Impact Factor ISSN: 1837-9664
J Cancer 2021; 12(15):4505-4512. doi:10.7150/jca.57831 This issue Cite
Review
Prognostic and Therapeutic Significance of BTN3A Proteins in Tumors
1. School of Life Science, Jiangsu Normal University, Xuzhou, Jiangsu, China.
2. College of Health Science, Jiangsu Normal University, Xuzhou, Jiangsu, China.
3. Department of Ultrasonic Medicine, Xuzhou First People's Hospital, Jiangsu, China.
# These authors contributed equally to this work.
Received 2021-1-4; Accepted 2021-5-9; Published 2021-5-27
Abstract
The Butyrophilin 3A (BTN3A) family is a type I transmembrane protein belonging to the immunoglobulin (Ig) superfamily. The family contains three members: BTN3A1, BTN3A2 and BTN3A3, which share 95% homology in the extracellular domain. The expression of BTN3A family members is different in different types of tumors, which plays an important role in tumor prognosis. Among them, there are many studies on tumor immunity of BTN3A1, which shows that it is essential for the activation of Vγ9Vδ2 T cells, while BTN3A3 is expected to become a potential therapeutic target for breast cancer. Recent studies have shown that the BTN3A family is closely related to the occurrence and development of tumors. Now the BTN3A family has become one of the research hotspots and is expected to become new tumor prediction and treatment targets.
Keywords: Butyrophilin 3A, Tumor, Prognosis
1. Introduction
1.1. Origin of the BTN3A family
The Butyrophilin (BTN) family is a type I transmembrane protein belonging to the immunoglobulin (Ig) superfamily [1-3]. It has structural homology with members of the B7 family at the extracellular domain level, that is, an Ig-like domain with IgV and IgC domains [1, 3, 4]. Butyrophilin (BTN) protein was first isolated from milk fat globule in the milk of human, bovine, goat and other species by Werner W. Franke. Because of its close relationship with yellow grease, it was named Butyrophilin (in Latin, butyro means butter) [5].
The BTN3A family, also known as CD277, is a subfamily of Butyrophilin (BTN) molecules, including BTN3A1, BTN3A2 and BTN3A3 [6, 7], which are three isomers produced by two successive duplications of the BTN3 gene [8]. These three isotypes are encoded by three different genes found in humans and some non-human primates [9]. They are expressed in most human immune cell subsets, including T cells, B cells, monocytes, dendritic cells and natural killer (NK) cells [4, 6, 10], as well as hematopoietic and non-hematopoietic tumor cell lines [11, 12]. The specific antibody 20.1 can bind to the extracellular domain of BTN3A molecule and induce the activation of Vγ9Vδ2 T cells, while antibody 103.2 can inhibit this process [13, 14].
1.2. The structure of the BTN3A family
Each member of the BTN3A family has an extracellular N-terminal IgV domain and a near-membrane IgC domain connected to a one-way transmembrane domain [13, 15, 16]. The extracellular domains of the three BTN3A subtypes are structurally similar with 95% homology [9], and there is only a slight angular difference between the IgV and IgC domains [13, 17]. The extracellular domains of BTN3A1 and BTN3A3 have 97% sequence identity [2]. There are two possible conformations in its static extracellular domain, namely, “head-tail” conformation and “V-shaped” conformation [18, 19], while “V-shaped” conformation is recognized and observed by more people.
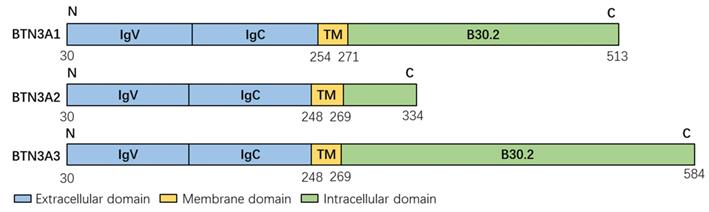
The intracellular domains of the three members of the BTN3A family are different (Figure 1). BTN3A1 and BTN3A3 have an intracellular B30.2 domain [6, 11, 20]. BTN3A1 has a unique pocket lined with basic amino acid residues, including histidine (His351 and His378), arginine (Arg412, Arg418 and Arg469) and lysine (Lys393), providing a highly positively charged environment that can complement the negative charge of the pyrophosphate portion but position 351 of the B30.2 structure of BTN3A3 is arginine [17, 21]. Apart from this, BTN3A3 has a unique C-terminalextension of 70 amino acids [17]. However, BTN3A2 lacks the B30.2 domain [6, 17, 22-24]. The reason for this phenomenon is that an Alu sequence is translocated into B30.2 exon of BTN3A2 during a stage of primate evolution, resulting in the interruption of reading frames with different intergroup lengths [25].
