Impact Factor ISSN: 1837-9664
J Cancer 2021; 12(14):4408-4417. doi:10.7150/jca.52993 This issue Cite
Research Paper
Mutational Landscape of PI3K-AKT-mTOR Pathway in Breast Cancer: Implications for Targeted Therapeutics
1. Department of Breast Cancer, Cancer Center, Guangdong Provincial People's Hospital, Guangdong Academy of Medical Sciences, Guangzhou, China.
2. Department of Breast, Foshan Women and Children Hospital, Foshan, China.
3. Burning Rock Biotech, Guangzhou, China.
# These authors made equal contributions.
Received 2020-9-7; Accepted 2021-1-17; Published 2021-5-27
Abstract
Background: Comprehensive analysis of PI3K-AKT-mTOR pathway gene alterations in breast cancer may be helpful for targeted therapy.
Methods: We performed targeted sequencing using a panel of 520 cancer-related genes to investigate gene alterations in the PI3K-AKT-mTOR pathway from 589 consecutive Chinese women diagnosed with stage I-III breast cancer. Analyses of overall survival (OS) were performed using the publicly available clinical and genomic data from METABRIC.
Results: PI3K-AKT-mTOR pathway gene alterations were detected in 62.6% (369/589) of our cohort. The most commonly altered genes were PIK3CA (45%), PTEN (7.5%), AKT1 (5.9 %), PIK3R1 (2.7%), and PIK3CG (2%). Four PIK3CA mutations (E545K, H1047R, E542K, and H1047L) were detected in all the breast cancer molecular subtypes. Seven PIK3CA mutations (E545G, E418_L422delinsV, E726K, E110del, G1049R, G118D, and D350G) were only detected in HR+ subtypes. Two PIK3CA mutations (C420R and N345K) were only detected in non-triple-negative subtypes. Most cases with PTEN mutation were HR+/HER2- subtype (77.3%), followed by triple-negative subtype (18.2%). In the METABRIC breast cancer dataset, no significant OS difference was observed between the PIK3CA-mutant and wild-type groups. However, patients with multiple PIK3CA mutations (mOS: 131 vs. 159 months, P= 0.029), or PIK3CA mutations located in the C2 domain had significantly shorter OS (mOS, 130 vs. 154 months, P=0.020) than those without the mutations.
Conclusions: Our study reveals the heterogeneity in PI3K-AKT-mTOR pathway among the breast cancer molecular subtypes in our cohort. Moreover, the number and specific sites of PIK3CA mutations have distinct prognostic impact.
Keywords: Breast cancer, PI3K-AKT-mTOR pathway, gene alteration, molecular subtypes, prognosis
Introduction
The PI3K-AKT-mTOR pathway is among the most common activation signals in various cancer types [1-3]. This pathway is involved in the regulation of cell proliferation, apoptosis, metabolism, migration, and invasion [4, 5]. The activation of this pathway begins with the binding of the ligand to the corresponding receptor tyrosine kinase, phosphoinositide 3-kinase (PI3K). The regulatory subunit of PI3K becomes phosphorylated and leads to the activation of p110, its catalytic subunit. The activated PI3K produces phosphatidylinositol 3, 4, 5-triphosphate (PIP3), a lipid second messenger, and leads to further activation of downstream effectors such as protein kinase B (AKT) and mammalian target of rapamycin (mTOR) [4]. On the other hand, phosphatase and tensin homolog (PTEN) counteracts the function of PI3K by dephosphorylating PIP3 and is regarded as the major natural inhibitor of the PI3K-AKT-mTOR pathway [4].
Breast cancer is the most common malignancy in women worldwide. The PI3K-AKT-mTOR signaling pathway plays an important role in the development of breast cancer [6-9]. Various inhibitors have been developed for targeting the three major components of the PI3K-AKT-mTOR pathway and have shown promising therapeutic effects in breast cancer [7, 10]. For example, a specific inhibitor of PI3Kα, alpelisib (BLY719) in combination with fulvestrant in the SOLAR-1 clinical trial had shown encouraging results by prolonging progression-free survival (PFS) in patients with advanced PIK3CA-mutant breast cancer as compared with fulvestrant alone [11]. Moreover, the results of the PAKT trial confirmed that the addition of the AKT inhibitor capivasertib to first-line paclitaxel therapy can significantly prolong PFS and overall survival (OS) in patients with triple-negative breast cancer [12]. Similarly, combination therapy with mTOR inhibitor and exemestane was also shown to significantly prolong the OS of patients with hormone receptor (HR)-positive metastatic breast cancer [13, 14]. Therefore, targeting the PI3K-AKT-mTOR signaling pathway has become promising therapeutic option in the clinical treatment of breast cancer. With the advent of novel promising PI3K-AKT-mTOR pathway inhibitors, it is important to identify the patients who can benefit from these inhibitors. The comprehensive analysis of the PI3K-AKT-mTOR pathway gene aberrations in breast cancer may help choose the proper inhibitors to precisely treat certain molecular subset of patients. Next-generation sequencing (NGS) technology has made it possible to quickly and accurately identify such genetic aberrations. In this study, we systematically analyzed the genetic aberrations in the PI3K-AKT-mTOR pathway in a large cohort of 589 Chinese women with early-stage breast cancer by NGS methods.
Materials and methods
Patients and specimens
Women diagnosed with breast cancer (American Joint Committee on Cancer (AJCC) Tumor, Node, Metastasis (TNM) stage IA to III) at the Department of Breast Cancer in Guangdong Provincial People's Hospital who had tissue samples submitted for NGS were included in this study. Breast tumor tissues were obtained by surgical resection and processed as formalin-fixed paraffin-embedded (FFPE) blocks. Expression status for estrogen receptor (ER), progesterone receptor (PR), and human epidermal growth factor receptor 2 (HER2) were examined according to the guidelines of the American Society of Clinical Oncology (ASCO) /College of American Pathologists (CAP) [15]. All tumor samples were reviewed by breast pathologists, and clinical data were obtained from medical records. This study was approved by the institutional review board of the Guangdong Provincial People's Hospital. Written informed consent were provided by all patients.
