Impact Factor ISSN: 1837-9664
J Cancer 2020; 11(1):57-68. doi:10.7150/jca.37033 This issue Cite
Research Paper
High expression levels of TLR9 and PD-L1 indicates a poor prognosis in patients with angioimmunoblastic T-cell lymphoma: a retrospective study of 88 cases in a single center
1. Department of Clinical Laboratory, Harbin Medical University Cancer Hospital, Harbin, Heilongjiang 150081, P. R. China.
2. Department of Pathology, Harbin Medical University Cancer Hospital, Harbin, Heilongjiang 150081, P. R. China.
3. Department of Clinical Laboratory, The First Hospital of Harbin, Harbin, Heilongjiang 150010, P. R. China.
4. Department of Medical Oncology, Harbin Medical University Cancer Hospital, Harbin, Heilongjiang 150081, P. R. China.
#These authors contributed equally to this work.
Received 2019-5-27; Accepted 2019-9-22; Published 2020-1-1
Abstract
Background: The role of TLR9 expressed by tumor cells in evading immune surveillance was confirmed. PD-L1 expression in tumor cells plays a key role in tumor immune escape, which is associated with poor prognosis. However, the clinical relevance of TLR9 and PD-L1 expression in angioimmunoblastic T-cell lymphoma (AITL) has not been evaluated.
Materials and methods: In this study, we identified differentially expressed genes in AITL samples by bioinformatic analysis, and we first examined TLR9 and PD-L1 expression by immunohistochemical staining in patients with AITL and compared these data with clinical features and survival time.
Results: It was found that the expression of PD-L1 and multiple TLRs was higher in AITL than normal T-cell samples, and TLR9 and PD-L1 expression displayed complex interactions by bioinformatic analysis. The rates of TLR9 and PD-L1 high expression were 69% and 50%, respectively. High expression of either TLR9 or PD-L1 indicated a poor survival rate for patients with AITL. Multivariate analysis further confirmed that high expression levels of TLR9 and PD-L1 were unfavorable prognostic factors for AITL. We further found inferior overall survival in AITL with clinical features of ECOG status ≥ 2, advanced-stage, elevated serum LDH levels, elevated serum β2-MG levels, and high IPI score.
Conclusion: TLR9 and PD-L1 expression may be a novel predictor of prognosis for patients with AITL and may serve as potential therapeutic strategy.
Keywords: TLR9, PD-L1, angioimmunoblastic T-cell lymphoma, prognosis
Introduction
Angioimmunoblastic T-cell lymphoma (AITL) is an age-related aggressive lymphoma and belongs to the nodal T-cell lymphoma with T follicular helper (Tfh) phenotype [1]. It displays a unique morphologic appearance and is biologically complex [2]. Despite treatment with various therapies, the survival rate of AITL remains bleak with a 5-year overall survival (OS) rate of 30% [3]. Data on the prognostic factors of AITL are limited, and innovative therapeutic targets remain to be determined.
Programmed cell death protein-1 (PD-1) is a member of the CD28/CTLA-4 family of co-stimulatory receptors and conveys inhibitory signals to T cells [4]. Programmed death ligand-1 (PD-L1) is a ligand for PD-1 and is reported to be expressed on tumor cells. The high expression of PD-L1 is associated with poor clinical prognosis in some solid tumors [5,6] and in some subtypes of T- and B-cell lymphomas, including adult T-cell leukemia/lymphoma and diffuse large B-cell lymphoma [7,8]. However, the clinical prognostic implications of PD-1 and PD-L1 expression in AITL are unknown.
Toll-like receptor9 (TLR9), a member of the toll-like receptor family of pattern recognition receptors, binds unmethylated CpG DNA motifs present in microbial nucleic acids and activates the immune response, and is currently under investigation as an adjuvant in anti-PD-1 therapy [9,10]. Recent studies reported that TLR9 is also expressed in some types of tumor cells, including oral cancer, breast cancer, glioma and pancreatic cancer cells. Some researchers suggested that high expression of TLR9 is associated with tumor growth and migration, and it has been recognized as a new prognostic biomarker in cancer [11,12]. To date, there has been limited information about the expression and clinical significance of TLR9 in AITL.
In this study, we identified differentially expressed genes in AITL samples, and using immunohistochemical staining, we analyzed TLR9 and PD-L1 expression in patients with AITL. And we further evaluated the relationship between TLR9 and PD-L1 expression, and compared these data with clinical features and survival time, which may disclose clinical significance of TLR9 and PD-L1 expression in AITL.
Material and methods
Genes expression profiles
Gene expression profiles GSE6338 were downloaded from the Gene Expression Omnibus database (GEO; Affymetrix GPL570 platform, Affymetrix Human Genome U133 Plus 2.0 Array; https://www.ncbi.nlm.nih.gov/geo/) database. We compared gene expression profiles between 5 AITL samples (GSM146170-GSM146174) and 5 normal T-cell samples (GSM146182-GSM146186). Those data were analyzed by the R Statistical Package (R version 3.4.3) to identified differentially expressed genes (DEGs). Each sample has significantly different gene expression profiles. Fold change (FC) > 2, adjusted P-value (adj. P) <0.001, and false discovery rate (FDR) < 0.01 were considered statistically significant.
Functional bioinformatics microarray analysis
To analyze the function of DEGs, biological analyses were performed using The Database for Annotation, Visualization and Integrated Discovery online database (DAVID, version 6.8; https://david.ncifcrf.gov/). In this online database, we input DEGs into the Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) databases to annotate genes and analyze potential functions. The protein-protein interaction (PPI) network was predicted using the Search Tool for the Retrieval of Interacting Genes online database (STRING, version 10.5; https://string-db.org/), and was constructed using Cytoscape (version 3.6.1). Interactions with a combined score > 0.4 were considered statistically significant.
Patient information
A total of 88 patients diagnosed with AITL at Harbin Medical University Cancer Hospital from January 2008 to June 2018 were enrolled. The diagnosis of AITL was based on criteria established by the World Health Organization criteria (WHO 2016) [1]. Risk stratification was assigned according to international prognostic index (IPI) scores (low risk: 0-1; low-intermediate risk: 2; high-intermediate risk: 3; high risk: 4-5). The responses of treatments were defined according to the International Working Group response criteria (published in 2007) [13] or revised response criteria (published as the Lugano classification, 2014) [14].
Immunohistochemistry staining
Immunohistochemical staining was performed on each of paraffin-embedded biopsy specimens obtained from 88 patients with AITL, a 3-μm thick formalin-fixed paraffin-embedded section was submitted. Immunohistochemical staining was performed, as previously described [15]. The primary antibodies used for staining include: anti-TLR9 (ab134368, 1:100; Abcam, Cambridge, UK), anti-PD-L1 (ab205921, 1:100; Abcam, Cambridge, UK), and anti-PD-1 (ab52587, 1:100; Abcam, Cambridge, UK).