The structure of the members of the BTN3A family. Extracellular domain, blue; Membrane domain, yellow; Intracellular domain, green. The data in this figure is from the Universal Protein Resource (UniProt) website (https://www.uniprot.org/). TM: Transmembrane domain.
2. BTN3A1 and Cancer
2.1. BTN3A1 and prognosis of cancer
The BTN3A family is closely related to many cancers, but its prognostic role is different in different cancers. BTN3A1 is related to the prognosis of ovarian cancer, breast cancer, bladder cancer, pancreatic ductal adenocarcinoma and renal cell carcinoma. BTN3A1 was highly expressed in advanced ovarian cancer [2], but down-regulated in breast cancer [26]. The expression of BTN3A1 was positively correlated with the overall survival rate of patients with bladder cancer [27], but negatively correlated with the overall survival rate of patients with pancreatic ductal adenocarcinoma [28]. Recently, studies have shown that the response of metastatic clear cell carcinoma to nivorumab can be predicted by the level of BTN3A1 to distinguish between responders who are already at the baseline of treatment and non-responders [29].
2.2. BTN3A1 kills tumor cells through the activation of Vγ9Vδ2 T cells
2.2.1. Phosphate antigens are selectively recognized and bound by the intracellular binding domain B30.2 of BTN3A1
Vγ9Vδ2 T cells mainly exist in peripheral blood and can resist microbial infection and cancer [14, 30]. Due to the imbalance of mevalonate pathway or the accumulation of intracellular phosphate antigen (pAg) in tumor cells after microbial infection, Vγ9Vδ2 T cells can target tumor cells [31, 32]. pAgs include isopentenyl pyrophosphate (IPP), (E)-4-hydroxy-3-methyl-but-2-enyl pyrophosphate (HMBPP) and some synthetic pAgs, such as phosphorylated bromohydrin (BrHPP). The first step in this complex process is the combination of pAg and BTN3A1. Vavassori et al. believe that the V-like distal domain of BTN3A1 specifically binds phosphorylated antigens in a shallow groove and then transmits it to Vγ9Vδ2 T cells [14, 18]. However, more people think that pAg binds directly to the B30.2 domain of BTN3A1 [12, 21, 33-37], but not to the extracellular IgV-like domain. It is worth noting that when treated with 20.1 agonist antibody, BTN3A1, BTN3A2 and BTN3A3 all gave stimulation signals to Vγ9Vδ2 T cells, indicating that their extracellular domains were involved in the activation process [14, 21].
However, only BTN3A1 mediates pAg-induced activation [21], which indicates the importance of the intracellular domain B30.2. Homologues of the B30.2 domain have been found in all primates carrying Vγ9Vδ2 T cells [9]. This domain is generally considered to be a protein-protein interaction module, but their binding chaperones are various and there is no obviously conservative binding interface [21]. The B30.2 domain in the intracellular domain of BTN3A3 shares 87% homology with BTN3A1 [21], but BTN3A3 cannot stimulate Vγ9Vδ2 T cells in a pAg-dependent manner [21, 36]. The difference of a pocket residue H351 (His351) between the B30.2 domain of BTN3A1 and BTN3A3 may determine the pAg binding [21]. In addition, Gu et al. also mentioned that the pocket-side Y352 (Tyr352) in the B30.2 domain is very important for the binding of pAg to the B30.2 domain [19].
There is a positively charged surface bag in the B30.2 domain of BTN3A1, which combines a series of negatively charged small molecules, including pAg [33]. Through the nuclear magnetic resonance (NMR) experiment, Salim et al. found that BTN3A1 selectively detected pAg, to distinguish pAg from non-antigenic small molecules through the conformational antigen sensor in its B30.2 domain [33]. Through the detection of NMR spectra, Gu et al. found that the conformation of B30.2 domain in BTN3A1 cells changed after binding to pAg, and most of the residues in or near the pocket of B30.2 domain and a small part of residues located in other regions of B30.2 domain experienced chemical shift perturbation (CSP) [19]. The binding of the B30.2 domain of BTN3A1 to pAg can also induce the immobilization of BTN3A1 on the cell surface, which is also a small step in the process of stimulating Vγ9Vδ2 T cells [21]. In addition, BTN3A2 and BTN3A3 may enhance the stimulation of BTN3A1 perception by increasing BTN3A1 levels and experiencing the same extracellular domain changes as BTN3A1, but they are not absolutely necessary [38].
