Impact Factor ISSN: 1837-9664
J Cancer 2020; 11(24):7312-7319. doi:10.7150/jca.46630 This issue Cite
Research Paper
Amide-type local anesthetics may suppress tumor cell proliferation and sensitize Human Hepatocellular Carcinoma Cells to Cisplatin via upregulation of RASSF1A expression and demethylation
1. Department of Anesthesiology, Sun Yat-Sen University Cancer Center, State Key Laboratory of Oncology in South China, Collaborative Innovation Center for Cancer Medicine, Guangzhou 510060, China.
2. Department of Anesthesiology, Huizhou Municipal Central Hospital, Huizhou 516001, China.
3. Department of Anesthesiology, Peking University Shenzhen Hospital, Shenzhen 518000, China.
4. Department of Hepatobiliary Oncology, Sun Yat-Sen University Cancer Center, State Key Laboratory of Oncology in South China, Collaborative Innovation Center for Cancer Medicine, Guangzhou 510060, China.
*These authors equally contributed to this work.
Received 2020-4-3; Accepted 2020-10-10; Published 2020-10-23
Abstract
Background: It has been reported that local anesthetics are toxic to various types of cells. Furthermore, several local anesthetics have been confirmed to exert demethylation effects and regulate the proliferation of human cancer cells. Our previous findings suggest that lidocaine may exert potential antitumor activity and enhance the sensitivity of cisplatin to hepatocellular carcinoma in vitro and in vivo. A recent study proved that lidocaine sensitizes breast cancer cells to cisplatin via upregulation of RASSF1A, a promotor of tumor suppressive gene (TSG) demethylation. We sought to determine whether amide-type local anesthetics (lidocaine, ropivacaine and bupivacaine) exert growth-inhibitory effects on human hepatoma cells and to determine whether amide-type local anesthetics sensitize human hepatoma cells to cisplatin-mediated cytotoxicity via upregulation of RASSF1A expression.
Methods: Human hepatoma cell lines HepG2 and BEL-7402 were incubated with lidocaine, ropivacaine and bupivacaine. The viability of local anesthetic-treated cells with or without cisplatin was investigated. Further, we evaluated RASSF1A expression after treatment of HepG2 and BEL-7402 cells with three local anesthetics and determined the influence of RASSF1A expression on the toxicity of cisplatin to these cells.
Results: The viability of HepG2 and BEL-7402 cells was significantly decreased by treatment with amide-type local anesthetics (lidocaine, ropivacaine and bupivacaine). In these cells, the combination treatment with cisplatin and local anesthetics exhibited a stronger reduction in viability. Lidocaine, ropivacaine and bupivacaine promoted a significant increase in RASSF1A expression and a decrease in RASSF1A methylation. The combined treatment with both local anesthetics and cisplatin resulted in a significantly lower level of HepG2 and BEL-7402 cell viability than that with singular local anesthetics or cisplatin treatment. Moreover, local anesthetics enhanced the cytotoxicity of cisplatin against HepG2 and BEL-7402 cells, accompanied by an increase in RASSF1A expression.
Conclusions: These data indicated that amide-type local anesthetics (lidocaine, ropivacaine and bupivacaine) have growth-inhibitory and demethylation effects in human hepatoma cells. We also found that these amide local anesthetics may enhance the cytotoxicity of cisplatin in human hepatocellular carcinoma cells possibly via upregulation of RASSF1A expression and demethylation.
Keywords: amide-type local anesthetics, DNA demethylation, cisplatin, RASSF1A, human hepatoma cells
Introduction
Recently, an increasing number of studies have shown the potential mechanisms of local anesthetic-induced tumor suppression. Several pathways have been described in the literature. On the one hand, local anesthetics, at high doses, are cytotoxic in vitro and in vivo [1-4], and on the other hand, they may induce sensitization of tumor cells to chemotherapeutics [1, 5] and heat [6].
Another potential mechanism whereby local anesthetics may influence tumor growth is by interaction with the tumor epigenome [3, 7, 8]. Studies have indicated that the silencing of tumor suppressor genes (TSGs) through methylation of their promoters is one of the causes of tumor development [9]. As one of the natural covalent modifications of chromatin, defined as epigenomic modifications [10], the silencing of TSGs by DNA methylation induces mechanisms responsible for promoting tumor development, such as metastasis, avoidance of apoptosis, and uncontrolled cell growth or maintaining angiogenesis. In malignant tumors, increased methylation often leads to downregulation of tumor suppressor genes, which is conducive to tumor progression [11].