Determination of TLR9 and PD-L1
TLR9 and PD-L1 expression was determined by immunohistochemical staining of adjacent tissue sections from the same biopsy specimen. The IHC scoring used for TLR9 and PD-L1 was evaluated independently by two experienced pathologists. The IHC score was acquired by a semi-quantitative method that takes staining intensity and cell proportion into account, and final IHC scores were consensus-based. For TLR9, staining intensity was divided into four categories according to the color of the immune reaction: negative, 0; light brown, 1; brown, 2; and dark brown, 3. The proportion of positively stained cells was determined and scored as: ≤ 5%, 0; 6-25%, 1; 26-50%, 2; 51-75%, 3; and 76-100%, 4. The expression level of TLR9 was obtained by multiplying the intensity and proportion score. And a final score of 0-2 was considered as negative expression (-, noted as 0); 3-4: weak positive expression (+, noted as 1); 6-8: moderate positive expression (++, noted as 2); and 9-12 was strong positive expression (+++, noted as 3) [15]. For PD-L1, based on previous studies, the samples with stained tumor cells < 5% was defined as negative (-, scored as 0). For ≥ 5% positive stained samples, the expression level was further classified into weak positive (+, noted as 1), moderate positive (++, noted as 2), and strong positive expression (+++, noted as 3) according to staining intensity from light brown to brown and dark brown, correspondingly [16]. Samples with negative and weak positive expression were defined as low expression, and samples with moderate and strong positive expression were defined as high expression for TLR9 and PD-L1.
Statistical analysis
Clinicopathological characteristics of the patients were compared using the chi-square test or Fisher's two-sided exact test. The Kaplan-Meier method was used to estimate the overall survival (OS) and progression-free survival (PFS), and the Log-rank test was performed to determine significant differences. Univariate and multivariate Cox proportional regression models were used to evaluate the proposed prognostic factors. P-values < 0.05 were considered statistically significant. Data were analyzed using IBM SPSS software (version 22.0).
Results
Differentially expressed genes in AITL
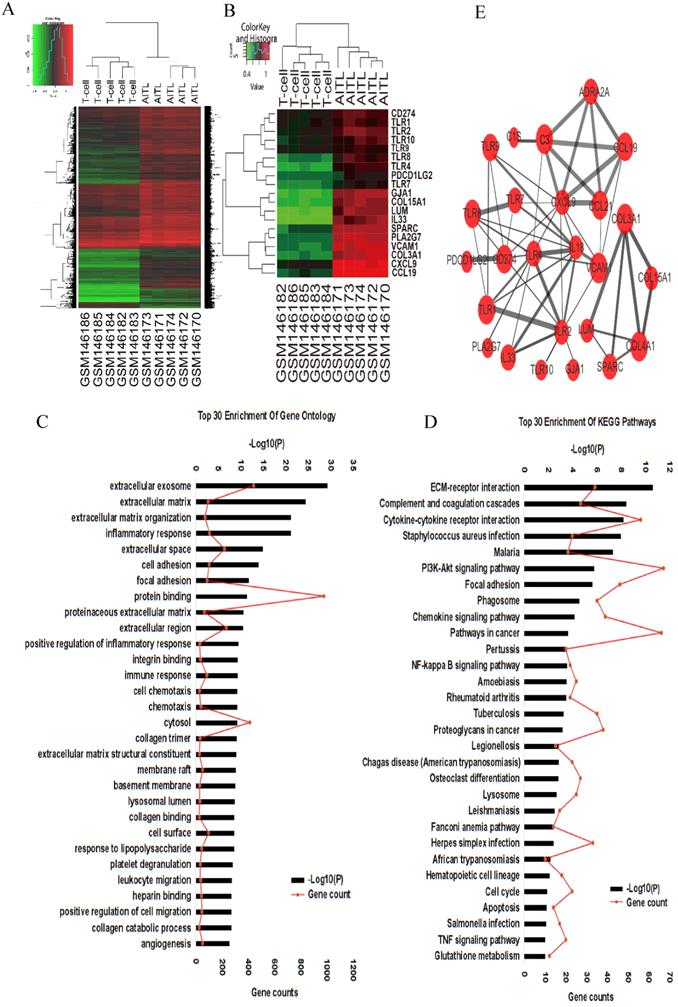
We compared gene expression profiles between 5 AITL samples and 5 normal T cell samples by bioinformatic analysis. A total of 4,439 DEGs in AITL samples were identified, of which 4,182 were upregulated and 257 were downregulated (absolute FC > 2, FDR < 0.01, P < 0.001; Fig. 1A). We assessed the predictive functions of DEGs by GO enrichment analysis, several biological processes such as extracellular exosome, inflammatory response, immune response, cell chemotaxis, cell migration, and angiogenesis were enriched (Fig. 1C). Based on GO enrichment analysis, we suggested some key DEGs are enriched in inflammatory response (IL18, TNF/TNFRSF, TLR9, CXCL10, S1PR3, NLRC4, MYD88), cell chemotaxis (CCL-2, -8, -5, 18, 19, 21, CXCL-9, -10, -12, -14, CXCR-2, -3, -6), cell migration (PDGFRA, JAMA, PTPN6, CD274), and angiogenesis (VEGF, IL18, CXCR3, TGFβ1, ACVRL1, COL15A1) in AITL. Furthermore, we found DEGs related to immune function, which were over-represented in AITL samples, and under-represented in T samples, such as CD274 (PD-L1), PDCD1LG2 (PD-L2), and multiple TLRs (TLR1, TLR2, TLR4, TLR8, TLR9, TLR10) (Fig.1B). Those may be related to the development, tumor microenvironment and treatment sensitivity of AITL. KEGG functional enrichment analysis suggested that DEGs in AITL samples were mainly enriched in the ECM-receptor interaction, cytokine-cytokine receptor interaction, PI3K-Akt signaling pathway, NF- κB signaling pathway, cell cycle, apoptosis, and TNF signaling pathway (Fig. 1D). To further explore the relationship between protein and protein, we constructed a PPI network of 25 DEGs based on GO and KEGG pathway analyses (Supplementary document 1), including the most significant central genes in the network: TLR2, TLR4, and CXCL9. Furthermore, we found that PD-L1 had complex interactions with TLR9 in the network (combined score: 0.449; Fig. 1E; Table 3).
Gene expression profiling analysis in AITL. (A) Hierarchical clustering analysis of 5 AITL samples and 5 normal T cell samples was constructed using the R Statistical Package. A total of 10 samples were clustered according to the expression of 4,439 DEGs. (B) A total of 10 samples were clustered according to the expression of 19 DEGs related to immunological functions. (C-D) Top 30 enrichment GO terms and KEGG pathways for DEGs. (E) PPI network of DEGs was constructed.
The correlation between PD-L1 expression and TLR9 expression on the protein-protein interaction network
| Protein 1 expression | Protein 2 expression | Combined score* |
|---|---|---|
| PD-L1 | TLR9 | 0.449 |
* The combined score of >0.4 was considered to be statistically significant using Cytoscape.