2.2.2. Change in the near membrane domain of BTN3A1
The change of BTN3A1 near membrane domain is also an important participant in the process of Vγ9Vδ2 T cell recognition, which strongly affects the activation of Vγ9Vδ2 T cells [39]. The change of BTN3A3 near membrane domain can significantly enhance or decrease the reactivity of γδ T cells. This region is identified as a possible dimerization interface and located near the starting position of the B30.2 domain [39], which implies the importance of the transmembrane domain. Some researchers believe that Ser/Thr296/297 and Thr304 play an important role in the near-membrane domain. The role of Ser/Thr296/297 may lie in its ability to stabilize some local folding or its ability to participate in the formation of binding interface with B30.2, while Thr304 is involved in binding HMBPP [40]. The near membrane domain is flexible in the natural state, but once B30.2 is clamped in combination with HMBPP, it is likely to cause the near membrane domain to become a rigid spiral shape consistent with the compression direction of the whole domain [40]. The transmembrane region of BTN3A1 has no specific contribution to the development of Vγ9Vδ2 T cell response [39].
The accumulation of intracellular phosphate antigen leads to the relocation of RhoB from the nucleus or nuclear membrane to the proximal region of the plasma membrane, where it can directly interact with BTN3A1 [31]. At the same time, RhoB will complete the transition from GDP-bound state to GTP-bound state, that is, from inactive state to active state [31]. RhoB can also regulate the membrane mobility of BTN3A1 on cancer cells [31]. Recently, it has been confirmed that the decrease of transmembrane protein BTN3A1 migration on target tumor cells is a key determinant of Vγ9Vδ2 T cell activation [31]. BTN3A2 regulates the subcellular localization of BTN3A1, and BTN3A2 recombines with BTN3A1 in an IgC-dependent manner [41]. If there is endogenous BTN3A1 expression in Vγ9Vδ2 T cells, but lack of BTN3A2 expression, the cells show impaired activation [41], suggesting that BTN3A2 also plays a role in the activation of Vγ9Vδ2 T cells. In addition, the full functional activity of BTN3A1 required BTN3A2, with some capacity of BTN3A3 to substitute for BTN3A2 [41].
2.2.3. The extracellular domain of BTN3A1 changed and is recognized by Vγ9Vδ2 T cells
BTN3A1 can form homodimers through its IgC domain [41]. BTN3A1 exists on the surface of cancer cells in the form of dimers [39], and can also form heterodimers with BTN3A2 [19]. RhoB interacts with BTN3A1 homodimer in cancer cells recognized by Vγ9Vδ2 T-cell receptor (TCR) [31]. Other studies have shown that the accumulation of phosphate antigen is related to the conformational change of BTN3A1 dimer [31]. The increase of intracellular phosphate antigen level can induce the extracellular change of BTN3A1 dimer, which may act as a molecular feature recognized by Vγ9Vδ2 TCR [31]. On the other hand, RhoB may dissociate from the fixed BTN3A1 after the binding of phosphorylated antigen to B30.2 and the change of extracellular conformation [31].
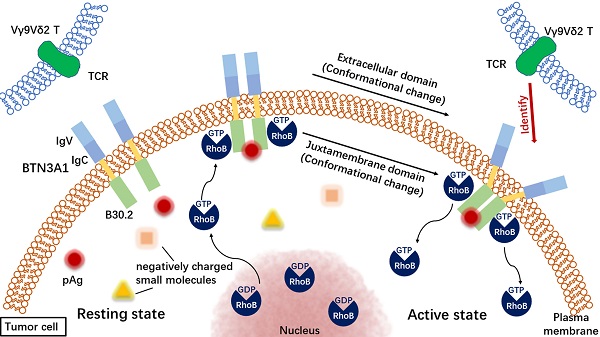
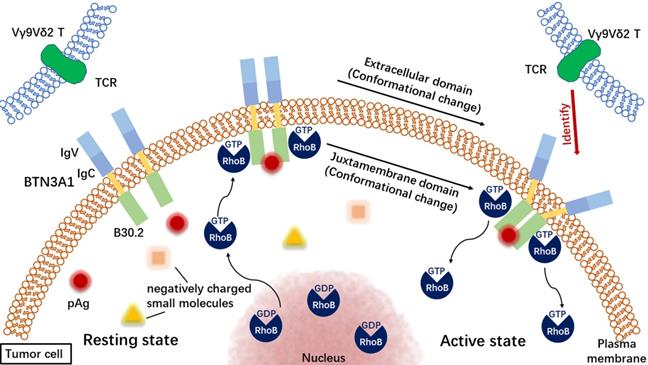
In general, pAg binding to BTN3A1 and activating Vγ9Vδ2 T cells is an “inside-out” signal transduction mechanism [39, 42] (Figure 2). Vγ9Vδ2 T cells activated by phosphorylated antigenicity can recognize and kill a variety of tumor cell lines, including breast cancer [43], pancreatic cancer [44], renal cell carcinoma [45-47], melanoma [48], neuroblastoma [49], oral squamous cell carcinoma [50] and multiple myeloma [51, 52]. Cancer therapy, which focuses on tumor immune response mediated by γδ T cells, is considered to be a promising treatment [22, 53-55].
