into a pre-cooled plastic tube, which is the extracted nucleoprotein.
Statistical analysis
SPSS 22.0 software was utilized for statistical analyses in this study. Each assay was performed at least 3 times. The data were expressed as mean ± SD and the significance of differences in multiple comparisons was determined by Student's t-test. p<0.05 was considered to be statistically significant.
Results
The Screening of lung cancer cell lines
First, we detected the expression of LKB1 protein in three lung cancer cell lines (NCI-H460, A549, LK2) and normal bronchial epithelial cell lines (HBE) by western blotting. We found that the expression of LKB1 in lung cancer cell lines was lower than that in normal bronchial epithelial cells. The expression of LKB1 protein was relatively higher in NCI-H460 lung cancer cell lines, but lower in A549 and LK2 cell lines (Figure S). Based on these data, A549 and LK2 cell lines were selected for transfection and NCI-H460 cell lines were selected for interference.
LKB1 causes the increase in pYAP expression and inhibits hTERC expression
Due to transient transfection of pcDNA3-LKB1-His to A549 and LK2 two low expression cell lines, the expression levels of LKB1 increased significantly. The elevated levels of LKB1 down-regulated the expression of hTERC at mRNA level (Fig. 1B), and up-regulated pYAP expression at protein level (Fig. 1A). There were no significant changes of YAP mRNA expression level and protein level (Fig. 1A and B).
Effects of LKB1 on the regulation of pYAP, YAP and hTERC expression in lung cancer cell lines. (A) Expression of LKB1, pYAP and YAP were demostrated by western blotting in lung cancer cells lines. (B) Expression of LKB1, YAP and hTERC were demostrated by RT-qPCR in lung cancer cells lines. Mock: mock transfection; vector: empty vector; ns: no significance (*p<0.05; **p<0.0l).
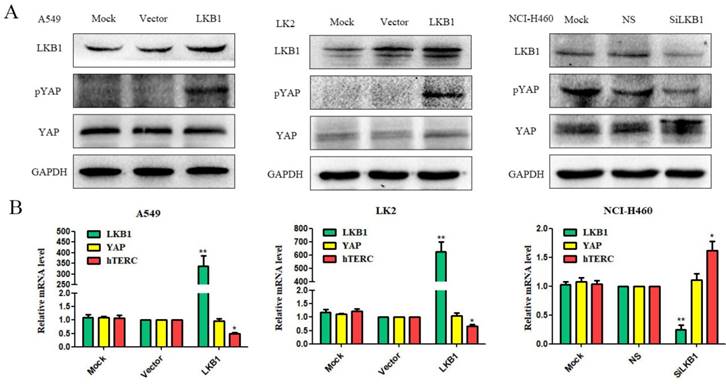
LKB1 depletion down-regulates pYAP expression and up-regulates hTERC expression
Using LKB1-specific siRNA techniques, we knocked down the expression of LKB1 in NCI-H460 cell lines, which resulted in down-regulation of pYAP at protein level, up-regulation of YAP at protein level (Fig. 1B), and up-regulation of hTERC at mRNA level (Fig. 1C). There were no significant changes in YAP mRNA expression level (Fig. 1C).