As an example, methylation of promoter regions of TSGs, such as VHL, CDKN2A and BRCA1, renders them inactive in cancer cells [9, 12]. Moreover, DNA methylation may facilitate the mutation of TSGs. The tumor suppressive p53 gene is mutated in more than 50% of solid tumors, and 25% of the mutations result from methylated cytosine to thymine changes in the CpG dinucleotides of this gene [13]. Quantitative analysis of DNA methylation profiles for more cancer-related genes also indicates a strong association of TSG hypermethylation with lung cancers [14] and colorectal cancers [15]. Specific promoter hypermethylation of these genes is dependent on tumor type [16-18]. Unlike genetic alterations, epigenetic gene silencing through DNA hypermethylation is potentially reversible, and this probably can provide new opportunities for the clinical treatment of malignant tumors.
Several local anesthetics have been confirmed to exert demethylation effects and to regulate the proliferation of human cancer cells. Procaine promotes DNA demethylation and inhibits the growth of the human breast cancer cell line MCF-7 [16] and of human hepatoma cell lines [3]. Recently, a study confirmed the demethylation effect of lidocaine in breast cancer cells and found that the demethylation of Ras association domain family 1A (RASSF1A) sensitized breast cancer cells to cisplatin-mediated cytotoxicity [8].
Our previous findings suggested that lidocaine may exert potential antitumor activity and enhance the sensitivity of hepatocellular carcinoma to cisplatin in vitro and in vivo [1]. However, whether other commonly used amide local anesthetics, such as ropivacaine and bupivacaine, have antitumor effects in hepatocellular carcinoma has not been proven. The tumor suppressive gene RASSF1A [19-22] has also been shown to be silenced by promoter methylation in hepatocellular carcinoma. Therefore, whether local anesthetics sensitize hepatocellular carcinoma cells to cisplatin via upregulation of RASSF1A expression is unclear.
The aim of this study was therefore to test two hypotheses: (i) the amide-type local anesthetics lidocaine, ropivacaine and bupivacaine may exert potential antitumor activity in hepatocellular carcinoma and (ii) the three amide-type local anesthetics may sensitize hepatocellular carcinoma cells to cisplatin-mediated cytotoxicity via upregulation of RASSF1A expression.
Materials and Methods
Cell Culture and Treatment with Reagents
HepG2 and BEL-7402 hepatocellular carcinoma cells were obtained from the Chinese Type Culture Collection (China). Cells were routinely cultured in high glucose in Dulbecco's modified Eagle's medium supplemented with 10% (v/v) fetal bovine serum (Invitrogen, USA) and maintained at 37 °C in a humidified atmosphere of 5% CO2 plus 95% air. Lidocaine, ropivacaine, bupivacaine, 5-aza-2'-deoxycytidine (DAC, a demethylation agent as a positive control) and cis-diammineplatinum dichloride (cisplatin) were purchased from Sigma-Aldrich (USA).
Cell Counting Kit-8 (CCK-8) Assay
Cell viability was determined by the Cell Counting Kit-8 assay kit (Dojindo Molecular Technologies, Japan) according to the manufacturer's instructions. HepG2 and BEL-7402 cells were seeded at 3,000 cells per well in 96-well plates and were incubated at 37 °C for 24 h to approximately 85% confluence, and then cells were treated with amide-type local anesthetics (one of lidocaine, ropivacaine and bupivacaine) or cisplatin (alone or in combination) at the indicated concentrations. Cell Counting Kit-8 was added to each well, and the plate was incubated for 3 h at 37 °C. The absorbance value (OD) was read at 450 nm using a spectrophotometer, and the data were analyzed using Softmax Pro software (Molecular Devices LLC, USA). Each experiment was performed in triplicate.
Methylation Analysis of TSG RASSF1A
The DNA methylation was analyzed using MethyLight method [23], to quantify the methylation level of RASSF1A, which has been reported to be hypermethylated in breast cancer [24]. The methylation status of individual CpG sites in this gene within the promoter region was determined by the sodium bisulfite-sequencing assay as described previously [25]. Each experiment was performed in triplicate.
RNA Extraction and RT-qPCR
Total cellular mRNA samples were extracted from cultured cells using the RNeasy Mini Kit (Qiagen, Valencia, CA, USA). Quantification of RASSF1A expression was conducted using the QuantiTect SYBR Green PCR Kit (Qiagen, Hilden, Germany) and an ABI 7900HT Sequence Detector System (Applied Biosystems). The sequences of the primers for RASSF1A were as follows: forward primer: 5'-AGCC TGAG CTCA TTGA GCTG-3', reverse primer: 5'-ACCA GCTG CCGT GTGG-3'. All mRNA expression levels were normalized to β-actin (forward primer: 5'-GATG AGAT TGGC ATGG CTTT-3', reverse primer: 5'-GTCA CCTT CACC GTTC CAGT-3'), and the ΔΔCt method was used for relative quantification [26]. Each experiment was performed in triplicate.