Clinical characteristics
The clinical characteristics of the 88 patients diagnosed with AITL are summarized in Table 1. The patients included 56 males (64%) and 32 females (36%), with a median age of 58 years (range, 17-81 years). Among the patients, systemic B symptoms were described in 53 patients (60%), and 88% of patients had advanced-stage (III and IV) disease by Ann Arbor classification. Almost all the patients had more than one extranodal site, and bone marrow (38%) was the most commonly involved extranodal site, followed by spleen (36%), skin (22%), and liver (13%). Autoimmune hemolytic anemia was observed in 39% of patients. Most AITL patients show elevated serum lactate dehydrogenase (LDH) levels, elevated serum β2-microglobulin (β2-MG) levels, and elevated C-reactive protein levels (84%, 66%, and 33%, respectively). Data on autoantibodies, cryoglobulinemia, and circulating immune complexes were not available for all patients, but antinuclear antibody was found in 33% of patients. Patients were stratified by IPI score; a total of 60% of patients had high-intermediate or high risk. Additionally, Epstein-Barr virus (EBV)-positive status was detected by quantitative PCR in 20 of 25 (80%) patients.
Clinical characteristics at diagnosis of 88 patients with AITL
| Characteristic | No. of Patients | |
|---|---|---|
| (n/N) | % | |
| Age, year, median (rang) | 58(17-81) | |
| < 60 y | 46/88 | 52 |
| ≥ 60 y | 42/88 | 48 |
| Sex | ||
| Male | 56/88 | 64 |
| Female | 32/88 | 36 |
| ECOG score | ||
| 0-1 | 77/88 | 88 |
| 2-4 | 11/88 | 12 |
| B-symptom | ||
| Absent | 35/88 | 40 |
| Present | 53/88 | 60 |
| Tumor Size | ||
| < 7.5cm | 84/88 | 95 |
| ≥ 7.5cm | 4/88 | 5 |
| Primary site | ||
| Lymph node | 83/88 | 94 |
| Extranodal | 5/88 | 6 |
| Ann Arbor stage | ||
| I or II | 11/88 | 12 |
| III or IV | 77/88 | 88 |
| Extranodal sites > 1 | 88/88 | 100 |
| Mediastinal lymphadenopathy | 45/88 | 51 |
| Extranodal involvement | ||
| Liver | 11/88 | 13 |
| Splen | 32/88 | 36 |
| Lung | 5/88 | 6 |
| Skin | 19/88 | 22 |
| Kidney | 1/88 | 1 |
| Breast | 1/88 | 1 |
| Gastrointestinal tract | 1/88 | 1 |
| BM involvement | 33/88 | 38 |
| WBC count > 10×109/L | 27/88 | 31 |
| Neutrophil count > 8×109/L | 25/88 | 28 |
| Lymphocyte count < 0.7×109/L | 25/88 | 28 |
| Platelet counts < 100×109/L | 21/88 | 24 |
| Anemia (HGB < 120 g/L) | 34/88 | 39 |
| Elevated LDH ( > 200 U/L) | 74/88 | 84 |
| Elevated β2-MG (> 3 mg/L) | 58/88 | 66 |
| Elevated CRP (> 20mg/L) | 14/42 | 33 |
| Coombs test (positive) | 3/9 | - |
| ANA (positive) | 4/12 | - |
| EBV DNA (positive) | 20/25 | - |
| IPI score | ||
| 0 or 1 (low risk) | 8/88 | 9 |
| 2 (low-intermediate risk) | 27/88 | 31 |
| 3 (high-intermediate risk) | 37/88 | 42 |
| 4 or 5 (high risk) | 16/88 | 18 |
| Treatment | ||
| Chemotherapy | 79/88 | 90 |
| Adjuvant radiation | 5/79 | - |
| Transplantation | 7/88 | 8 |
| No chemotherapy or radiation | 2/88 | 2 |
| CR or CR (u) | 28/88 | 32 |
Abbreviations: ECOG, Eastern Cooperative Oncology Group; BM, bone marrow; WBC, white blood cell; HGB, Hemoglobin count; LDH, lactate dehydrogenase; β2-MG, β2-microglobulin; CRP, C-reactive protein; ANA, antinuclear antibody; EBV, Epstein-Barr virus; IPI, international prognostic index; CR complete response/remission.
Pathological characteristics
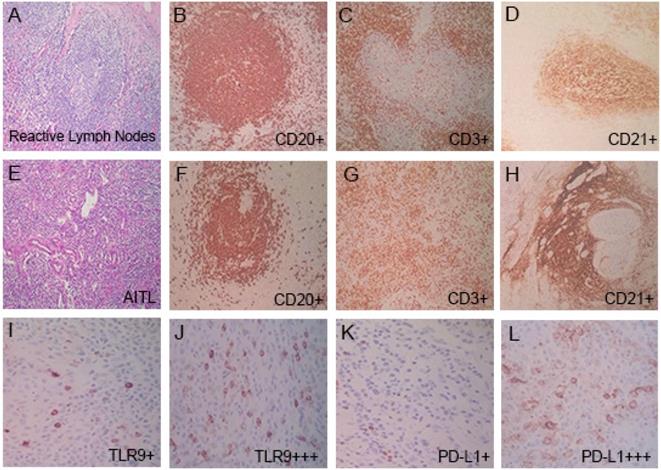
Reactive lymph node samples displayed follicular hyperplasia, the germinal centers were mainly composed of CD20-positive B cells, and the interfollicular areas were mainly composed of CD3-positive T cells (Fig. 2A-2D). In AITL, nodal biopsy exhibited vascular proliferation and numerous intermediate-sized lymphoid cells with irregular nuclei and clear cytoplasm (Fig. 2E-2H). The presence of numerous CD3-positive T cells, CD20 highlighted rare B cells, and an expanded CD21-positive follicular dendritic cell (FDC) meshwork associated with neoplastic T cell proliferation was shown, which is a characteristic feature of AITL. Immunohistochemical examination of tumor cells showed CD3 was positive in 98% (86/88) of patients, CD4 was positive in 92% (81/88), CD10 in 57% (50/88), CD30 in 80% (70/88), BCL-6 in 56% (49/88), and CXCL13 in 88% (77/88) in patients with AITL, and the high expression rates of Ki67 (≥ 45%) were 59% (52/88) (Table 2 ).
Pathologic features in reactive lymph nodes and AITL tissues. (A-D) In reactive lymph nodes, the presence of numerous CD20+ B cells (B), CD3+ T cells (C), and CD21+ FDC (D). (E-L) In AITL tissues, nodal biopsy exhibited vascular proliferation and numerous intermediate-sized lymphoid cells. The presence of rare CD20+ B cells (F), numerous CD3+ T cells (G), and an expanded CD21+ FDC meshwork in AITL tissues (H). TLR9 low and high expression (I, J), and PD-L1 low and high expression (K, L) in AITL tissues.