LKB1 inhibits YAP nuclear localization by the phosphorylation of YAP
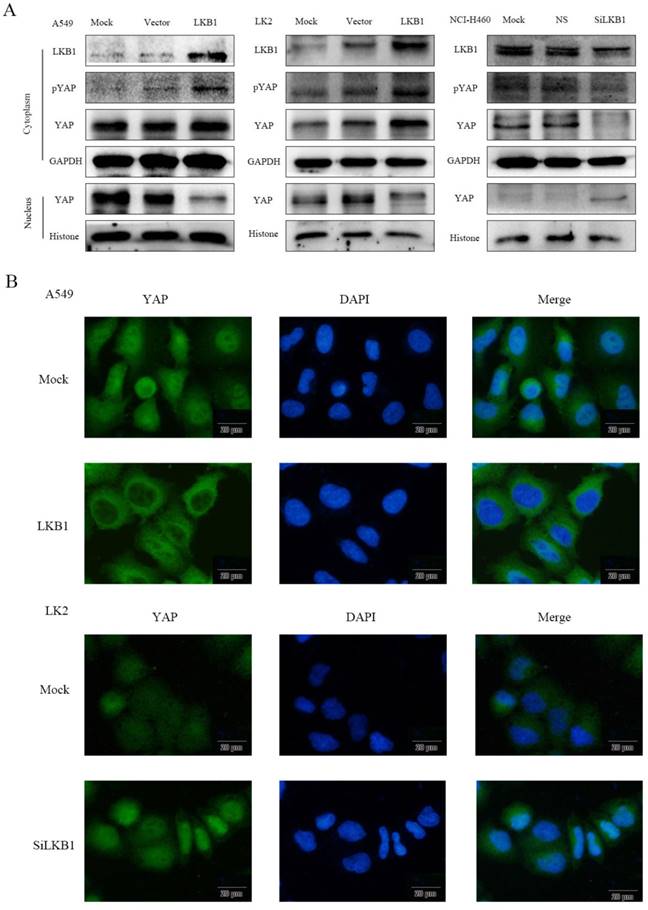
To determine whether the localization of YAP is related to the regulation of LKB1, we used western blotting (Fig. 2A) to detect the expression of YAP and pYAP in nucleic and cytoplasmic fractions. The subcellular expression of YAP was examined by immunofluorescence (Fig. 2B). After transfection of LKB1, the expression of pYAP and YAP in cytoplasm increased significantly, which were consistent with that of pYAP. YAP in the nucleus decreased significantly. After interfering LKB1, the expression of pYAP in cytoplasm decreased significantly, and the expression of YAP decreased, which was consistent with that of pYAP. YAP in the nucleus was significantly increased. However, the expression of YAP mRNA after transfection or interfered with LKB1 was barely altered (Fig. 1C), so we speculated that LKB1 phosphorylated YAP to inhibit its entry into the nucleus. At the same time, we use immunofluorescence test to verify this hypothesis. Immunofluorescence results showed that the expression of YAP in cytoplasm was significantly increased after transfection of LKB1. After interfering with LKB1, we found that the expression of YAP was high in the nucleus.
LKB1 inhibits YAP nuclear localization by the phosphorylation of YAP in lung cancer lines. (A) The relative levels of LKB1, pYAP, YAP, GAPDH, and Histone were measured by western blotting. Cytoplasmic LKB1, pYAP and YAP expression levels were higher but nuclear YAP expression levels were lower in A549 cells and LK2 cells transiently expression LKB1. Cytoplasmic LKB1, pYAP and YAP expression levels were lower but nuclear YAP expression levels were higher in NCI-H460 cells transfected with LKB siRNA. (B) Immunofluorescence showed that LKB1 downregulated YAP nuclear localization while LKB1 siRNA upregulated YAP nuclear localization. Scale bar, 20μm. Mock: mock transfection; vector: empty vector; ns: no significance.
Knocking down YAP suppressed expression and amplifictation of hTERC
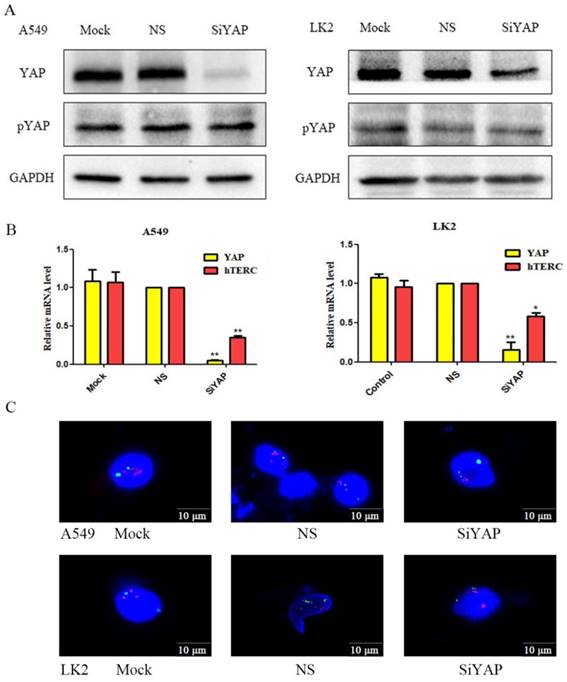
After interfering with YAP, we found that hTERC mRNA decreased significantly and inhibited the amplification of hTERC (Fig.3A and B). We suspect that LKB1 inhibited hTERC expression and amplification by phosphorylating YAP. Therefore, we use Western blot to detect the expression of YAP and pYAP after interference. We found that the expression of pYAP was essentially unchanged, while the expression of YAP was decreased significantly. The experimental results confirmed our hypothesis.
