Impact Factor
ISSN: 1837-9664
J Cancer 2021; 12(12):3626-3647. doi:10.7150/jca.52170 This issue Cite
Research Paper
Five-gene signature associating with Gleason score serve as novel biomarkers for identifying early recurring events and contributing to early diagnosis for Prostate Adenocarcinoma
1. Department of Laboratory Medicine, The Second Affiliated Hospital of Zhejiang University School of Medicine, Hangzhou, 310009, China.
2. Department of Biochemistry and Molecular Biology, Bengbu Medical College, Anhui 233030, China.
*These authors contributed equally to this work.
Abstract
Background: Compared to non-recurrent type, recurrent prostate adenocarcinoma (PCa) is highly fatal, and significantly shortens the survival time of affected patients. Early and accurate laboratory diagnosis is particularly important in identifying patients at high risk of recurrence, necessary for additional systemic intervention. We aimed to develop efficient and accurate diagnostic and prognostic biomarkers for new PCa following radical therapy.
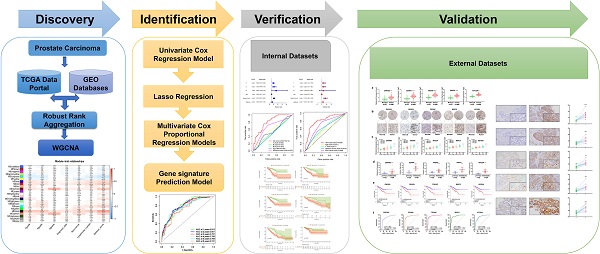
Methods: We identified differentially expressed genes (DEGs) and clinicopathological data of PCa patients from Gene Expression Omnibus (GEO) datasets and The Cancer Genome Atlas (TCGA) repositories. We then uncovered the most relevant clinical traits and genes modules associated with PCa prognosis using the Weighted gene correlation network analysis (WGCNA). Univariate Cox regression analysis and multivariate Cox proportional hazards (Cox-PH) models were performed to identify candidate gene signatures related to Disease-Free Interval (DFI). Data for internal and external cohorts were utilized to test and validate the accuracy and clinical utility of the prognostic models.
Results: We constructed and validated an accurate and reliable model for predicting the prognosis of PCa using 5 Gleason score-associated gene signatures (ZNF695, CENPA, TROAP, BIRC5 and KIF20A). The ROC and Kaplan-Meier analysis revealed the model was highly accurate in diagnosing and predicting the recurrence and metastases of PCa. The accuracy of the model was validated using the calibration curves based on internal TCGA cohort and external GEO cohort. Using the model, patients could be prognostically stratified in to various groups including TNM classification and Gleason score. Multivariate analysis revealed the model could independently predict the prognosis of PCa patients and its utility was superior to that of clinicopathological characteristics. In addition, we fund the expression of the 5 gene signatures strongly and positively correlated with tumor purity but negatively correlated with infiltration CD8+ T cells to the tumor microenvironment.
Conclusions: A 5 gene signatures can accurately be used in the diagnosis and prediction of PCa prognosis. Thus this can guide the treatment and management prostate adenocarcinoma.
Keywords: prostate cancer, gene signatures, robust rank aggregation, Weighted Gene Co-expression Network Analysis, disease-free interval, inflammation landscape

 Global reach, higher impact
Global reach, higher impact