Pathologic features in 88 patients with AITL
| Immunohistochemistry | NO.(n) | % |
|---|---|---|
| CD3 | 86 | 98 |
| CD4 | 81 | 92 |
| CD5 | 63 | 72 |
| CD20 | 69 | 78 |
| CD30 | 70 | 80 |
| CD21 | 80 | 91 |
| CD10 | 50 | 57 |
| CXCL13 | 77 | 88 |
| PD-1 | 74 | 84 |
| BCL-6 | 49 | 56 |
| MUM-1 | 46 | 52 |
| CD43 | 13 | 15 |
| CD79a | 8 | 9 |
| PD-L1 | 44 | 50 |
| TLR9 | 61 | 69 |
| Ki67 (≥ 45%) | 52 | 59 |
Treatment and outcome
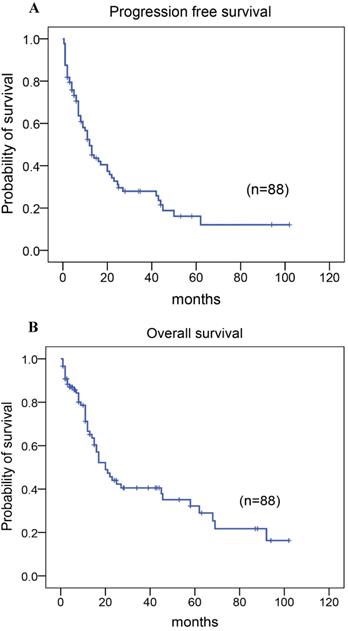
Among the 88 patients, 79 patients received chemotherapy with CHOP, CHOPE, or Hyper CVAD regimen, including five patients who received adjuvant radiotherapy following chemotherapy as the first-line treatment, two patients who did not receive any therapy, and seven patients who had autologous stem cell transplantation as consolidation therapy after achieving the first complete response/remission (CR) (Table 1). A total of 28 patients (32%) achieved CR or unconfirmed CR. The 5-year progression-free survival (PFS) rate was 22%, the median time of PFS was 12.00 months (95% CI: 8.539-15.461; Fig. 3A). And the 5-year overall survival (OS) rate for 88 patients with AITL was 37%, the median time of OS was 20.00 months (95% CI:13.558-26.442; Fig. 3B). At the close of the study, 55% of patients had died.
Kaplan-Meier analysis for PFS and OS in patients with AITL. (A) PFS. (B) OS.
TLR9 and PD-L1 expression in AITL
The expression of TLR9 and PD-L1 in AITL tissues by immunohistochemical staining were shown (Table 2; Figure 2). Of the 88 patients, 84% (74/88) patients had PD-1 high expression. Immunohistochemistry with PD-L1 showed that 50% (44/88) patients had PD-L1 high expression and the rest (50%) were classified as PD-L1 low expression. 69% (61/88) patients were had TLR9 high expression, and 31% (27/88) patients had TLR9 low expression. We further discussed the relationship between TLR9 and PD-L1 expression. Among PD-L1 high expression group (n=44), 70% (31/44) patients had TLR9 high expression, and 30% (13/44) patients had TLR9 low expression. However, there was no significant correlation between TLR9 and PD-L1 expression in AITL (Spearman 'r = 0.025, P = 0.820).
Expression of PD-L1 and TLR9 correlated with reduced OS in AITL
The correlations between TLR9 expression and clinicopathological features were analyzed in this study (Table 4). Compared to the TLR9 low expression group, TLR9 expression was significantly associated with age (P=0.006), gender (P=0.013), ECOG score (P=0.044), and serum β2-MG levels (P=0.019). In addition, the relationships between PD-L1 expression and clinicopathological features were also summarized in Table 4. We found that PD-L1 expression was associated with B symptoms (P=0.050). No statistical associations were observed between PD-L1 expression and age, gender, ECOG score, IPI score, LDH levels, leukocyte count, or Ki-67 expression.
Correlations between TLR9 and PD-L1 expression and clinicopathological features in patients with AITL
| Characteristics | TLR9 | P-value | PD-L1 | P-value | ||
|---|---|---|---|---|---|---|
| Low (n=27) | High (n=61) | Low (n=44) | High (n=44) | |||
| Age(years) | ||||||
| < 60 | 20 | 26 | 0.006 | 21 | 25 | 0.393 |
| ≥ 60 | 7 | 35 | 23 | 19 | ||
| Gender | ||||||
| Male | 12 | 44 | 0.013 | 30 | 26 | 0.375 |
| Female | 15 | 17 | 14 | 18 | ||
| Primary site | ||||||
| Nodal | 24 | 59 | 0.335 | 42 | 41 | 1.000 |
| Extra nodal | 3 | 2 | 2 | 3 | ||
| Tumor size | ||||||
| < 7.5cm | 27 | 57 | 0.420 | 40 | 44 | 0.125 |
| ≥ 7.5cm | 0 | 4 | 4 | 0 | ||
| Ann Arbor stage | ||||||
| I-II | 3 | 8 | 1.000 | 5 | 6 | 0.747 |
| III-IV | 24 | 53 | 39 | 38 | ||
| B symptoms | ||||||
| Absent | 11 | 24 | 0.902 | 13 | 22 | 0.050 |
| Present | 16 | 37 | 31 | 22 | ||
| ECOG score | ||||||
| 0-1 | 27 | 50 | 0.044 | 37 | 40 | 0.334 |
| 2-4 | 0 | 11 | 7 | 4 | ||
| IPI score | ||||||
| Low risk | 4 | 4 | 0.097 | 7 | 1 | 0.141 |
| low-intermediate risk | 9 | 18 | 14 | 13 | ||
| high-intermediat risk | 13 | 24 | 16 | 21 | ||
| High risk | 1 | 15 | 7 | 9 | ||
| LDH level (U/L) | ||||||
| < 200 | 6 | 8 | 0.447 | 9 | 5 | 0.244 |
| ≥ 200 | 21 | 53 | 35 | 39 | ||
| HGB level (g/L) | ||||||
| < 120 | 8 | 26 | 0.248 | 17 | 17 | 1.000 |
| ≥ 120 | 19 | 35 | 27 | 27 | ||
| Platelet count(10^9/L) | ||||||
| < 100 | 6 | 15 | 0.810 | 14 | 7 | 0.080 |
| ≥ 100 Leukocyte count(10^9/L) | 21 | 46 | 30 | 37 | ||
| < 7.2 | 15 | 35 | 0.874 | 24 | 26 | 0.667 |
| ≥ 7.2 | 12 | 26 | 20 | 18 | ||
| Lymphocyte count(10^9/L) | ||||||
| < 0.7 | 8 | 17 | 0.866 | 11 | 14 | 0.478 |
| ≥ 0.7 | 19 | 44 | 33 | 30 | ||
| β2-MG level | ||||||
| < 3.0 | 14 | 16 | 0.019 | 13 | 17 | 0.368 |
| ≥ 3.0 | 13 | 45 | 31 | 27 | ||
| Ki-67 | ||||||
| < 45 | 19 | 33 | 0.152 | 22 | 30 | 0.083 |
| ≥ 45 | 8 | 28 | 22 | 14 | ||
Abbreviations: ECOG, Eastern Cooperative Oncology Group; IPI, international prognostic index; LDH, lactate dehydrogenase; HGB, Hemoglobin; β2-MG, β2-microglobulin.