Targeted sequencing and analysis
Sample processing, NGS library construction and subsequent sequencing analysis were performed at Burning Rock Biotech, a CAP-accredited and Clinical Laboratory Improvement Amendments (CLIA)-certified clinical laboratory as described previously [16]. In brief, tumor tissue DNA were extracted from FFPE tumor samples using QIAamp DNA FFPE tissue kit (Qiagen, Hilden, Germany), according to the manufacturer's instructions. The DNA concentration was quantified by Qubit 2.0 fluorimeter with the dsDNA high sensitivity assay kit (Life Technologies, Carlsbad, CA, USA). A minimum of 50 ng DNA was used for the NGS library construction. Indexed samples were sequenced on an Illumina NextSeq 500 instrument (Illumina, Inc., Hayward, CA, USA) with paired-end reads and average sequencing depth of 1,000X using a commercially available panel targeting 520 cancer-related genes, spanning 1.64 megabases (Mb) of the human genome (OncoScreen Plus, Burning Rock Biotech, Guangzhou, China). The panel was designed to capture whole exons of 312 genes and critical exons, introns, and promoter regions of the remaining 208 genes. The panel interrogates genes included in the PI3K-AKT-mTOR pathway, including all exons of AKT1-3, MTOR, PIK3CA/B/G, PIK3R1-2, and PTEN and critical exons, introns, and promoter region of PIK3C2B/G, PIK3C3, PIK3CD, and PIK3R3, which allows accurate detection of single nucleotide variants (SNV) and copy number variants. Paired white blood cells from each patient were used to filter out germline mutations. Sequencing data were analyzed using proprietary computational algorithms optimized for somatic variant calling.
Sequence data analysis
Sequence data were mapped to the human genome (hg19) using Burrows-Wheeler Aligner v.0.7.10. Local alignment optimization, variant calling, and annotation were performed using GATK v.3.2 and VarScan v.2.4.3. Variants were filtered using the VarScan fpfilter pipeline, with loci with depth less than 100 filtered out. At least 5 supporting reads were required for insertion-deletion variants; while 8 supporting reads were required for SNVs to be called. According to the ExAC, 1000 Genomes, dbSNP, and ESP6500SI-V2 databases, variants with population frequency over 0.1% were grouped as single nucleotide polymorphisms and excluded from further analysis. The remaining variants were annotated with ANNOVAR and SnpEff v.3.6. DNA translocation analysis was performed using Factera v.1.4.3. Copy number variation was detected using in-house bioinformatics pipeline based on the depth of coverage data of capture intervals.
Statistical analysis
Descriptive statistics were used to summarize patient features and sequencing results, including median and range for categorical variables, mean, and standard deviation for continuous variables. Chi-square test or Fisher's exact test was used to compare the differences in demographic, clinical, and pathological characteristics in different groups. All statistical tests were two-way. P value of less than 0.05 was considered significant.
Kaplan-Meier analysis with log-rank test was performed to analyze OS data obtained from the Molecular Taxonomy of Breast Cancer International Consortium (METABRIC) [17]. OS was defined from the date of initial diagnosis until death. A total of 2,388 breast cancer patients with TNM stage I-III from the METABRIC dataset were included in the survival analysis, including TNM stage I (n = 214), stage II (n = 976), and stage III (n = 1198). P-values were adjusted with clinical data including age, tumor stage, histological subtype, menopausal status, therapy received (radiotherapy, hormone therapy, or chemotherapy).
Results
Baseline patient characteristics
Our study included 589 Chinese women between the ages of 22 and 85 years who were diagnosed with stage I-III breast cancer. The clinicopathological characteristics of the patients are listed in Table S1. A total of 321 (54.5%) patients were HR+/HER2-, 111 (18.8%) were HR+/HER2+, and 64 (10.9%) were HR-/HER2+, while 68 (11.5%) had triple-negative breast cancer (TNBC, HR-/HER2-).
Somatic mutations of PI3K-AKT-mTOR pathway
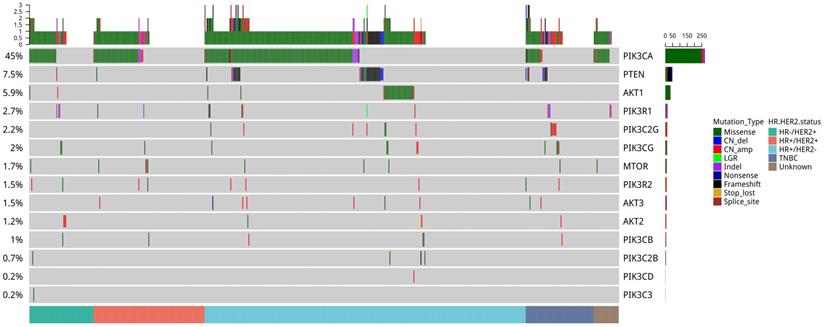
To understand the mutational pattern of genes involved in the PI3K-AKT-mTOR pathway, we analyzed the somatic mutation landscape of our breast cancer cohort. In total, 62.6% (369/589) of patients had at least one mutation in the PI3K-AKT-mTOR pathway. Among all the genes analyzed, PIK3CA was the most frequently mutated gene (45.0%, 265/589), followed by PTEN (7.5%, 44/589), AKT1 (5.9%, 35/589), PIK3R1 (2.7%, 16/589), and PIK3CG (2.0%, 12/589) (Figure 1). Mutations in other genes were less common. In the METABRIC cohort, PIK3CA was also the most frequently mutated gene (41.3%, 1035/2,509), and multiple PIK3CA mutations accounted for 15.5% (160/1,035). The frequencies of copy number amplifications in PIK3C2B and AKT3 in the METABRIC cohort were 21.6% and 20%, respectively, which were significantly higher than in our cohort. Subsequently, the mutation rates were 5.9% for PTEN, 5.0% for AKT1, and 2.7% for PIK3R1 in the METABRIC cohort, which were similar to our cohort (Figure S1).
Mutation profile of genes in the PI3K-AKT-mTOR pathway of 589 Chinese patients with breast cancer. Tumor samples were grouped according to molecular subtype: HR+/HER2- (n = 321), HR+/HER2+ (n = 111), HER2-rich (n = 64), and TNBC (n = 68) as indicated by the annotation at the bottom. The mutation frequency for each gene is shown on the left. Colors indicate the mutation types.
