3.2
Impact Factor
ISSN: 1837-9664
J Cancer 2021; 12(12):3715-3725. doi:10.7150/jca.51059 This issue Cite
Research Paper
A Novel Set of Immune-associated Gene Signature predicts Biochemical Recurrence in Localized Prostate Cancer Patients after Radical Prostatectomy
1. Department of Urology, The First Affiliated Hospital of Nanjing Medical University, Nanjing, China.
2. The Affiliated Kezhou People's Hospital of Nanjing Medical University, Kezhou, Xinjiang, China.
*These authors contributed equally to this work.
Abstract
Background: Decision-making regarding biochemical recurrence (BCR) in localized prostate cancer (PCa) patients after radical prostatectomy (RP) mainly relies on clinicopathological parameters with a low predictive accuracy. Currently, accumulating evidence suggests that immune-associated genes (IAGs) play irreplaceable roles in tumorigenesis, progression and metastasis. Considering the critical role of immune in PCa, we therefore attempted to identify the novel IAGs signature and validate its prognostic value that can better forecast the risk for BCR and guide clinical treatment.
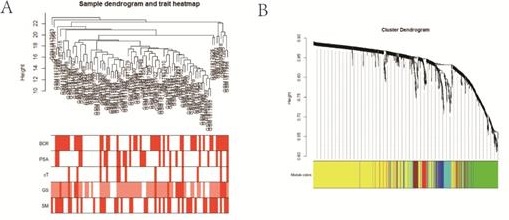
Methods: RNA-sequencing and corresponding clinicopathological data were downloaded from the Gene Expression Omnibus (GEO) database and the Cancer Genome Atlas (TCGA) database. Weighted gene co-expression network analysis (WGCNA) was utilized to screen out the candidate module closely related to BCR, and univariate and LASSO Cox regression analyses were performed to build the gene signature. Kaplan-Meier (KM) survival analysis, time-dependent receiver operating curve (ROC), independent prognostic analysis and nomogram were also applied to evaluate the prognostic value of the signature. Besides, Gene ontology analysis (GO), Kyoto encyclopedia of genes and genomes (KEGG) and gene set enrichment analysis (GSEA) were used to explore potential biological pathways.
Results: A total of six IAGs (SSTR1, NFATC3, NRP1, TUBB3, IL1R1, GDF15) were eventually identified and used to establish a novel IAGs signature. The Kaplan-Meier analysis revealed that patients with low-risk scores had longer recurrence-free survival (RFS) than those with high-risk scores in both GSE70769 and TCGA cohorts. Further, our signature was also proven to be a valuable independent prognostic factor for BCR. We also constructed a nomogram based on the gene signature and related clinicopathologic features, which excellently predict 1-year, 3-year and 5-year prognosis of localized PCa patients after RP. Moreover, functional enrichment analysis demonstrated the vital biological processes, and stratified GSEA revealed that a crucial immune-related pathway (T cell receptor signaling pathway) was notably enriched in the high-risk group.
Conclusions: We successfully developed a novel robust IAGs signature that is powerful in BCR prediction in localized PCa patients after RP, and created a prognostic nomogram. In addition, the signature might help clinicians in selecting high-risk subpopulation, predicting survival status of patients and promoting more individualized therapies than traditional clinical factors.
Keywords: immune-associated gene signature, prostate cancer, biochemical recurrence, radical prostatectomy, prognosis

 Global reach, higher impact
Global reach, higher impact