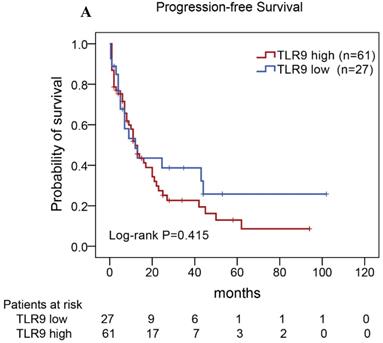
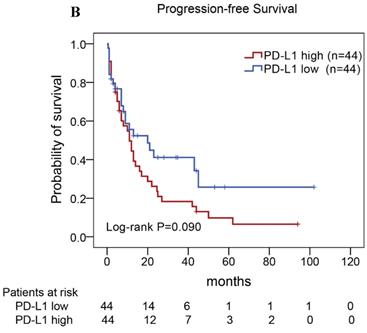
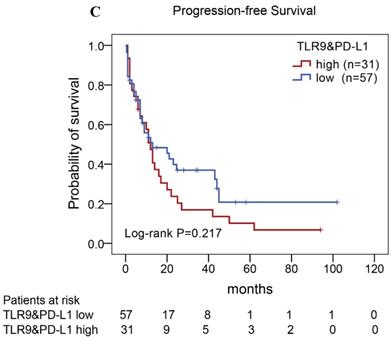
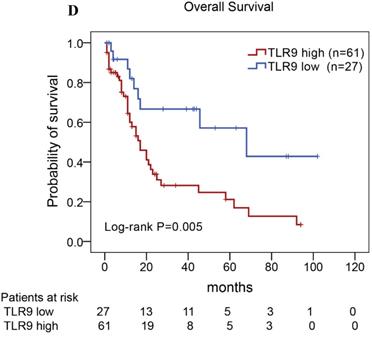
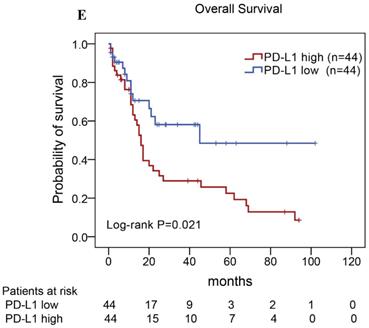
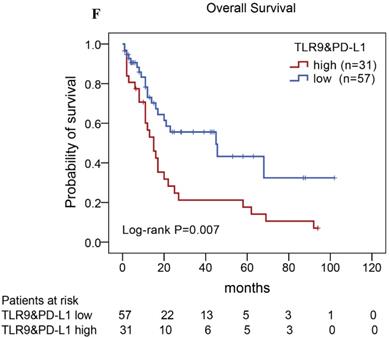
Of the 88 patients with AITL, we found that the 3-year PFS rate for AITL patients with TLR9 high expression was 22%, the median time of PFS was 12 months (Fig. 4A). The 3-year PFS rate for AITL patients with PD-L1 high expression was 19%, the median time of PFS was 11 months (Fig. 4B). And the 3-year PFS rate for AITL patients with both high expression of TLR9 and PD-L1 was 17%, the median time of PFS was 12 months, respectively (Fig. 4C). Although some of the P-value of comparison between the high expression group and the low expression group for PFS was larger than 0.05, the median PFS time was consistently shorter and the PFS time for patients with high expression group was shorter than that for patients with the low expression group. In addition, we further analyzed the correlations between TLR9 and PD-L1 expression and over survival in patients with AITL. AITL patients with high TLR9 expression had a shorter OS compared with low TLR9 expression (17 vs. 68 months; Log-rank P= 0.005; Fig. 4D). AITL patients with high PD-L1 expression had worse OS (16 vs. 45 months; Log-rank P=0.021; Fig. 4E) compared with low PD-L1 expression. Furthermore, AITL patients with the high expression of both TLR9 and PD-L1 had the worst overall survival rate than patients with the single-high and double low expression (15 vs. 45 months; Log-rank P=0.007; Fig. 4F).
Kaplan-Meier analysis for PFS and OS according to TLR9 and PD-L1 expression in patients with AITL. (A-C) Kaplan-Meier analysis for PFS according to TLR9 expression (A), PD-L1 expression (B), and both TLR9 and PD-L1 expression (C) in patients with AITL. (D-F) Kaplan-Meier analysis for OS according to TLR9 expression (D), PD-L1 expression (E), and both TLR9 and PD-L1 expression (F) in patients with AITL.
Univariate and multivariate analyses of prognostic factors in AITL
We performed univariate and multivariate analyses of OS using the Cox proportional hazards model to evaluate the prognostic factors (Table 5). Univariate analysis revealed that TLR9 high expression (Hazard Ratio, HR: 2.677, P=0.008), PD-L1 high expression (HR: 2.037, P=0.026), PD-1 high expression (HR: 3.078, P=0.013), ECOG status ≥2 (HR: 2.672, P=0.032), Ann Arbor stage III or IV (HR: 3.404, P=0.021), elevated serum LDH levels (HR: 3.442, P=0.019), elevated serum β2-MG levels (HR: 1.959, P=0.036), and high IPI score (HR: 5.570,P=0.029) were an individual unfavorable prognostic factors. Multivariate analysis also indicated that TLR9 high expression, PD-L1 high expression, ECOG status ≥ 2, Ann Arbor stage III or IV, elevated serum β2-MG levels were poor prognosis factors in patients with AITL.