Protein Isolation and Western Blot Analysis
Total cellular protein samples were extracted using a radioimmunoprecipitation assay buffer containing a protease inhibitor cocktail (Roche Boehringer Mannheim Diagnostics, Switzerland) and quantified using a bicinchoninic acid protein assay kit (Thermo Scientific) according to the manual. Equal amounts of proteins (40 μg/lane) were separated by 8%~10% sodium dodecyl sulfate-polyacrylamide gel electrophoresis and transferred onto polyvinylidene fluoride membranes (Millipore, USA). The membranes were incubated with primary antibodies at 4 °C overnight and then incubated with horseradish peroxidase-conjugated secondary antibodies at room temperature for 1 h. The blots were detected using an electrochemiluminescence system (Pierce Biotechnology, USA).
Statistical analysis
Data are shown as the mean ± SEM. To determine differences between three or more means, one-way ANOVA with Bonferroni post hoc tests was performed. A P value of < 0.05 was considered statistically significant.
Results
Effect of amide-type local anesthetics on cell proliferation
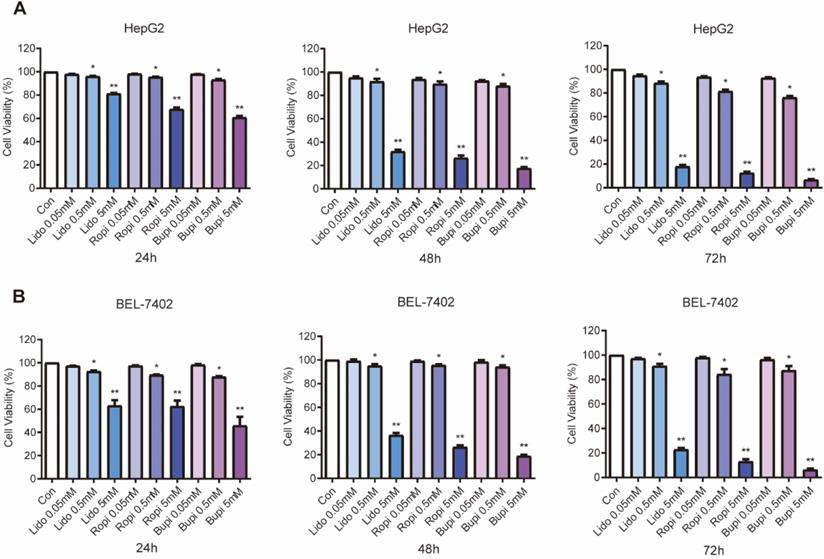
Separate treatment with lidocaine, ropivacaine and bupivacaine at 0.05, 0.5, or 5 mM resulted in significant reductions in cell number, while lower concentrations of local anesthetics had no effect. Anesthetics inhibited the growth of HepG2 (Figure 1A) and BEL-7402 (Figure 1B) cells in a dose-dependent manner.
Effect of lidocaine, ropivacaine and bupivacaine on cell proliferation of HepG2 and BEL-7402 human hepatocellular carcinoma cells. (A) HepG2 cell numbers after lidocaine, ropivacaine and bupivacaine treatment. (B) BEL-7402 cell numbers after lidocaine, ropivacaine and bupivacaine treatment. The results are expressed as the mean (SD). Statistical significance is shown as * P ˂ 0.05, or ** P ˂ 0.01, ns: no significance.