Silencing of YAP results in decreased the expression and amplification of hTERC. (A) The relative levels of YAP, pYAP, and GAPDH were measured by western blotting. Silencing of YAP had no effect on pYAP expression. (B) The relative levels of YAP and hTERC were measured by RT-qPCR. Silencing of YAP downregulated the expression of hTERC. (*p<0.05; **p<0.0l) (C) The amplification of hTERC was measured by FISH. Silencing of YAP downregulated the amplification of hTERC. Mock: mock transfection; Vector: empty vector; ns: no significance.
Discussion
In the previous work, we found that LKB1 could inhibit the expression of hTERT [7], but whether LKB1 could inhibit the expression of hTERC was not clear. As an important component of telomerase, the expression of hTERC is increased in many malignant tumors, including lung cancer [18]. The abnormal expression of hTERC is closely related to the activity of telomerase. Normal people generally have no expression of telomerase except stem cells and active cells such as germinal layer of skin, hair follicle, intestinal mucosa and bone marrow [19]. Therefore, a comprehensive understanding of the regulatory mechanism of telomerase activity in lung cancer cells is essential.
In this study, western blotting and qRT-PCR were used to detect the expression of LKB1, YAP, pYAP and hTERC in lung cancer cells to reveal the relationship between gene regulation, and immunofluorescence was used to analyze the factors affecting the localization of YAP. In our previous work, we found that the expression of LKB1 and hTERT was negatively correlated in bronchial brush cells of lung cancer and benign lesions. However, the regulatory relationship between LKB1 and hTERC has not been reported. This study found that LKB1 could inhibit the expression of hTERC by phosphorylating YAP, which is the continuation and deepening of previous research [16]. LKB1, a critical tumor suppressor gene, can repress YAP activity. YAP is a key effector molecule downstream of Hippo signaling pathway [8]. It regulates signal transduction inside and outside the nucleus through phosphorylation, and acts as a transcription co-activator to regulate the activity of transcription factors [8,9,10].
We hypothesized that LKB1 might inhibit the expression of hTERC through phosphorylation of YAP. In order to study LKB1's regulation on pYAP, YAP and hTERC in lung cancer, we transiently transferred LKB1 plasmid (pcDNA3-LKB1-His) to A549 cells and LK2 cells. LKB1 overexpression significantly inhibited the expression of hTERC at mRNA levels, significantly up-regulated the protein level of pYAP, but the total protein and mRNA of YAP were almost unchanged. LKB1 also promoted the migration of YAP from nucleus to cytoplasm. In contrast, LKB1 depletion increased expression hTERC at mRNA levels in NCI-H460 cells. After interfering with LKB1, the protein level of pYAP was significantly down-regulated, the total protein of YAP was slightly increased, and the expression of YAP at mRNA levels was almost unchanged. We believe that exogenous LKB1 can phosphorylate YAP without altering its total protein and mRNA, while interfering with endogenous LKB1 can dephosphorylate pYAP and increase its total protein, but the expression of YAP at mRNA levels will not change. We speculate that LKB1 only regulates the protein level of YAP. When we specifically interfered with YAP, we found that the expression of hTERC and its atypical amplification were significantly inhibited. Because LKB1 can inhibit hTERC, there was no effect on the change of YAP mRNA, we speculated that it was probably non-phosphorylated YAP that affected hTERC. To verify this hypothesis, we detected the expression of YAP and pYAP after YAP interference. The results showed that after YAP interference, the total protein of YAP decreased significantly, while the protein of pYAP basically unchanged. This result confirmed our hypothesis.
In conclusion, our results suggest that LKB1 inhibits the expression and amplification of hTERC through the axis of LKB1-pYAP(YAP)-hTERC. LKB1 may be a key regulator driving YAP's nuclear and cytoplasmic shuttle in cells. What other regulators exist between them needs to be further studied in the future. YAP promotes the expression and atypical amplification of hTERC , but whether YAP promotes hTERC expression directly or indirectly, and whether there are other regulatory factors between YAP and hTERC remains to be further studied.
Supplementary Material
Supplementary figure.