PIK3CA mutations
A total of 265 patients had PIK3CA mutations, including 301 mutations, with some patients harboring more than 1 (multiple) mutations. Missense mutations accounted for 93.0% (280/301) of the PIK3CA mutations. The PIK3CA mutation frequency significantly differed among the four breast cancer molecular subtypes, with 48.3% (155/321) in HR+/HER2-, 45% (50/111) in HR+/HER2+, 42.2% (27/64) in HR-/HER2+, and 25% (17/68) in HR-/HER2- patients (Table 1). The mutation sites were differentially distributed among the subtypes (Figure 2). A total of 58 PIK3CA mutations were identified in our cohort; 47 of these mutations were detected in HR+/HER2- subtype, 18 were in HR+/HER2+ subtype, 9 were in HR-/HER2+ subtype, and 7 were in the HR-/HER2- subtype. Four mutations, namely E545K, H1047R, E542K, and H1047L, were consistently detected in all four subtypes. Two mutations, C420R and N345K, were detected in three subtypes except in TNBC. Seven mutations, namely E545G, E418_L422delinsV, E726K, E110del, G1049R, G118D, and D350G, were only detected in HR+/HER2- and HR+/HER2+ subtypes. There were 40 PIK3CA mutations only detected in any one subtype. Of them, 32 mutations were only detected in HR+/HER2- subtype, 4 mutations (S499F, P449_E453delinsQ, D1017V, and P539R) were only detected in HR+/HER2+ subtype, 2 mutations (E39K and H1048L) were only detected in HR-/HER2+ subtype, and 2 mutations (A1066fs and E798N) were only detected in the HR-/HER2- subtype.
Clinicopathological features of patients according to PIK3CA mutations.
| Variables | Number of patients (n=589) | PIK3CA mutation | P-value | |
|---|---|---|---|---|
| Positive (n=265) | Negative (n=324) | |||
| Age (median 58, range 26-90) | 0.002 | |||
| ≤40 years | 130 | 43(16.2%) | 87(26.9%) | |
| >40 years | 459 | 222(83.8%) | 237(73.1%) | |
| Menopausal status | 0.004 | |||
| Premenopause | 336 | 134(50.6%) | 202(62.3%) | |
| Postmenopause | 253 | 131(49.4%) | 122(37.7%) | |
| Tumor (T) stage | 0.409 | |||
| T1 | 228 | 107(40.4%) | 121(37.3%) | |
| T2 | 327 | 146(55.1%) | 181(55.9%) | |
| T3 | 24 | 7(2.6%) | 17(5.2%) | |
| T4 | 10 | 5(1.9%) | 5(1.5%) | |
| Lymph node metastasis | 0.316 | |||
| Negative (N0) | 253 | 120(45.3%) | 133(41.0%) | |
| Positive (N1-3) | 336 | 145(54.7%) | 191(59.0%) | |
| ER status | 0.000 | |||
| Negative | 155 | 51(19.2%) | 104(32.1%) | |
| Positive | 434 | 214(80.8%) | 220(67.9%) | |
| PR status | 0.012 | |||
| Negative | 183 | 68(25.7%) | 115(35.5%) | |
| Positive | 406 | 197(74.3%) | 209(64.5%) | |
| HER-2/neu status | 0.149 | |||
| Negative | 389 | 172(64.9%) | 217(67.0%) | |
| Positive | 175 | 77(29.1%) | 98(30.2%) | |
| Equivocal | 25 | 16(6.0%) | 9(2.8%) | |
| Subtypes | 0.003 | |||
| HR+/HER2- | 321 | 155(56.5%) | 166(51.2%) | |
| HR+/HER2+ | 111 | 50(18.9%) | 61(18.8%) | |
| HR-/HER2+ | 64 | 27(10.2%) | 37(11.4%) | |
| HR-/HER2- | 68 | 17(6.4%) | 51(15.7%) | |
| Unknown | 25 | 16(6.0%) | 9(2.8%) | |
| Ki67 score | 0.001 | |||
| ≤ 14% | 137 | 80(30.2%) | 57 (17.6%) | |
| >14% | 448 | 184(69.4%) | 264(81.5%) | |
| Unknown | 4 | 1(0.4%) | 3(0.9%) | |
Distribution of PIK3CA mutation types in our cohort according to molecular subtypes. Colored boxes in the bottom indicate the five PIK3CA domains, including an adaptor binding domain (ABD), a Ras binding domain (RBD), a C2 domain, a helix domain, and a PI4K kinase catalytic domain. The distribution of mutations is according to specific mutation site. Each mutation type is indicated by color. The pie charts on the right summarize the distribution of mutation types for each molecular subtype.
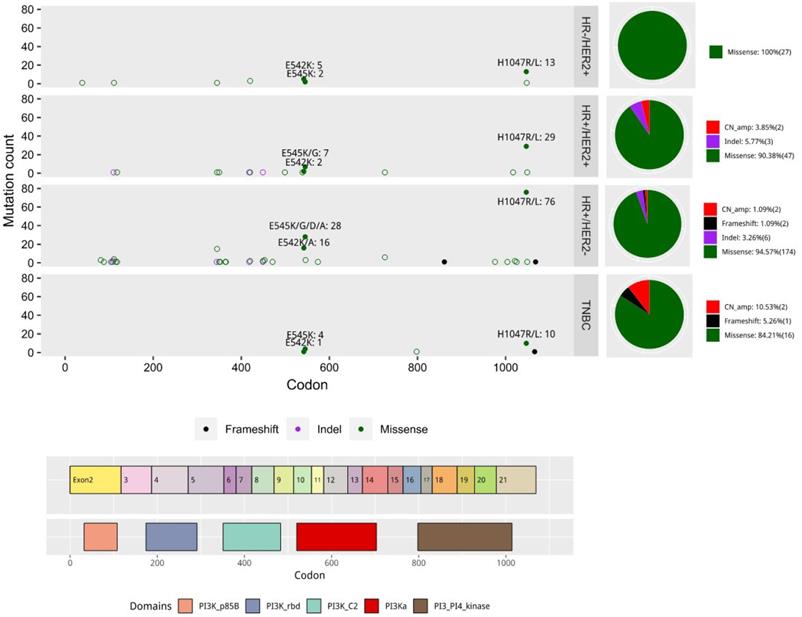
PIK3CA H1047R in exon 20 affecting the catalytic subunit of PIK3CA was the most commonly mutated site (41.9%, 126/301). Mutations in exon 9 that affect the helix binding domain, E545K (13.0%, 39/301) and E542K (7.6%, 23/301), were also identified in our cohort (Figure 2). In addition to these common and typical “major-mutant” hotspots, some “minor-mutant” sites, including N345K (6.0%), H1047L (4.0%), E726K (2.3 %), and C420R (2.0%), were also detected in our cohort.