Amide-type local anesthetics increase the expression and demethylation of RASSF1A gene in HepG2 and BEL-7402 cells
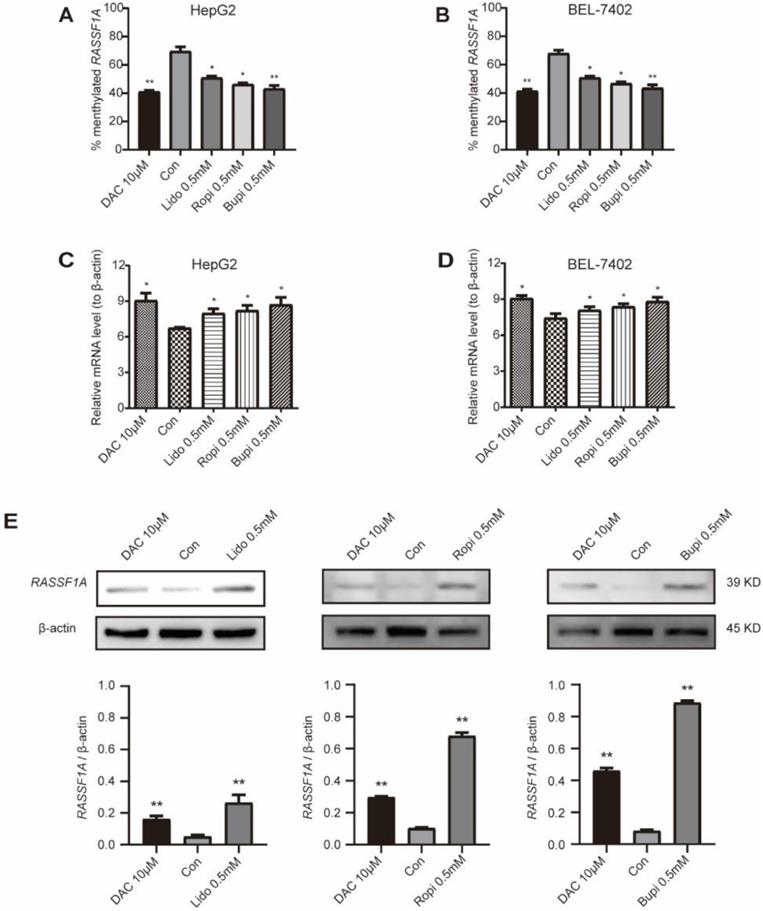
Ras association domain family 1A (RASSF1A) [28,29], a classical TSG, was found to interact with molecules such as CNK1 [33], Nore1 [34], MDM2 [35], and MAP1S [36] to induce cell death. Methylation of this TSG has been reported to silence its expression and promote the tumorigenesis of hepatocellular carcinoma [19-22] and breast cancers [27-29]. We evaluated the methylation levels of RASSF1A in HepG2 and BEL-7402 cells with lidocaine, ropivacaine and bupivacaine treatment separately. Figure 2A and B indicates the methylation level of this gene compared to that of the global genomic CpG islands in HepG2 (Figure 2A) and BEL-7402 (Figure 2B) cells; as shown in Figure 2A and B, 10 μM 5-aza-2'-deoxycytidine (DAC, a demethylation agent as a positive control) treatment for 48 h significantly reduced the methylation of the RASSF1A gene in HepG2 and BEL-7402 cells compared to that in the control (P < 0.05). To examine the influence of promoter demethylation in HepG2 and BEL-7402 cells, the expression of RASSF1A mRNA was examined using real-time quantitative PCR (RT-qPCR), and the expression of RASSF1A protein was assessed by western blotting. As shown in Figure 2C and D, there was a significant increase in RASSF1A mRNA levels either by 10 μM 5-aza-2'-deoxycytidine (P < 0.05) or by 0.5 mM lidocaine, ropivacaine or bupivacaine treatment for 48 h (P < 0.05, respectively). Furthermore, the western blot assay reconfirmed the upregulation of RASSF1A at the protein level by demethylation, which was promoted by 0.5 mM lidocaine, ropivacaine or bupivacaine separate treatment for 48 h in HepG2 (Figure 2E; P < 0.05) cells. Therefore, the demethylation of RASSF1A promotes the expression of this gene.
Local anesthetics upregulate the expression and demethylation of RASSF1A. (A) Methylation levels of RASSF1A in HepG2 cells treated for 48 h with 0.5 mM lidocaine, ropivacaine, bupivacaine, or with 10 µM DAC, respectively. (B) Methylation levels of RASSF1A in BEL-7402 cells treated for 48 h with 0.5 mM lidocaine, ropivacaine, bupivacaine or with 10 µM DAC, respectively. (C) mRNA level of RASSF1A in HepG2 cells post treatment with 0.5 mM lidocaine, ropivacaine, bupivacaine, or with 10 µM DAC for 48 h, respectively. (D) mRNA level of RASSF1A in BEL-7402 cells post treatment with 0.5 mM lidocaine, ropivacaine, bupivacaine or with 10 µM DAC for 48 h, respectively. (E) Western blot assay showing the protein level of RASSF1A in HepG2 cells with or without 0.5 mM lidocaine, ropivacaine, bupivacaine, or 10 µM DAC treatment for 48 h, respectively. Each value is expressed as the mean (SD) of three independent tests. Statistical significance is shown as * P < 0.05, or ** P < 0.01, ns: no significance.
Amide-type local anesthetics enhance the cytotoxicity of cisplatin against HepG2 and BEL-7402 cells possibly via upregulating RASSF1A expression
Cisplatin has been clinically utilized for hepatocellular carcinoma treatment for decades [27-29], and its antiproliferation effect has been confirmed to be abrogated [30] or enhanced [1, 31] by other molecules. To investigate the regulation of local anesthetics on the cisplatin-mediated antitumor effect, we evaluated the proliferation of HepG2 and BEL-7402 cells by CCK-8 assay after treatment with 0.5 mM of each local anesthetic separately, 10 μM cisplatin, or combinations of cisplatin and each anesthetic. It is shown in Figure 3A and B that HepG2 (Figure 3A) and BEL-7402 (Figure 3B) cells with or without the abovementioned treatment resulted in significant reductions in cell number (P < 0.05) and increases in cisplatin cytotoxicity (P < 0.01).