Prognostic factors affecting the overall survival of patients with AITL
| Characteristics | Univariate analysis | Multivariate analysis 1* | Multivariate analysis 2** | ||||||
|---|---|---|---|---|---|---|---|---|---|
| HR | (95%CI) | P-value | HR | (95%CI) | P-value | HR | (95%CI) | P-value | |
| TLR9: High vs. Low | 2.677 | (1.291-3.552) | 0.008 | 2.526 | (1.157-5.514) | 0.020 | |||
| PD-L1: High vs. Low | 2.037 | (1.089-3.811) | 0.026 | 2.620 | (1.344-5.107) | 0.005 | |||
| PD-1: High vs. Low | 3.078 | (1.271-7.451) | 0.013 | ||||||
| Age (years): ≥ 60 vs. < 60 | 1.450 | (0.818-2.570) | 0.203 | ||||||
| Gender: Male vs. female | 1.410 | (0.762-2.606) | 0.274 | ||||||
| Primary site: Extranodal vs. Nodal | 0.430 | (0.059-3.134) | 0.405 | ||||||
| Tumour size: ≥ 7.5 cm vs. < 7.5 cm | 0.871 | (0.119-6.352) | 0.892 | ||||||
| Ann Arbor stage: III-IV vs. I-II | 3.404 | (1.202-9.640) | 0.021 | 3.793 | (1.309-10.986) | 0.014 | 3.347 | (1.169-9.579) | 0.024 |
| B symptoms: Present vs. Absent | 1.601 | (0.866-2.961) | 0.134 | ||||||
| ECOG score: 2-4 vs. 0-1 | 2.672 | (1.087-6.569) | 0.032 | 1.673 | (0.676-4.143) | 0.266 | 3.063 | (1.184-7.925) | 0.021 |
| IPI score | |||||||||
| 4/5 vs. 0/1 | 2.188 | (0.493-9.709) | 0.303 | ||||||
| 4/5 vs. 2 | 3.444 | (0.810-14.639) | 0.094 | ||||||
| 4/5 vs. 3 | 5.570 | (1.188-26.115) | 0.029 | ||||||
| LDH levels (U/L): ≥ 200 vs. < 200 | 3.442 | (1.229-9.635) | 0.019 | ||||||
| HGB levels (g/L): < 120 vs. ≥ 120 | 1.083 | (0.597-1.962) | 0.793 | ||||||
| Platelet count (10^9/L): ≥ 150 vs. < 150 | 1.179 | (0.550-2.526) | 0.673 | ||||||
| Leukocyte count(10^9/L): < 7.20 vs. ≥ 7.20 | 0.895 | (0.498-1.611) | 0.712 | ||||||
| Lymphocyte count(10^9/L): < 0.7 vs. ≥ 0.7 | 1.634 | (0.850-3.140) | 0.141 | ||||||
| β2-MG level (mg/L): ≥ 3.0 vs. < 3.0 | 1.959 | (1.046-3.670) | 0.036 | 1.455 | (0.755-2.804) | 0.263 | 2.004 | (1.068-3.762) | 0.030 |
| Ki-67: ≥ 45 vs. < 45 | 0.550 | (0.296-1.024) | 0.059 | ||||||
Abbreviations: HR, hazard ratio; CI, confidence interval; ECOG, Eastern Cooperative Oncology Group; IPI, international prognostic index; LDH, lactate dehydrogenase; HGB, Hemoglobin count; β2-MG, β2-microglobulin. *The variables included in multivariate analysis 1 for OS were TLR9 expression, Ann Arbor stage, ECOG score and β2-MG level. **The variables included in multivariate analysis 2 for OS were PD-L1 expression, Ann Arbor stage, ECOG score and β2-MG level.
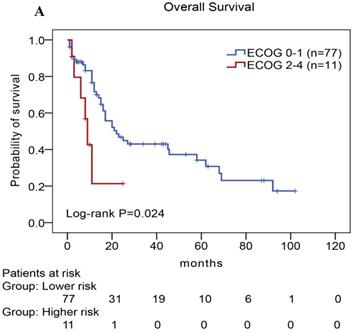
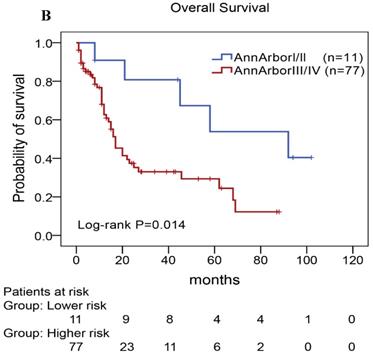
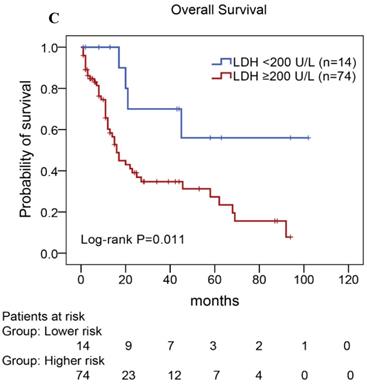
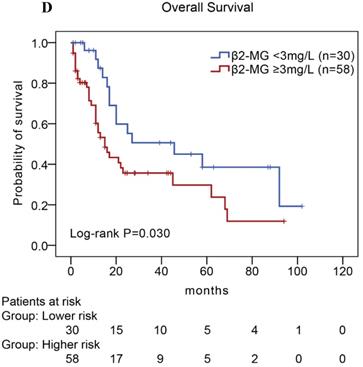
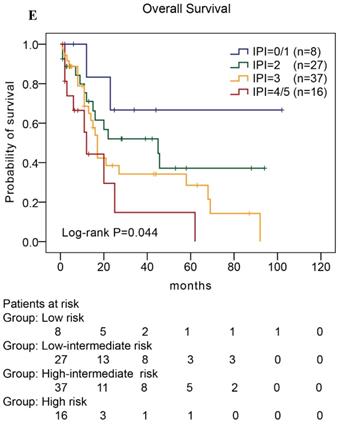
In our study, we further found inferior overall survival in AITL with clinical features of ECOG status ≥ 2, advanced-stage, elevated serum LDH levels, elevated serum β2-MG levels, and high IPI score (Fig. 5). And we further demonstrated that AITL patients with IPI scores of 0 or 1, 2, 3, and 4 or 5 had 2-year OS rates of 68%, 52%, 47%, and 29%, respectively, and high IPI score predicted shorter OS in patients with AITL (Log-rank P=0.044; Fig. 5E).
Kaplan-Meier analysis for OS according to ECOG score (A), Ann Arbor stage (B), serum LDH levels (C), serum β2-MG levels (D), and IPI score (E) in patients with AITL.
Discussion
Due to low incidence, the clinical characteristics and prognosis evaluation of AITL remain poorly understood. In this study, we identified DEGs in AITL by bioinformatics analysis, which may help distinguish AITL from other T-cell lymphoma subtypes. Meanwhile, we studied 88 AITL patients who received treatment at Harbin Medical University Cancer Hospital, which is a large study for a single center in China. And we explored TLR9 and PD-L1 expression in AITL tissues by immunohistochemical staining, and evaluated the association between those expression and clinicopathological features, which may disclose clinical significance of TLR9 and PD-L1 expression in AITL.
Recent study with gene expression profiling of 114 AITL reported that the cytokine environment includes angiogenesis, inflammatory, immunosuppressive and many chemokines/receptors, indicating a complex immune network in AITL [17], which are consistent with our results. In our study, we further identified several DEGs related to immune function including PD-L1, PD-L2, and multiple TLRs, which may play a critical role in the progression of AITL. Based on KEGG pathway analysis, we also evaluated several cancer-related pathways in AITL, including PI3K-Akt signaling pathway, NF-κB signaling pathway, cell cycle, apoptosis, and TNF signaling pathway. Previous study reported that the NF-κB pathway, immunosuppressive pathways (TGFβ pathway) and IL-6 signaling were enriched in AITL [18]. And Zhou LL et al. suggested that activation of PI3K/AKT, JAK-STAT, and NF-κB signaling pathway are associated with processes of T-cell lymphoma [19]. These pathways may play an important role in invasion, metastasis, and therapeutic applications for AITL.