As shown in Table 1, PIK3CA mutations were more likely to be detected in patients over 40 years old (P = 0.002), those who are postmenopausal (P = 0.004), and had ER-positive (P = 0.002), and PR-positive (P = 0.012) types of breast cancer. Interestingly, the proportion of patients with PIK3CA mutations having Ki67 scores of >14% was significantly lower than that of PIK3CA wild-type (WT) patients. However, there were no significant differences between PIK3CA-mutant and WT patients in terms of HER2 status, tumor size, and lymph node metastasis (Table 1).
Double PIK3CA-mutant tumors are frequent in breast cancer
As described above, 265 PIK3CA-mutant tumors were identified in our cohort. Among them, 34 (12.8%) cases harbored more than one PIK3CA mutations, and a vast majority (94.1%, 32/34) of such tumors had two PIK3CA mutations. In addition, the most common double PIK3CA mutation detected in our cohort consisted of a typical hotspot mutation (such as E542, E545, or H1047) and a second minor site. Tumors that harbored multiple PIK3CA mutations were HR-positive; of which, 5.9% (2/34) had HR+/HER2+ subtype.
PTEN mutations
A total of 48 PTEN mutations were detected from 44 patients, with 4 patients detected with double PTEN mutations. Frame-shift mutations (50%, 24/48) and missense mutations (16.7%, 8/48) were the two most common PTEN mutation types. The mutation frequencies of PTEN were also significantly different across breast cancer molecular subtypes. PTEN mutations were predominantly detected among HR+/HER2- subtype (77.3%, 34/44), followed by TNBC subtype (18.2%, 8/44). Only a very low percentage of PTEN mutated cases were HR-/HER2+ (2.27%, 1/44) and HR+/HER2+ (2.27%, 1/44) subtypes. No specific hotspot mutations were observed for PTEN, reflecting the diversity of mutation sites (Figure S2). Breast cancers with PTEN mutations were more likely to be HER2-negative. However, no significant differences were observed between PTEN-mutant and WT patients in terms of age, ER status, PR status, tumor size, lymph node metastasis, and Ki67 scores (Table 2).
Clinicopathological features of patients according to PTEN mutations.
| Variables | Number of patients (n=589) | PTEN mutation | P-value | |
|---|---|---|---|---|
| Positive (n=44) | Negative (n=545) | |||
| Age (median 58, range 26-90) | 0.705 | |||
| ≤40 years | 130 | 8(18.2%) | 122(22.4%) | |
| >40 years | 459 | 36(81.8%) | 423(77.6%) | |
| Menopausal status | 0.429 | |||
| Premenopause | 336 | 28(63.6%) | 308(56.5%) | |
| Postmenopause | 253 | 16(36.4%) | 237(43.5%) | |
| Tumor (T) stage | 0.622 | |||
| T1 | 228 | 16(36.4%) | 212(38.9%) | |
| T2 | 327 | 25(56.8%) | 302(55.4%) | |
| T3 | 24 | 3(6.8%) | 21(3.9%) | |
| T4 | 10 | 0(0.0%) | 10(1.8%) | |
| Lymph node metastasis | 0.268 | |||
| Negative (N0) | 253 | 15(34.1%) | 238(43.7%) | |
| Positive (N1-3) | 336 | 29(65.9%) | 307(56.3%) | |
| ER status | 0.72 | |||
| Negative | 155 | 10(22.7%) | 145(26.6%) | |
| Positive | 434 | 34(77.3%) | 400(73.4%) | |
| PR status | 0.402 | |||
| Negative | 183 | 11(25.0%) | 172(31.6%) | |
| Positive | 406 | 33(75.0%) | 373(68.4%) | |
| HER-2/neu status | ||||
| Negative | 389 | 42(95.5%) | 347(63.7%) | 0.000 |
| Positive | 175 | 2(4.5%) | 173(31.7%) | |
| Equivocal | 25 | 0(0.0%) | 25(4.6%) | |
| Subtypes | 0.002 | |||
| HR+/HER2- | 321 | 34(77.3%) | 287(52.7%) | |
| HR+/HER2+ | 111 | 1(2.3%) | 110(20.2%) | |
| HR-/HER2+ | 64 | 1(2.3%) | 63(11.6%) | |
| HR-/HER2- | 68 | 8(18.2%) | 60(11.0%) | |
| Unknown | 25 | 0(0.0%) | 25(4.6%) | |
| Ki67 score | 0.385 | |||
| ≤ 14% | 137 | 11(25.0%) | 126(23.1%) | |
| >14% | 448 | 32(72.7%) | 416(76.3%) | |
| Unknown | 4 | 1(2.3%) | 3(0.6%) | |
AKT1 mutations
In our cohort, a total of 35 patients were identified to harbor 36 AKT1 mutations. The majority (94.3%, 33/35) of AKT1 mutations were detected among patients with HR+/HER2- breast tumor. AKT1 mutations were predominantly missense mutations (91.7%, 33/36), and the remaining 8.3% (3/36) were copy number amplification. AKT1 E17K was the hotspot mutation, accounting for 86.1% (31/36, Figure S3). Breast cancers with AKT1 mutations were more likely to be ER-positive (P = 0.016), PR-positive (P = 0.002), and HER2-negative (P = 0.001). However, patients with AKT1 mutations did not statistically differ from AKT1 WT patients in terms of age, tumor size, lymph node metastasis, and Ki67 score (Table 3).
Clinicopathological features of patients according to AKT1 mutations.