Local anesthetics enhance the cytotoxicity of cisplatin against HepG2 and BEL-7402 cells possibly via upregulating RASSF1A expression. (A) HepG2 cell numbers after treatment with 10 µM cisplatin or (and) 0.5 mM local anesthetics for 48 h. (B) BEL-7402 cell numbers after treatment with 10 µM cisplatin or (and) 0.5 mM local anesthetics for 48 h. (C) Western blot analysis of RASSF1A expression in HepG2 cells without or with 0.5 mM local anesthetics and/or 10 µM cisplatin treatment for 48 h. The results are expressed as the mean (SD). Statistical significance is shown as *P < 0.05, or **P < 0.01 different from the corresponding control group; &P < 0.05, or &&P < 0.01 local anesthetics groups versus local anesthetics plus cisplatin groups; #P < 0.05, or ##P < 0.01 cisplatin group versus local anesthetics plus cisplatin groups; ns: no significance.
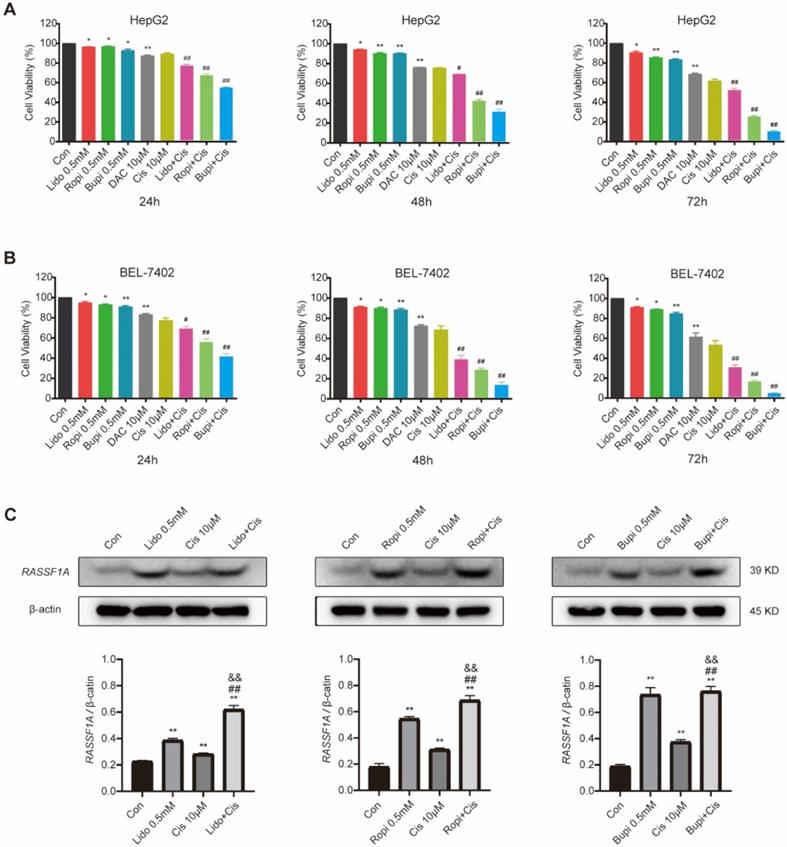
To further investigate the association of the local anesthetic-induced expression of RASSF1A with HepG2 cell proliferation inhibition, we analyzed RASSF1A expression in HepG2 cells without or with amide-type local anesthetics and/or cisplatin treatment by western blot assay (Figure 3C). The results showed that the protein expression level of RASSF1A was upregulated when 0.5 mM lidocaine, ropivacaine or bupivacaine were separately combined with cisplatin for 48 h in HepG2 cells (P < 0.01).
Discussion
Studies have indicated that silencing of tumor suppressor genes (TSGs) by promoter methylation contributes to tumor development [9] that the inactivation of TSGs through the hypermethylation of the 5'-CpG islands in gene promoter regions induces processes such as uncontrolled cell growth, metastasis or apoptosis inhibition and also supports the maintenance of angiogenesis [9, 12]. As opposed to genetic alteration, epigenetic gene silencing through DNA hypermethylation is potentially reversible, and this presents new opportunities for the clinical management of malignant tumors. DAC (5-aza-2'-deoxycytidine), a demethylating agent, is currently being tested in clinical trials of cancer treatment. DNA demethylation by DAC effectively abolishes 17 β-estradiol-induced cell growth of human breast cancer cells [32]; DAC decreases the incidence of gastric cancers and inhibits their growth [33, 34]; miRNA-34b inhibits prostate cancer through demethylation, activating chromatin modifications [35]. However, the clinical utility of these drugs has not been fully realized because of their adverse effects, such as myelosuppression and mutagenicity [36]. The mechanisms of the antitumor effects of DNA demethylation agents still need to be clarified.
