Impact Factor
ISSN: 1837-9664
J Cancer 2021; 12(14):4264-4276. doi:10.7150/jca.53831 This issue Cite
Research Paper
Development and Validation of an Immune-Related Long Non-coding RNA Prognostic Model in Glioma
1. Beijing University of Chinese Medicine Third Affiliated Hospital, Beijing, China.
2. Institute of Acupuncture and Moxibustion, Shaanxi University of Chinese Medicine, Shaanxi, China.
#These authors contributed equally to this work.
Abstract
Background: Long non-coding RNAs (lncRNAs) play an important role in the immune processes of glioma. Immune related lncRNAs (IRlncRs) may be a critical prognosis in patients with glioma. The current study aimed to construct a glioma immune-related prognosis model by IRlncRs.
Methods: Transcriptome RNA-sequencing data of glioma were obtained from The Cancer Genome Atlas (TCGA) and an immune‑related risk score (IRRS) model was constructed by Lasso and multivariate Cox regression analysis. Receiver Operating Characteristic (ROC) curves were used to assess the sensitivity and specificity of the prognosis on IRRS. A predictive nomogram and a time-dependent ROC curve was performed in training and validation cohort. We explored the relationships between survival‑related IRlncRs (sIRlncRs) and clinicopathologic parameters. Functional annotation of the sIRlncRs was investigated by gene set enrichment analysis (GSEA) and principal component analysis (PCA). The relationships between IRRS model and immune cell infiltration and co-expression network analysis among the sIRlncRs were performed for molecular mechanism study.
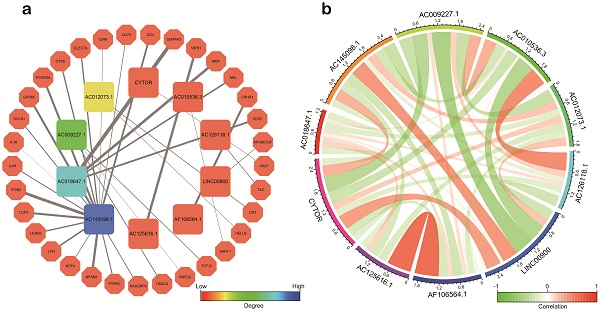
Results: A total of 10 sIRlncRs were enrolled to build IRRS model. The IRRS was identified as an independent prognostic factor and correlated with the overall survival (AUC =0.880). The nomogram was constructed successfully with IRRS, age and grade as variables. Immune cell infiltration analysis indicated that B cells, neutrophil, dendritic and macrophage cells were positively correlated with IRRS. PCA and GSEA illustrated that the lncRNA signature enrolled the IRRS model was closely related to immune status. Additionally, co-expression network showed that there was a strong correlation between 10 sIRlncRs at the transcriptional level.
Conclusion: We successfully constructed a remarkable clinical model of sIRlncRs with potential prognostic value for glioma patients, which provides an insight into immunological research and treatment strategies of glioma.
Keywords: glioma, immune, LncRNA, prognosis

 Global reach, higher impact
Global reach, higher impact