| Variables | Number of patients (n=589) | AKT1 mutation | P-value | |
|---|---|---|---|---|
| Positive (n=35) | Negative (n=554) | |||
| Age (median 58, range 26-90) | 0.205 | |||
| ≤40 years | 130 | 11(31.4%) | 435(78.5%) | |
| >40 years | 459 | 24(68.6%) | 119(21.5%) | |
| Menopausal status | 0.164 | |||
| Premenopause | 336 | 24(68.6%) | 312(56.3%) | |
| Postmenopause | 253 | 11(31.4%) | 242(43.7%) | |
| Tumor (T) stage | 0.157 | |||
| T1 | 228 | 19(54.3%) | 209(37.7%) | |
| T2 | 327 | 15(42.9%) | 312(56.3%) | |
| T3 | 24 | 0(0.0%) | 24(4.3%) | |
| T4 | 10 | 1(2.9%) | 9(1.6%) | |
| Lymph node metastasis | 0.168 | |||
| Negative (N0) | 253 | 11(31.4%) | 242(43.7%) | |
| Positive (N1-3) | 336 | 24(68.6%) | 312(56.3%) | |
| ER status | 0.016 | |||
| Negative | 155 | 3(8.6%) | 152(27.4%) | |
| Positive | 434 | 32(91.4%) | 402(72.6%) | |
| PR status | 0.002 | |||
| Negative | 183 | 3(8.6%) | 180(32.5%) | |
| Positive | 406 | 32(91.4%) | 374(67.5%) | |
| HER-2/neu status | 0.001 | |||
| Negative | 389 | 33(94.3%) | 356(64.3%) | |
| Positive | 175 | 2(5.7%) | 173(31.2%) | |
| Equivocal | 25 | 0(0.0%) | 25(4.5%) | |
| Subtypes | 0.002 | |||
| HR+/HER2- | 321 | 33(94.3%) | 288(52.0%) | |
| HR+/HER2+ | 111 | 2(5.7%) | 111(20.0%) | |
| HR-/HER2+ | 64 | 0(0.0%) | 62(11.2%) | |
| HR-/HER2- | 68 | 0(0.0%) | 68(12.3%) | |
| Unknown | 25 | 0(0.0%) | 25(4.5%) | |
| Ki67 score | 0.385 | |||
| ≤ 14% | 137 | 15(42.9) | 122(22.0) | |
| >14 | 448 | 20(57.1) | 428(77.3) | |
| Unknown | 4 | 0(0.0) | 4(0.7) | |
Coexistence of PIK3CA, PTEN, and AKT1 mutations
As shown in Figure S4, 5.3% of patients (14 of 265) with PIK3CA mutation also had concurrent PTEN mutation; 1.1% of patients (3 of 265) with PIK3CA mutation also had concurrent AKT1 mutation. No patients were detected with concurrent PTEN and AKT1 mutations.
Clinical significance of PI3K-AKT-mTOR pathway mutations
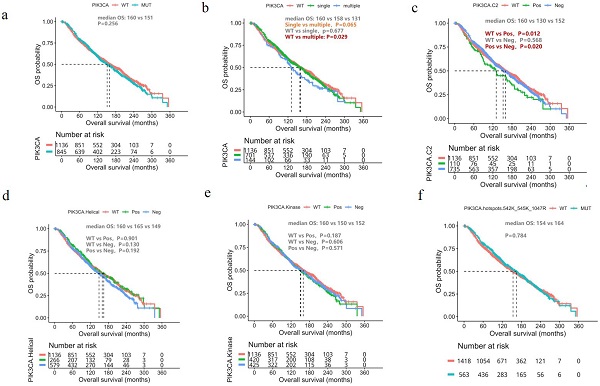
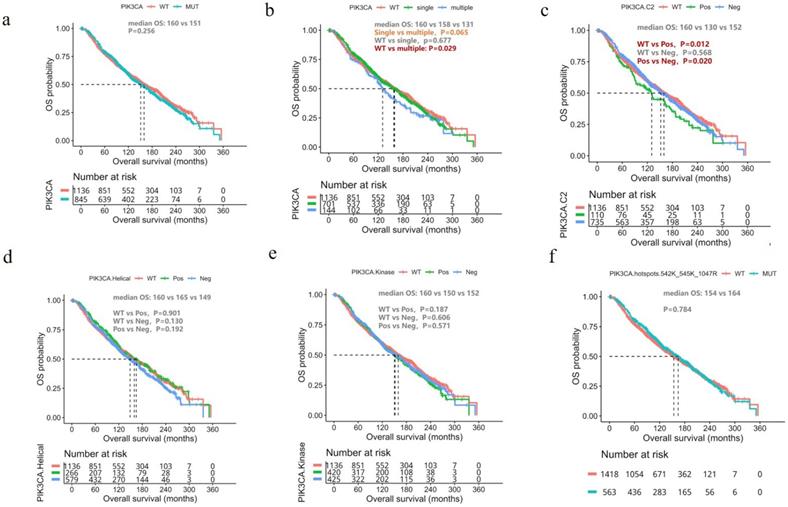
Since survival data for our cohort is unavailable, we analyzed the molecular and survival data of the breast cancer cohort from the METABRIC dataset [17]. In general, OS was similar between patients with and without PIK3CA mutations (mOS, 152 vs. 159 months, P=0.256, Figure 3A). We then divided the patients with PIK3CA mutations into two groups according to the number of PIK3CA mutations they harbored: those with single and multiple PIK3CA mutations. Patients with multiple PIK3CA mutations had significantly shorter OS than those with PIK3CA WT (mOS, 131 vs. 159 months, P=0.029, Figure 3B) and a trend of shorter OS, but statistical significance was not reached, than those with single PIK3CA mutation (mOS, 131 vs. 162 months, P=0.065, Figure 3B). We also performed a subgroup analysis based on the location of the PIK3CA mutations. Interestingly, we found that patients with PIK3CA mutations located in the C2 domain had significantly shorter OS than patients with non-C2 domain mutations (mOS, 130 vs. 154 months, P=0.020, Figure 3C). However, there was no significant difference in prognosis between patients with non-C2 domain mutations and patients with WT PIK3CA (mOS, 152 vs. 160 months, P=0.568, Figure 3C). There was no difference observed in survival between patients with PIK3CA mutations located in either the helical domain (Figure 3D) or kinase domain (Figure 3E). Finally, patients with PIK3CA H1047R mutations also had significantly shorter OS than patients with non-H1047 mutations (mOS, 152 vs. 157 months, P=0.021, Figure 3F).
Prognostic impact of PIK3CA mutations. Kaplan Meier curves comparing the overall survival of the breast cancer cohort from the METABRIC dataset according to (A) The presence/absence of PIK3CA mutation (PIK3CA-mut vs. PIK3CA-WT); (B) The number of PIK3CA mutations (multiple, single, WT); (C) Mutations located in the PIK3CA-C2 domain vs. non-C2 domain; (D) Mutations located in the helical domain vs. non-helical domain; (E) Mutations in the kinase domain vs. non-kinase domain; (F) H1047R vs. non-H1047R mutations. WT, wild-type.