Recently, procaine, known as an ester-type local anesthetic, has been revealed to be a nonnucleoside inhibitor of DNA methylation and has a growth-inhibitory effect in human hepatoma cells [3]. Lidocaine, known as an amide-type local anesthetic, has been indicated to promote DNA demethylation [8, 37] and increase the cytotoxicity of cisplatin in breast cancer cells via upregulation of RASSF1A demethylation [8]. In our previous study, we demonstrated that lidocaine may exert potential antitumor activity and enhance the sensitivity of hepatocellular carcinoma to cisplatin in vitro and in vivo [1].
In the current study, we first used the CCK-8 assay to investigate whether amide-type local anesthetics (lidocaine, ropivacaine and bupivacaine) reduced the viability of human hepatoma cells. Treatment with 0.05, 0.5, or 5 mM lidocaine, ropivacaine or bupivacaine led to a decrease in viable HepG2 and BEL-7402 cells, and the local anesthetics reduced the cell viability in a dose-dependent manner in these cells.
Ras association domain family 1A (RASSF1A), a well-recognized tumor suppressive gene in various types of tumors, is a putative tumor suppressor gene located on the 3p21.3 locus [38-40]. The hypermethylation of CpG islands in RASSF1A occurs in a large percentage of human hepatocellular carcinomas [19-22].
The results here indicate that hypermethylation in this gene was reduced significantly by treatment with three local anesthetics. The three local anesthetics restored the expression of RASSF1A at both the mRNA and protein levels in HepG2 and BEL-7402 cells.
Interestingly, treatment with lidocaine, ropivacaine or bupivacaine for 48 h attenuated the reduction in cell viability promoted by 10 μM cisplatin in both cell lines. The sensitization effect of the three local anesthetics on the antitumor effect of cisplatin was confirmed by CCK-8 assay; combined treatment with 0.5 mM one of the three local anesthetics and 10 μM cisplatin significantly reduced the viability in HepG2 cell lines compared to the singular 0.5 mM one of the three local anesthetics or singular 10 μM cisplatin treatment. Furthermore, we demonstrated by western blot analysis that local anesthetics can enhance the cytotoxicity of cisplatin against hepatoma cells in vitro possibly via upregulate of RASSF1A expression.
Although the concentrations of local anesthetics are far too high to be clinically relevant, these concentrations are very similar to those used in previous studies under in vitro conditions [37, 41, 42]. One of our previous studies also used similar concentrations in vitro [1]. It is not appropriate to extrapolate our findings here to humans. Nevertheless, our findings suggest the anti-hepatic tumor potential of amide-type local anesthetics.
Taken together, these data indicate that the amide-type local anesthetics (lidocaine, ropivacaine and bupivacaine) have growth-inhibitory effects on hepatoma cells in vitro. Each of the three local anesthetics can increase the expression and demethylation of RASSF1A gene in HepG2 and BEL-7402 cells. Amide-type local anesthetics may sensitize human hepatocellular carcinoma cells to cisplatin via upregulation of RASSF1A expression and demethylation.
Acknowledgements
Funding
This work was supported by grants from the National Natural Science Foundation of China (grant 81702394 to WX and grant 81902490 to DTC).