Several studies have demonstrated strong PD-1 expression on tumor-infiltrating lymphocytes in the majority of patients with AITL, and PD-1 expression is the most sensitive marker for AITL [20]. In our study, we reported PD-1 and PD-L1 expression in AITL tissue by immunohistochemical staining. However, we found 14 patients with PD-1 low expression, and further found that these patients had CXCL13, CD10, BCL-6 and other Tfh-related antigen expression, meanwhile, combined with pathomorphological features to confirm the diagnosis of AITL, which was consistent with previous reports [20,21]. Moreover, we demonstrated that patient with high PD-1 expression had poorer OS compared with low PD-1 expression, and high expression of both PD-1 and PD-L1 had the worst survival rate when compared with the single-high and double low expression. These suggested that the PD-1/PD-L1 expression may contribute to a worse clinical outcome in AITL and may be useful as a clinical prognostic indicator to select appropriate therapeutic approaches for individual patients.
In studies on TLRs and cancer, TLR9 is also expressed in lymphoma cells including oral cancer, breast cancer, glioma and pancreatic cancer cells. Several studies reported that the high expression of TLR9 is associated with tumor growth and migration, and it has been recognized as a new prognostic biomarker in cancer. However, the prognostic significance of TLR9 expression in malignancy is still controversial. Previous studies suggested that the high expression of TLR9 was significantly associated with a shorter OS time [22, 23]. However, Leppänen et al. suggested that TLR9 high expression indicated a more favorable prognosis [24]. In this study, we analyzed the expression and clinical relevance of TLR9 in AITL and demonstrated that TLR9 expression was an independent prognostic factor, indicating poor prognosis, which suggested that abnormal TLR9 levels may potentially influence the progression of AITL.
In this study, we further evaluated the correlation between TLR9 expression and PD-L1 expression in a relatively large population with AITL. We constructed a PPI network by bioinformatic analysis, and intuitively shown that TLR9 expression and PD-L1 expression displayed complex interactions. Furthermore, using immunohistochemical staining, we found a potentially relevant trend between TLR9 expression and PD-L1 expression in patients with AITL. Among PD-L1 high expression group, 70% patients had high TLR9 expression, which the number of patients was significantly higher than that of patients with TLR9 low expression. In addition, recent studies have reported the possible potential association between TLR9 and PD-L1 expression. Some in vitro studies demonstrated that TLR9 stimulation can induce PD-L1 expression in mouse tumor cells [25]. In lung cancer, Chen et al. suggested that TLR9 activation in combination with irradiation regulated PD-L1 expression via the NF-kB signaling pathway [26]. Wang D et al. also reported that TLR9 stimulation increased immune checkpoint gene expression, such as PD-L1, in the murine syngeneic A20 lymphoma [27]. Those indicated that PD-L1 expression may be increased or induced by TLR9 ligands. And this may explain that AITL patients with the high expression of both TLR9 and PD-L1 had the worst overall survival rate than patients with the single-high and double low expression in our study. Our found was consistent with previous studies, and indicated the possible potential association between TLR9 and PD-L1 expression. In addition, we will expand the sample size to further confirm our conclusion, and functional studies on the correlation between TLR9 and PD-L1 expression in AITL are still further explored.
In this study, we also analyzed the correlations between TLR9 and PD-L1 expression and survival time in patients with AITL. We found that the median PFS time for patients with low or high expression of those markers was consistently shorter and the PFS time for patients with high expression was shorter than that for patients with the low expression group. Those indicated that AITL patients with high expression had disease progression at an early stage. In addition, the 5-year OS rate for AITL patients with PD-L1 high expression was only 24%, indicating that the disease has strong clinical aggressive characteristics, which was consistent with the characteristics of AITL reported in the previous literature. Thus, exploring the prognostic factors in AITL remain an area of clinical interest. We reported that IPI scores were significant factors in predicting the survival of patients with AITL. We further reported that ECOG performance status, Ann Arbor stage III or IV, elevated serum LDH levels, and elevated serum β2-MG levels were prognostic factors for a shorter OS. However, the application of IPI and PIT scores for AITL is still controversial. Some studies suggested that IPI is not a good prognostic model for AITL [28], while others have supported the prognostic value of IPI in AITL [29,30]. In addition, factors not used in IPI, such as IgA, anemia, bone marrow involvement, also significantly affected prognosis [30]. Additionally, study has suggested that circulating EBV DNA levels were a prognostic factor for survival [31]. A large population and multicenter study is needed to confirm the findings.
In conclusion, we demonstrated that TLR9 and PD-L1 expression indicates a poor prognosis in patients with AITL. The identification of specific prognostic factors for patients with AITL may improve our ability to identify alternative therapies. With the development of novel and complex technologies in genomics and proteomics, significant advances in AITL treatment can be expected in the future.
Acknowledgements
The authors thank Dr. Ken H. Young, MD. PhD, department of Hematopathology, the University of Texas MD Anderson Cancer Center, who provided medical guidance and critical review of the article.
This study is supported by a University of Texas MD Anderson Cancer Center Sister Institution Network Fund Research Grant, the Natural Science Foundation of Heilongjiang Province (grant number H2018045) and Heilongjiang Provincial Returned Scholars Science Fund (grant number LC2015035).
Compliance with ethical standards
All procedures performed involving human participants in our study were in accordance with the ethical standards of the Institutional and/or research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards. The research was approved by the ethic committee of Harbin Medical University Cancer Hospital. Informed consent was obtained from all patients and complied with the Declaration of Helsinki.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Swerdlow SH, Campo E, Pileri SA, Harris NL, Stein H, Siebert R. et al. The 2016 revision of the World Health Organization classification of lymphoid neoplasms. Blood. 2016;127:2375-90
2. Broccoli A, Zinzani PL. Angioimmunoblastic T-Cell Lymphoma. Hematol Oncol Clin North Am. 2017;31:223-38
3. Lunning MA, Vose JM. Angioimmunoblastic T-cell lymphoma: the many-faced lymphoma. Blood. 2017;129:1095-102
4. Pedoeem A, Azoulay-Alfaguter I, Strazza M, Silverman GJ, Mor A. Programmed death- pathway in cancer and autoimmunity. Clin Immunol. 2014;153:145-52
5. Liu CQ, Xu J, Zhou ZG, Jin LL, Yu XJ, Xiao G. et al. Expression patterns of programmed death ligand 1 correlate with different microenvironments and patient prognosis in hepatocellular carcinoma. Br J Cancer. 2018;119:80-8
6. Kim J, Kim S, Lee HS, Yang W, Cho H, Chay DB. et al. Prognostic implication of programmed cell death 1 protein and its ligand expressions in endometrial cancer. Gynecol Oncol. 2018;149:381-7
7. Kataoka K, Iwanaga M, Yasunaga JI, Nagata Y, Kitanaka A, Kameda T. et al. Prognostic relevance of integrated genetic profiling in adult T-cell leukemia/ lymphoma. Blood. 2018;131:215-25
8. Georgiou K, Chen L, Berglund M, Ren W, de Miranda NF, Lisboa S. et al. Genetic basis of PD-L1 overexpression in diffuse large B-cell lymphomas. Blood. 2016;127:3026-34