Discussion
In this study, we performed a comprehensive analysis of the genetic aberrations associated with the PI3K-AKT-mTOR pathway in a large cohort of Chinese women with early-stage breast cancer (n = 589). Based on our results, PIK3CA was the most frequently mutated gene (45.0%), which is consistent with a similarly high frequency (43.6%) reported by Chen et al. [9], but numerically higher than the data from the Cancer Genome Atlas (TCGA) breast cancer dataset (36%) [20]. Similar to previous reports [9, 20], the PIK3CA hotspot mutations observed in our cohort were also H1047R, E545K, and E542K mutations. Among them, the H1047R substitution in exon 20 accounted for 41.9% (126/301), the E545K substitution in exon 9 accounted for 13.0% (39/301), and the E542K substitution in exon 9 accounted for 7.6% (23/301) of all the PIK3CA mutations. Since H1047 is in the kinase domain and near the C-terminus of the activation loop, the substitution of histidine (H) to arginine (R) at this position may alter the conformation of the activation loop and result in the activation of the PI3K pathway [21, 22]. H1047R mutation is sufficient to induce tumors in animal models and may be associated with a favorable response to PI3K inhibitors [21, 22]. On the other hand, E545 is within the helical domain and interacts with the N-terminal SH2 domain of the p85 regulatory subunit [23]. The change from glutamic acid (E) to lysine (K) causes charge reversal and disrupts its inhibitory interaction with p85 [24]. In addition to the three common and typical “major-mutant” hotspots (E542K, E545K, or H1047R), we also identified some “minor-mutant” sites, including N345K, H1047L, E726K, and C420R. Consistent with our findings, Martínez-Sáez et al. previously reported a PIK3CA mutation rate of 35.7% in their breast cancer cohort, with H1047R (35%), E545K (17%), E542K (11%), N345K (6%), and H1047L (4%) as the most common mutation sites [25]. H1047L and N345K mutations are located in the kinase domain with relatively higher mutation frequency, and N345K and C420R mutations are located in the PIK3CA C2 domain, which are relatively rare.
In addition, we found that 12.8% of patients had two or more PIK3CA mutations. This data is also consistent with a previous report by Vasan et al [26]. Analysis of 3,740 PIK3CA-mutated tumors identified about 12% of patients who harbored multiple PIK3CA mutations [26]. The frequency of multiple PIK3CA mutations in our cohort is also largely consistent with those reported in the breast cancer cohort of the METABRIC dataset (13%) [27] and the TCGA dataset (11%) [20]. In the METABRIC and TCGA datasets, a vast majority of breast tumors (88% to 96%) with multiple PIK3CA mutations also had double mutations. Through in vivo and in vitro experiments, Vasan et al. demonstrated that the presence of double PIK3CA mutations on the same allele resulted in increased PI3K activity, which led to enhanced downstream signal transduction, cell proliferation, and tumor growth [26]. The molecular mechanism of PIK3CA double mutation includes increasing the destruction of the binding of p110α to the inhibitory subunit p85α, thereby reducing its catalytic inhibitory effect and enhancing the binding of p110α membrane lipids. As compared to single mutations, double PIK3CA mutations activate PI3K signaling pathway to a higher degree and promote the malignant progression of tumors [26]. This can explain, to a certain extent, the poorer prognosis of patients with multiple PIK3CA mutations than those with only one PIK3CA mutation.
In our cohort, women with PIK3CA mutations were more likely to be older than 40 years of age (P = 0.002), postmenopausal (P = 0.004), ER-positive (P = 0.002), and PR-positive (P = 0.012). Among patients with PIK3CA mutations, the proportion of patients with Ki67 score of >14% was significantly lower than that of WT patients (Table 1). However, there were no significant differences between patients with or without PIK3CA mutations in terms of HER2 status, tumor size, and lymph node metastasis. Consistent with our findings, a pooled analysis of 10,319 women with early-stage breast cancer from 19 studies showed a PIK3CA mutation rate of 32% with the detection of PIK3CA mutations significantly associated with ER-positive status, increased age, lower tumor grade, and smaller body size (all P <0.001) [28]. Tao et al. sequenced the plasma of 234 Chinese women with metastatic breast cancer and reported a PIK3CA mutation frequency of 31%, which were significantly higher in HR+/HER2+, HR+/HER2-, and HR-/HER2+ subtypes and significantly less common in TNBC (P<0.01) [29]. Our results also found that PIK3CA mutations were the least in TNBC than the other subtypes.
In the PI3K-AKT-mTOR pathway, our study identified PTEN as the second most commonly mutated gene. PTEN mutations occurred in 7.5% (44/589) of patients, and the incidence rate of double-PTEN mutations was 9.09% (4/44). Millis et al. reported a PTEN mutation rate of 6% across 19,784 diverse solid tumors [1], while the PTEN mutation rate was about 10% in the TCGA cohort [20]. However, the incidence of negative PTEN expression using immunohistochemistry of breast tumors was reported to be as high as 30% [1], which suggests that somatic mutations in PTEN are only a small proportion of breast tumors with PTEN loss-of-function. In our cohort, we observed a difference in the subtype distribution of PTEN and PIK3CA mutations. PTEN mutations occurred most frequently in HR+/HER2- subtype, and second most in TNBC subtype. In addition, for PTEN mutations also lack typical hotspot mutations. Since PTEN is a tumor suppresser gene, any loss-of-function mutation would inactivate it in tumors. However, in the case of PIK3CA, only a few gain-of-function mutations would be selected during tumorigenesis. Another interesting observation in our cohort is that breast cancers with PTEN mutations were more likely to be HER2-negative, while no significant differences between patients with/without PTEN mutations in terms of age, ER status, PR status, tumor size, lymph node metastasis, and Ki67 score.
AKT1 (5.9%) ranks as the third most commonly mutated gene in the PI3K-AKT-mTOR signaling pathway in our cohort. Almost all (94.3%, 33/35) of AKT1 mutations were identified in HR+/HER2- breast tumors, with only one typical hotspot mutation. AKT1 E17K is a single amino acid mutation in the pleckstrin homology (PH) domain of the AKT1 protein, which has been reported to function as a carcinogenic driver mutation in breast cancer and other solid cancers [18, 19]. In our cohort, breast cancers with AKT1 mutations were significantly associated with positive ER- (P = 0.016) and PR (P = 0.002) but negative HER2- (P = 0.001) status. However, women with/without AKT1 mutations were similar in terms of age, tumor size, lymph node metastasis, and Ki67 score.