Acknowledgements
This work was supported by grants from the National Natural Science Foundation of China to Guang-Ping Wu, Grant No.81171650 and 81672082.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A. Global cancer statistics, 2012. CA Cancer J Clin. 2015;65:87-108
2. Hemminki A, Markie D, Tomlinson I, Avizienyte E, Roth S, Loukola A. et al. A serine/threonine kinase gene defective in Peutz-Jeghers syndrome. Nature. 1998;391:184-7
3. Momcilovic M, Shackelford DB. Targeting LKB1 in cancer - exposing and exploiting vulnerabilities. Br J Cancer. 2015;113:574-84
4. Ji H, Ramsey MR, Hayes DN, Fan C, McNamara K, Kozlowski P. et al. LKB1 modulates lung cancer differentiation and metastasis. Nature. 2007;448:807-10
5. Tanwar PS, Kaneko-Tarui T, Zhang L, Tanaka Y, Crum CP, Teixeira JM. Stromal liver kinase B1 [STK11] signaling loss induces oviductal adenomas and endometrial cancer by activating mammalian Target of Rapamycin Complex 1. PLoS Genet. 2012;8:e1002906
6. Lattouf H, Kassem L, Jacquemetton J, Choucair A, Poulard C, Trédan O. et al. LKB1 regulates PRMT5 activity in breast cancer. Int J Cancer. 2019;144:595-06
7. Yang JH, Li XY, Wang X, Hou WJ, Qiu XS, Wang EH. et al. Long-term persistent infection of HPV 16 E6 up-regulate SP1 and hTERT by inhibiting LKB1 in lung cancer cells. Plos One. 2017;12:e0182775
8. Harvey KF, Hariharan IK. The hippo pathway. Cold Spring Harb Perspect Biol. 2012;4:a011288
9. LeBlanc L, Lee BK, Yu AC, Kim M, Kambhampati AV, Dupont SM. et al. Yap1 safeguards mouse embryonic stem cells from excessive apoptosis during differentiation. Elife. 2018;7:e40167
10. Dong J, Feldmann G, Huang J, Wu S, Zhang N, Comerford SA. et al. Elucidation of a universal size-control mechanism in Drosophila and mammals. Cell. 2007;130:1120-33
11. Lee Y, Kim NH, Cho ES, Yang JH, Cha YH, Kang HE. et al. Dishevelled has a YAP nuclear export function in a tumor suppressor context-dependent manner. Nat Commun. 2018;9:2301
12. Nguyen HB, Babcock JT, Wells CD, Quilliam LA. LKB1 tumor suppressor regulates AMP kinase/mTOR-independent cell growth and proliferation via the phosphorylation of Yap. Oncogene. 2013;32:4100-9
13. Cleal K, Norris K, Baird D. Telomere Length Dynamics and the Evolution of Cancer Genome Architecture. Int J Mol Sci. 2018;19:E482
14. Freitas-Simoes TM, Ros E, Sala-Vila A. Nutrients, foods, dietary patterns and telomere length: Update of epidemiological studies and randomized trials. Metabolis. 2016;65:406-15
15. Harpaz T, Abumock H, Beery E, Edel Y, Lahav M, Rozovski U. et al. The Effect of Ethanol on Telomere Dynamics and Regulation in Human Cells. Cells. 2018;7:E169
16. Fan YB, Ye L, Wang TY, Wu GP. Correlation between morphology and human telomerase gene amplification in bronchial brushing cells for the diagnosis of lung cancer. Diagn Cytopathol. 2010;38:402-6
17. Yu P, Shen X, Yang W, Zhang Y, Liu C, Huang T. ZEB1 stimulates breast cancer growth by up-regulating hTERT expression. Biochem Biophys Res Commun. 2018;495:2505-11
18. Kumaki F, Kawai T, Hiroi S, Shinomiya N, Ozeki Y, Ferrans VJ. et al. Telomerase activity and expression of human telomerase RNA component and human telomerase reverse transcriptase in lung carcinomas. Hum Pathol. 2001;32:188-95
19. Hannen R, Bartsch JW. Essential roles of telomerase reverse transcriptase hTERT in cancer stemness and metastasis. FEBS Lett. 2018;592:2023-31
Author contact
Corresponding author: Guang-Ping Wu, Ph.D.: Department of Pathology, The First Affiliated Hospital and College of Basic Medical Sciences, China Medical University, Shenyang 110001, China. Telephone and fax numbers: 00862483282177; E-mail: wugpcom

 Global reach, higher impact
Global reach, higher impact