Impact Factor ISSN: 1837-9664
J Cancer 2021; 12(12):3678-3685. doi:10.7150/jca.57114 This issue Cite
Research Paper
Blood leukocytes methylation levels analysis indicate methylated plasma test is a promising tool for colorectal cancer early detection
1. Department of Gastroenterology, Affiliated Hospital of Xuzhou Medical University, Xuzhou Jiangsu 221002, China
2. Institute of Digestive Diseases, Xuzhou Medical University, Xuzhou Jiangsu 221002, China
3. Zhejiang University Kunshan Biotechnology Laboratory, Zhejiang University Kunshan Innovation Institute, Kunshan Jiangsu 215300, China
4. State Key Laboratory of Bioelectronics, School of Biological Science and Medical Engineering, Southeast University, Nanjing 210009, China
5. Suzhou Institute of Biomedical Engineering and Technology, Chinese Academy of Sciences, Suzhou Jiangsu 215163, China
6. Suzhou VersaBio Technologies Co. Ltd., Kunshan Jiangsu 215300, China
† These authors contributed equally to this work.
Abstract
Background: A number of plasma methylated DNA biomarkers related to colorectal cancer (CRC) have been identified. However, the effect of methylation level in leukocytes on plasma-based methylation test was rarely reported.
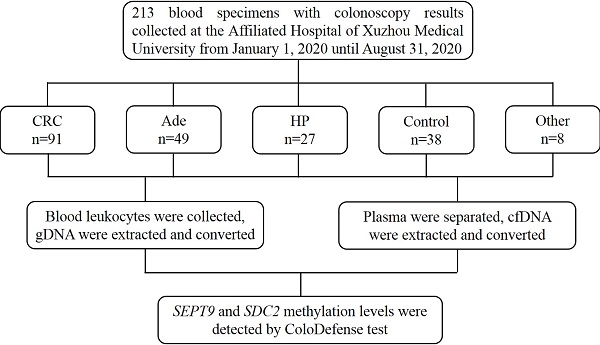
Methods: Blood samples from 213 individuals including 91 CRC patients were collected and separated into 3.5 mL of plasma and paired leukocyte fractions. DNA were extracted from plasma and leukocytes and bisulfite converted, followed by ColoDefense test that detects methylated SEPT9 (mSEPT9) and methylated SDC2 (mSDC2) simultaneously in a single qPCR reaction.
Results: Both mSEPT9 and mSDC2 levels in leukocytes exhibited no significant difference among CRC, benign tumors and healthy controls. However, mSEPT9 and mSDC2 levels in plasma were significantly higher in CRC group than those in other groups. The sensitivities of mSEPT9 and mSDC2 alone for detecting CRC with plasma samples were 75.8% and 60.4% with specificities of 94.7% and 86.8%, respectively. These two markers in combination exhibited an improved sensitivity of 85.7% for CRC detection with a specificity of 86.8%, mostly attributable to increased sensitivity of 81.8% for detecting stage 0-II CRC. AUC values for mSEPT9 and mSDC2 alone were 0.864 (95% CI: 0.798 - 0.929) and 0.796 (95% CI: 0.719 - 0.874), respectively, but improved to 0.972 (95% CI: 0.949 - 0.996) when combined for ColoDefense test.
Conclusions: The leukocytes gDNA will not affect the performance of plasma ColoDefense test, and plasma ColoDefense test exhibited high sensitivity and specificity in a validation set, demonstrating its potential as a non-invasive and cost-effective method for CRC early detection.
Keywords: Colorectal cancer, plasma, leukocyte, mSEPT9, mSDC2, early detection

 Global reach, higher impact
Global reach, higher impact