The prognostic impact of PIK3CA mutation in breast cancer is still controversial. Several studies have shown that breast cancer patients with PIK3CA mutations have a better prognosis [30, 31], while Sobhani et al. reported a significantly worse prognosis for women with PIK3CA-mutant breast tumors [32]. Gerratana et al. analyzed 88 cases of metastatic breast cancer by multivariate logistic regression and found that the detection of PIK3CA mutation in circulating tumor DNA was associated with lung metastasis (OR 3.74) [33]. Our findings using the METABRIC dataset may partly explain this controversy. In our survival analysis, we found that patients with multiple PIK3CA mutations, but not those with single PIK3CA mutations, had significantly worse prognosis than patients with WT PIK3CA. In addition, we also found that patients with mutations in the PIK3CA-C2 domain had significantly worse prognosis than patients with non-C2 domain mutations (mOS, 130 vs. 154 months, P=0.020, Figure 3C). Furthermore, patients with PIK3CA H1047R mutation also had worse prognosis than patients without H1047 mutations (mOS, 152 vs. 157 months, P=0.021, Figure 3F). However, prognosis was comparable between patients with non-C2 domain mutations and WT PIK3CA (mOS, 154 vs. 159 months, P=0.568, Figure 3C). These results suggest the distinct prognostic impact of the number and position of PIK3CA mutations.
There are some limitations in our study. First, since we do not have the long-term follow-up data for our cohort, correlation analysis for long-term survival outcomes and genetic data were only performed using publicly available METABRIC dataset. Second, as our current study did not include data on protein expression or other functional studies, the actual impact of the individual gene mutation on protein products and the activation status of the PI3K-AKT-mTOR signaling pathway largely rely on speculation. Third, most of the patients in our cohort did not receive inhibitors targeting the PI3K-AKT-mTOR signaling; hence, the therapeutic implication of the analyzed gene mutations was not evaluated.
In conclusion, our study demonstrated that the genetic alteration of the PI3K-AKT-mTOR pathway gene is very common in breast cancer, with a mutation rate of 62.6%. The frequencies and types of mutations in the PI3K-AKT-mTOR pathway genes were distinct across the breast cancer molecular subtypes. In addition, survival analysis using the data from the METABRIC cohort demonstrated the prognostic impact of different mutation sites and the number of mutations in PIK3CA. Nevertheless, the therapeutic value of the PI3K-AKT-mTOR pathway using various specific inhibitors warrants further exploration and deeper investigation of the genetic mutational landscape of the pathway.
Abbreviations
AKT: protein kinase B; BC: breast cancer; ER: Estrogen receptor; FFPE: formalin-fixed paraffin-embedded; HER2: human epidermal growth factor receptor 2; HR: Hormone receptors; M: metastasis; mOS: median overall survival; mTOR: mammalian target of rapamycin; N: lymph node involvement/metastasis; NGS: next-generation sequencing; OS: overall survival; PFS: progression-free survival; PI3K: phosphoinositide 3-kinase; PIP3: phosphatidylinositol 3,4,5-triphosphate; PR: Progesterone receptor; PTEN: phosphatase and tensin homolog; RTK: receptor tyrosine kinase; T: tumor; TNBC: triple-negative breast cancer; WT: wild-type.
Supplementary Material
Supplementary figure and tables.
Acknowledgements
We thank all patients and their families who participated in this study.
Availability of supporting data
Data supporting the results of this study can be obtained from the corresponding author under reasonable request.
Ethics approval and consent to participate
Ethics approval and consent to participate: All patients signed an informed consent form to participate in the study, and this study was approved by the Institutional Review Board of GDPH.
Funding Sources
This research is funded by the National Natural Science Foundation (approval No.: 8203066), Guangzhou basic and applied basic research project (approved in 2021), special science and Technology Fund (Doctoral entrepreneurship project) of Guangdong People's Hospital and Guangdong Medical Science and Technology Research Fund (a2021080).
Author Contribution
Conceptualization: NL and WKX; Data curation: WXK, GCZ and BC; Formal analysis: WXK, GCZ and BC; Funding acquisition: NL; Investigation: all the authors; Methodology: WXK, GCZ, BC, ML, HL, JL and HHZ; Project administration: NL and WKX; Resources: WXK, GCZ and BC; Software: WXK and ML; Supervision; NL and WKX;EAR; Validation: WXK, GCZ,ML and BC; Visualization: WXK, GCZ, ML and BC; Roles/Writing - original draft: WXK, GCZ and BC ; Writing - review & editing: All authors.
Competing Interests
Min Li, Hao Liu, Jing Liu, Han Han-Zhang and Analyn Lizaso are employees of Burning Rock Biotech. All the other authors declare no potential conflicts of interest.
References
1. Millis SZ, Ikeda S, Reddy S, Gatalica Z, Kurzrock R. Landscape of Phosphatidylinositol-3-Kinase Pathway Alterations Across 19784 Diverse Solid Tumors. JAMA Oncol. 2016;2:1565-73
2. Zhang Y, Kwok-Shing Ng P, Kucherlapati M, Chen F, Liu Y, Tsang YH. et al. A Pan-Cancer Proteogenomic Atlas of PI3K/AKT/mTOR Pathway Alterations. Cancer cell. 2017;31:820-32 e3
3. Willis O, Choucair K, Alloghbi A, Stanbery L, Mowat R, Charles Brunicardi F. et al. PIK3CA gene aberrancy and role in targeted therapy of solid malignancies. Cancer Gene Ther. 2020;27:634-44
4. Fruman DA, Chiu H, Hopkins BD, Bagrodia S, Cantley LC, Abraham RT. The PI3K Pathway in Human Disease. Cell. 2017;170:605-35
5. Tateishi K, Nakamura T, Juratli TA, Williams EA, Matsushita Y, Miyake S. et al. PI3K/AKT/mTOR Pathway Alterations Promote Malignant Progression and Xenograft Formation in Oligodendroglial Tumors. Clin Cancer Res. 2019;25:4375-87