Author contributions
WX and WAZ designed this study. DTC, YY, JDX, JHP and YHC performed all the experiments. DTC, YY, JDX and QL interpreted the data. WX, YFY and WAZ offered technical support. DTC, WX and WAZ drafted the manuscript. All authors polished the paper.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Xing W, Chen DT, Pan JH, Chen YH, Yan Y, Li Q. et al. Lidocaine Induces Apoptosis and Suppresses Tumor Growth in Human Hepatocellular Carcinoma Cells In vitro and in a Xenograft Model In vivo. Anesthesiology. 2017;126:868-81
2. Lucchinetti E, Awad AE, Rahman M, Feng J, Lou PH, Zhang L. et al. Antiproliferative effects of local anesthetics on mesenchymal stem cells: potential implications for tumor spreading and wound healing. Anesthesiology. 2012;116:841-56
3. Tada M, Imazeki F, Fukai K, Sakamoto A, Arai M, Mikata R. et al. Procaine inhibits the proliferation and DNA methylation in human hepatoma cells. Hepatol Int. 2007;1:355-64
4. Perez-Castro R, Patel S, Garavito-Aguilar ZV, Rosenberg A, Recio-Pinto E, Zhang J. et al. Cytotoxicity of local anesthetics in human neuronal cells. Anesth Analg. 2009;108:997-1007
5. Esposito M, Fulco RA, Collecchi P, Zicca A, Cadoni A, Merlo F. et al. Improved therapeutic index of cisplatin by procaine hydrochloride. J Natl Cancer Inst. 1990;82:677-84
6. Zwischenberger JB, Vertrees RA, Bedell EA, McQuitty CK, Chernin JM, Woodson LC. Percutaneous venovenous perfusion-induced systemic hyperthermia for lung cancer: a phase I safety study. Ann Thorac Surg. 2004;77:1916-24 discussion 25
7. Esteller M. Relevance of DNA methylation in the management of cancer. Lancet Oncol. 2003;4:351-8
8. Li K, Yang J, Han X. Lidocaine sensitizes the cytotoxicity of cisplatin in breast cancer cells via up-regulation of RARbeta2 and RASSF1A demethylation. Int J Mol Sci. 2014;15:23519-36
9. Lewandowska J, Bartoszek A. DNA methylation in cancer development, diagnosis and therapy-multiple opportunities for genotoxic agents to act as methylome disruptors or remediators. Mutagenesis. 2011;26:475-87
10. Kubicek S, Schotta G, Lachner M, Sengupta R, Kohlmaier A, Perez-Burgos L. et al. The role of histone modifications in epigenetic transitions during normal and perturbed development. Ernst Schering Res Found Workshop. 2006 p:1-27
11. Teschendorff AE, Menon U, Gentry-Maharaj A, Ramus SJ, Weisenberger DJ, Shen H. et al. Age-dependent DNA methylation of genes that are suppressed in stem cells is a hallmark of cancer. Genome Res. 2010;20:440-6
12. Ushijima T. Detection and interpretation of altered methylation patterns in cancer cells. Nat Rev Cancer. 2005;5:223-31
13. Singal R, Ginder GD. DNA methylation. Blood. 1999;93:4059-70
14. Vaissiere T, Hung RJ, Zaridze D, Moukeria A, Cuenin C, Fasolo V. et al. Quantitative analysis of DNA methylation profiles in lung cancer identifies aberrant DNA methylation of specific genes and its association with gender and cancer risk factors. Cancer Res. 2009;69:243-52
15. van Engeland M, Weijenberg MP, Roemen GM, Brink M, de Bruine AP, Goldbohm RA. et al. Effects of dietary folate and alcohol intake on promoter methylation in sporadic colorectal cancer: the Netherlands cohort study on diet and cancer. Cancer Res. 2003;63:3133-7
16. Villar-Garea A, Fraga MF, Espada J, Esteller M. Procaine is a DNA-demethylating agent with growth-inhibitory effects in human cancer cells. Cancer Res. 2003;63:4984-9
17. Costello JF, Fruhwald MC, Smiraglia DJ, Rush LJ, Robertson GP, Gao X. et al. Aberrant CpG-island methylation has non-random and tumour-type-specific patterns. Nat Genet. 2000;24:132-8
18. Esteller M, Corn PG, Baylin SB, Herman JG. A gene hypermethylation profile of human cancer. Cancer Res. 2001;61:3225-9
19. Schagdarsurengin U, Wilkens L, Steinemann D, Flemming P, Kreipe HH, Pfeifer GP. et al. Frequent epigenetic inactivation of the RASSF1A gene in hepatocellular carcinoma. Oncogene. 2003;22:1866-71
20. Retraction Note to. Role of RASSF1A promoter methylation in the pathogenesis of hepatocellular carcinoma: a meta-analysis of 21 cohort studies. Mol Biol Rep. 2015;42:1453
21. Liu ZJ, Huang Y, Wei L, He JY, Liu QY, Yu XQ. et al. Combination of LINE-1 hypomethylation and RASSF1A promoter hypermethylation in serum DNA is a non-invasion prognostic biomarker for early recurrence of hepatocellular carcinoma after curative resection. Neoplasma. 2017;64:795-802