9. Wagner H. The immunobiology of the TLR9 subfamily. Trends Immunol. 2004;25:381-6
10. Warming "Cold" Melanoma with TLR9 Agonists. Cancer Discov. 2018; 8: 670.
11. Pisani LP, Estadella D, Ribeiro DA. The Role of Toll Like Receptors (TLRs) in Oral Carcinogenesis. Anticancer Res. 2017;37:5389-94
12. Kauppila JH, Korvala J, Siirila K, Manni M, Makinen LK, Hagstrom J. et al. Toll-like receptor 9 mediates invasion and predicts prognosis in squamous cell carcinoma of the mobile tongue. J Oral Pathol Med. 2015;44:571-7
13. Cheson BD, Pfistner B, Juweid ME, Gascoyne RD, Specht L, Horning SJ. et al. Revised response criteria for malignant lymphoma. J Clin Oncol. 2007;25:579-86
14. Sykorova A, Pytlik R, Mocikova H, Belada D, Benesova K, Papajik T. et al. [Staging and Treatment Response Evaluation in Malignant Lymphomas - Czech Lymphoma Study Group Recommendations According to Criteria Revised in 2014 (Lugano Classification)]. Klin Onkol. 2016;29:295-302
15. Akira S, Takeda K. Toll-like receptor signalling. Nat Rev Immunol. 2004;4:499-511
16. Wilcox RA, Feldman AL, Wada DA, Yang ZZ, Comfere NI, Dong H. et al. B7-H1 (PD-L1, CD274) suppresses host immunity in T-cell lymphoproliferative disorders. Blood. 2009;114:2149-58
17. Iqbal J, Wright G, Wang C, Rosenwald A, Gascoyne RD, Weisenburger DD. et al. Gene expression signatures delineate biological and prognostic subgroups in peripheral T-cell lymphoma. Blood. 2014;123:2915-23
18. Iqbal J, Weisenburger DD, Greiner TC, Vose JM, McKeithan T, Kucuk C. et al. Molecular signatures to improve diagnosis in peripheral T-cell lymphoma and prognostication in angioimmunoblastic T-cell lymphoma. Blood. 2010;115:1026-36
19. Zhou LL, Xu XY, Ni J, Zhao X, Zhou JW, Feng JF. T-cell lymphomas associated gene expression signature: Bioinformatics analysis based on gene expression Omnibus. Eur J Haematol. 2018;100:575-83
20. Matsumoto Y, Nagoshi H, Yoshida M, Kato S, Kuroda J, Shimura K. et al. Expression of Master Regulators of T-cell, Helper T-cell and Follicular Helper T-cell Differentiation in Angioimmunoblastic T-cell Lymphoma. Intern Med. 2017;56:2851-6
21. Manso R, Gonzalez-Rincon J, Rodriguez-Justo M, Roncador G, Gomez S, Sanchez-Beato M. et al. Overlap at the molecular and immunohistochemical levels between angioimmunoblastic T-cell lymphoma and a subgroup of peripheral T-cell lymphomas without specific morphological features. Oncotarget. 2018;9:16124-33
22. Herrmann A, Cherryholmes G, Schroeder A, Phallen J, Alizadeh D, Xin H. et al. TLR9 is critical for glioma stem cell maintenance and targeting. Cancer Res. 2014;74:5218-28
23. Niu Z, Tang W, Liu T, Xu P, Zhu D, Ji M. et al. Cell-free DNA derived from cancer cells facilitates tumor malignancy through Toll-like receptor 9 signaling-triggered interleukin-8 secretion in colorectal cancer. Acta Biochim Biophys Sin (Shanghai). 2018;50:1007-17
24. Leppanen J, Helminen O, Huhta H, Kauppila JH, Isohookana J, Haapasaari KM. et al. High toll-like receptor (TLR) 9 expression is associated with better prognosis in surgically treated pancreatic cancer patients. Virchows Arch. 2017;470:401-10
25. Huang B, Zhao J, Li H, He KL, Chen Y, Chen SH. et al. Toll-like receptors on tumor cells facilitate evasion of immune surveillance. Cancer Res. 2005;65:5009-14
26. Chen X, Zhang Q, Luo Y, Gao C, Zhuang X, Xu G. et al. High-dose irradiation in combination with toll-like receptor 9 agonist CpG oligodeoxynucleotide 7909 downregulates PD-L1 expression via the NF-kappaB signaling pathway in non-small cell lung cancer cells. Onco Targets Ther. 2016;9:6511-8
27. Wang D, Jiang W, Zhu F, Mao X, Agrawal S. Modulation of the tumor microenvironment by intratumoral administration of IMO-2125, a novel TLR9 agonist, for cancer immunotherapy. Int J Oncol. 2018;53:1193-203
28. Federico M, Rudiger T, Bellei M, Nathwani BN, Luminari S, Coiffier B. et al. Clinicopathologic characteristics of angioimmunoblastic T-cell lymphoma: analysis of the international peripheral T-cell lymphoma project. J Clin Oncol. 2013;31:240-6
29. Kao HW, Lin TL, Shih LY, Dunn P, Kuo MC, Hung YS. et al. Clinical features, outcome and prognostic factors of 87 patients with angioimmunoblastic T cell lymphoma in Taiwan. Int J Hematol. 2016;104:256-65
30. Hong H, Fang X, Wang Z, Huang H, Lam ST, Li F. et al. Angioimmunoblastic T-cell lymphoma: a prognostic model from a retrospective study. Leuk Lymphoma. 2018;59:2911-6
31. Liang JH, Lu L, Zhu HY, Li W, Fan L, Li JY. et al. The Prognostic Role of Circulating Epstein-Barr Virus DNA Copy Number in Angioimmunoblastic T-Cell Lymphoma Treated with Dose-Adjusted EPOCH. Cancer Res Treat. 2019;51:150-7
Author contact
Corresponding authors: Wenhui Li, Department of Clinical Laboratory, Harbin Medical University Cancer Hospital, No.150 Haping Road, Nangang District, Harbin, Heilongjiang Province, 150081, China. Tel: 86-015344510273. Email: jykqjr1993com; Shu Zhao. Department of Medical Oncology, Harbin Medical University Cancer Hospital, No.150 Haping Road, Nangang District, Harbin, Heilongjiang Province, 150081, China. Tel: 86-013946158616. Email: zs_1881com.

 Global reach, higher impact
Global reach, higher impact