6. Dey N, De P, Leyland-Jones B. PI3K-AKT-mTOR inhibitors in breast cancers: From tumor cell signaling to clinical trials. Pharmacol Ther. 2017;175:91-106
7. Hyman DM, Smyth LM, Donoghue MTA, Westin SN, Bedard PL, Dean EJ. et al. AKT Inhibition in Solid Tumors With AKT1 Mutations. J Clin Oncol. 2017;35:2251-9
8. Toska E, Osmanbeyoglu HU, Castel P, Chan C, Hendrickson RC, Elkabets M. et al. PI3K pathway regulates ER-dependent transcription in breast cancer through the epigenetic regulator KMT2D. Science. 2017;355:1324-30
9. Chen L, Yang L, Yao L, Kuang XY, Zuo WJ, Li S. et al. Characterization of PIK3CA and PIK3R1 somatic mutations in Chinese breast cancer patients. Nat Commun. 2018;9:1357
10. Verret B, Cortes J, Bachelot T, Andre F, Arnedos M. Efficacy of PI3K inhibitors in advanced breast cancer. Ann Oncol. 2019;30(Suppl 10):x12-x20
11. Andre F, Ciruelos E, Rubovszky G, Campone M, Loibl S, Rugo HS. et al. Alpelisib for PIK3CA-Mutated, Hormone Receptor-Positive Advanced Breast Cancer. N Engl J Med. 2019;380:1929-40
12. Schmid P, Abraham J, Chan S, Wheatley D, Brunt AM, Nemsadze G. et al. Capivasertib Plus Paclitaxel Versus Placebo Plus Paclitaxel As First-Line Therapy for Metastatic Triple-Negative Breast Cancer: The PAKT Trial. J Clin Oncol. 2020;38:423-33
13. Jerusalem G, de Boer RH, Hurvitz S, Yardley DA, Kovalenko E, Ejlertsen B. et al. Everolimus Plus Exemestane vs Everolimus or Capecitabine Monotherapy for Estrogen Receptor-Positive, HER2-Negative Advanced Breast Cancer: The BOLERO-6 Randomized Clinical Trial. JAMA Oncol. 2018;4:1367-74
14. Giuliano M, Schettini F, Rognoni C, Milani M, Jerusalem G, Bachelot T. et al. Endocrine treatment versus chemotherapy in postmenopausal women with hormone receptor-positive, HER2-negative, metastatic breast cancer: a systematic review and network meta-analysis. Lancet Oncol. 2019;20:1360-9
15. Hammond ME, Hayes DF, Dowsett M, Allred DC, Hagerty KL, Badve S. et al. American Society of Clinical Oncology/College Of American Pathologists guideline recommendations for immunohistochemical testing of estrogen and progesterone receptors in breast cancer. J Clin Oncol. 2010;28:2784-95
16. Chen B, Zhang G, Wei G, Wang Y, Guo L, Lin J. et al. Heterogeneity of genomic profile in patients with HER2-positive breast cancer. Endocr Relat Cancer. 2020;27:153-62
17. Pereira B, Chin SF, Rueda OM, Vollan HK, Provenzano E, Bardwell HA. et al. The somatic mutation profiles of 2,433 breast cancers refines their genomic and transcriptomic landscapes. Nat Commun. 2016;7:11479
18. Rudolph M, Anzeneder T, Schulz A, Beckmann G, Byrne AT, Jeffers M. et al. AKT1 (E17K) mutation profiling in breast cancer: prevalence, concurrent oncogenic alterations, and blood-based detection. BMC Cancer. 2016;16:622
19. Kircher DA, Trombetti KA, Silvis MR, Parkman GL, Fischer GM, Angel SN. et al. AKT1(E17K) Activates Focal Adhesion Kinase and Promotes Melanoma Brain Metastasis. Mol Cancer Res. 2019;17:1787-800
20. Cancer Genome Atlas N. Comprehensive molecular portraits of human breast tumours. Nature. 2012;490:61-70
21. Janku F, Wheler JJ, Naing A, Falchook GS, Hong DS, Stepanek VM. et al. PIK3CA mutation H1047R is associated with response to PI3K/AKT/mTOR signaling pathway inhibitors in early-phase clinical trials. Cancer Res. 2013;73:276-84
22. Lee KH, Hwang HJ, Noh HJ, Shin TJ, Cho JY. Somatic Mutation of PIK3CA (H1047R) Is a Common Driver Mutation Hotspot in Canine Mammary Tumors as Well as Human Breast Cancers. Cancers (Basel). 2019 11
23. Hao Y, Wang C, Cao B, Hirsch BM, Song J, Markowitz SD. et al. Gain of interaction with IRS1 by p110alpha-helical domain mutants is crucial for their oncogenic functions. Cancer cell. 2013;23:583-93
24. Thirumal Kumar D, George Priya Doss C. Role of E542 and E545 missense mutations of PIK3CA in breast cancer: a comparative computational approach. J Biomol Struct Dyn. 2017;35:2745-57
25. Martinez-Saez O, Chic N, Pascual T, Adamo B, Vidal M, Gonzalez-Farre B. et al. Frequency and spectrum of PIK3CA somatic mutations in breast cancer. Breast Cancer Res. 2020;22:45
26. Vasan N, Razavi P, Johnson JL, Shao H, Shah H, Antoine A. et al. Double PIK3CA mutations in cis increase oncogenicity and sensitivity to PI3Kalpha inhibitors. Science. 2019;366:714-23
27. Curtis C, Shah SP, Chin SF, Turashvili G, Rueda OM, Dunning MJ. et al. The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature. 2012;486:346-52
28. Zardavas D, Te Marvelde L, Milne RL, Fumagalli D, Fountzilas G, Kotoula V. et al. Tumor PIK3CA Genotype and Prognosis in Early-Stage Breast Cancer: A Pooled Analysis of Individual Patient Data. J Clin Oncol. 2018;36:981-90
29. Tao Z, Li T, Feng Z, Liu C, Shao Y, Zhu M. et al. Characterizations of Cancer Gene Mutations in Chinese Metastatic Breast Cancer Patients. Front Oncol. 2020;10:1023
30. Loi S, Michiels S, Lambrechts D, Fumagalli D, Claes B, Kellokumpu-Lehtinen PL. et al. Somatic mutation profiling and associations with prognosis and trastuzumab benefit in early breast cancer. J Natl Cancer Inst. 2013;105:960-7
31. Bernichon E, Vallard A, Wang Q, Attignon V, Pissaloux D, Bachelot T. et al. Genomic alterations and radioresistance in breast cancer: an analysis of the ProfiLER protocol. Ann Oncol. 2017;28:2773-9
32. Sobhani N, Roviello G, Corona SP, Scaltriti M, Ianza A, Bortul M. et al. The prognostic value of PI3K mutational status in breast cancer: A meta-analysis. J Cell Biochem. 2018;119:4287-92
33. Gerratana L, Davis AA, Polano M, Zhang Q, Shah AN, Lin C. et al. Understanding the organ tropism of metastatic breast cancer through the combination of liquid biopsy tools. Eur J Cancer. 2021;143:147-57
Author contact
Corresponding author: Ning Liao, Department of Breast Cancer, Cancer Center, Guangdong Provincial People's Hospital, Guangdong Academy of Medical Sciences, Guangzhou, China. (Email: syliaoningedu.cn).

 Global reach, higher impact
Global reach, higher impact