22. Mansour LA, El Raziky M, Mohamed AA, Mahmoud EH, Hamdy S, El Sayed EH. Circulating Hypermethylated RASSF1A as a Molecular Biomarker for Diagnosis of Hepatocellular Carcinoma. Asian Pac J Cancer Prev. 2017;18:1637-43
23. Eads CA, Danenberg KD, Kawakami K, Saltz LB, Blake C, Shibata D. et al. MethyLight: a high-throughput assay to measure DNA methylation. Nucleic Acids Res. 2000;28:E32
24. Nole F, Munzone E, Mandala M, Catania C, Orlando L, Zampino MG. et al. Vinorelbine, cisplatin and continuous infusion of 5-fluorouracil (ViFuP) in metastatic breast cancer patients: a phase II study. Ann Oncol. 2001;12:95-100
25. Yan PS, Shi H, Rahmatpanah F, Hsiau TH, Hsiau AH, Leu YW. et al. Differential distribution of DNA methylation within the RASSF1A CpG island in breast cancer. Cancer Res. 2003;63:6178-86
26. Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods. 2001;25:402-8
27. Kim Y, Jang M, Lim S, Won H, Yoon KS, Park JH. et al. Role of cyclophilin B in tumorigenesis and cisplatin resistance in hepatocellular carcinoma in humans. Hepatology. 2011;54:1661-78
28. Schrader J, Gordon-Walker TT, Aucott RL, van Deemter M, Quaas A, Walsh S. et al. Matrix stiffness modulates proliferation, chemotherapeutic response, and dormancy in hepatocellular carcinoma cells. Hepatology. 2011;53:1192-205
29. Lo SJ, Fan LC, Tsai YF, Lin KY, Huang HL, Wang TH. et al. A novel interaction of nucleophosmin with BCL2-associated X protein regulating death evasion and drug sensitivity in human hepatoma cells. Hepatology. 2013;57:1893-905
30. Su YC, Davuluri GV, Chen CH, Shiau DC, Chen CC, Chen CL. et al. Galectin-1-Induced Autophagy Facilitates Cisplatin Resistance of Hepatocellular Carcinoma. PLoS One. 2016;11:e0148408
31. Hao J, Li Z, Zhang C, Yu W, Tang Z, Li Y. et al. Targeting NF-kappaB/AP-2beta signaling to enhance antitumor activity of cisplatin by melatonin in hepatocellular carcinoma cells. Am J Cancer Res. 2017;7:13-27
32. Singh KP, Treas J, Tyagi T, Gao W. DNA demethylation by 5-aza-2-deoxycytidine treatment abrogates 17 beta-estradiol-induced cell growth and restores expression of DNA repair genes in human breast cancer cells. Cancer Lett. 2012;316:62-9
33. Schneider BG, Peek RM Jr. Gastric cancer prevention by demethylation. Cancer Prev Res (Phila). 2013;6:253-6
34. Liu J, Xie YS, Wang FL, Zhang LJ, Zhang Y, Luo HS. Cytotoxicity of 5-Aza-2'-deoxycytidine against gastric cancer involves DNA damage in an ATM-P53 dependent signaling pathway and demethylation of P16(INK4A). Biomed Pharmacother. 2013;67:78-87
35. Majid S, Dar AA, Saini S, Shahryari V, Arora S, Zaman MS. et al. miRNA-34b inhibits prostate cancer through demethylation, active chromatin modifications, and AKT pathways. Clin Cancer Res. 2013;19:73-84
36. Santini V, Kantarjian HM, Issa JP. Changes in DNA methylation in neoplasia: pathophysiology and therapeutic implications. Ann Intern Med. 2001;134:573-86
37. Lirk P, Berger R, Hollmann MW, Fiegl H. Lidocaine time- and dose-dependently demethylates deoxyribonucleic acid in breast cancer cell lines in vitro. Br J Anaesth. 2012;109:200-7
38. Levesley J, Lusher ME, Lindsey JC, Clifford SC, Grundy R, Coyle B. RASSF1A and the BH3-only mimetic ABT-737 promote apoptosis in pediatric medulloblastoma cell lines. Neuro Oncol. 2011;13:1265-76
39. Yi M, Yang J, Chen X, Li J, Li X, Wang L. et al. RASSF1A suppresses melanoma development by modulating apoptosis and cell-cycle progression. J Cell Physiol. 2011;226:2360-9
40. Matallanas D, Romano D, Yee K, Meissl K, Kucerova L, Piazzolla D. et al. RASSF1A elicits apoptosis through an MST2 pathway directing proapoptotic transcription by the p73 tumor suppressor protein. Mol Cell. 2007;27:962-75
41. Sakaguchi M, Kuroda Y, Hirose M. The antiproliferative effect of lidocaine on human tongue cancer cells with inhibition of the activity of epidermal growth factor receptor. Anesth Analg. 2006;102:1103-7
42. Chang YC, Hsu YC, Liu CL, Huang SY, Hu MC, Cheng SP. Local anesthetics induce apoptosis in human thyroid cancer cells through the mitogen-activated protein kinase pathway. PLoS One. 2014;9:e89563
Author contact
Corresponding authors: Weian Zeng (E-mail: zengwasysu.edu.cn) or Wei Xing (E-mail: xingweiorg.cn).

 Global reach, higher impact
Global reach, higher impact