The process of recognizing tumor cells by Vγ9Vδ2 T cells. TCR: T-cell receptor.
2.3. Man-made mutation of BTN3A1
At present, no spontaneous mutation of BTN3A family members has been identified or characterized. However, in order to determine the binding region between pAg and BTN3A1 and the importance of some residues, some researchers have constructed mutants of BTN3A1. Salim M et al. constructed the mutant of BTN3A1 H381A, H351R, Y382A, H381R and Y382F, and found that compared with wild type (WT) BTN3A1, BTN3A1 H381A, H351R and Y382A reduced the activation of Vδ2+ T cells by more than 50% [33]. Besides, BTN3A1 H381R mutant reduced the activation of Vδ2+ T cells by more than 90%, while the BTN3A1 Y382F could support more than 60% of the activation observed in WT BTN3A1 [33]. Sandstrom A et al. carried out charge-reversal mutations on five basic residues in the positive charge pocket of BTN3A1 B30.2: H378D, K393D, R412E, R418E and R469E [21]. It was found that the mutant could not bind to pAg and could not support the stimulation of Vγ9Vδ2 T cells. Gu S et al. constructed the BTN3A1 Y352A mutant. Through isothermal titration calorimetric test, it was found that compared with BTN3A1 B30.2, the mutant Y352A had obvious defects in binding to 1-hydroxy-2-methylpent-2-enyl-pyrophosphonate (cHDMAPP) and IPP, resulting in a decrease in T cell activation [19]. Nguyen K et al. found that the BTN3A1 ST296AA and T304A mutants they constructed had a reduced ability to respond to intracellular ligand treatment to trigger the activation of Vγ9Vδ2 T cells, thus proving the importance of Ser/Thr296/297 and Thr304 in juxtamembrane region [40]. Through the construction of BTN3A1 H351R and BTN3A3 R351H mutants, Sandstrom A et al. found that the BTN3A1 H351R mutants could not bind to pAg, but the binding of BTN3A3 R351H and pAg was enhanced, so as to determine the importance of 351 position [21]. Gu S et al. found that locking BTN3A1 to the V-type dimer interface (D124 / S207C) could significantly prevent pAg-induced and 20.1-induced T cell activation by mutating D124 and S207 residues in the V-type dimer interface to cysteine. This shows that the V-conformation is the resting state of BTN3A1, which is not enough to activate T cells [19].
3. BTN3A2 and cancer
The research on BTN3A2 in cancer is mainly focused on the prognosis of cancer, and there is no report on the treatment of cancer. The expression level of BTN3A2 is higher in gastric cancer [56, 57], pancreatic cancer [58] and ovarian cancer [59], but lower in breast cancer [60]. The expression of BTN3A2 is related to the adverse progression of many tumors, and its overexpression is related to the increased proliferation and invasion of gastric cancer cell lines, as well as poorly differentiated tumors in patients with pancreatic ductal adenocarcinoma who show short-term survival [58]. However, the expression of BTN3A2 is also a marker of good prognosis in many cancer patients. In breast cancer, BTN3A2 is an independent prognostic marker for triple negative breast cancer patients [60]. BTN3A2 is positively correlated with immune cell infiltration of CD8+ T cells, T cell (general), DCs, Th1 cells, and T-cell exhaustion [60]. BTN3A2 can mediate immune infiltration by regulating T cell receptor interaction and NF-κB signal pathway [60]. The higher expression level of BTN3A2 is related to the good prognosis of HR negative, HER2 positive (HR-/HER2+) breast cancer patients, and also to the distant metastasis-free survival (DMFS) rate of breast cancer patients [61]. Additionally, the expression of BTN3A2 is related to the prognosis of patients with ovarian cancer [15, 42], overall survival and disease-free progression, and its high expression is closely related to the increased overall survival rate of patients [59].
4. BTN3A3 and cancer
4.1. BTN3A3 and prognosis of cancer
The single nucleotide polymorphism (SNP) in BTN3A3 was negatively correlated with the risk of ovarian cancer [62], and BTN3A3 was associated with a lower risk of ovarian cancer recurrence [15]. Changes in the expression of specific genomes show the response of women with advanced epithelial ovarian cancer to chemotherapy. The expression of BTN3A3 gene can predict early recurrence of ovarian cancer after platinum-paclitaxel chemotherapy (21 months), with an accuracy of 86% [62]. However, in breast cancer, some high-grade malignant human breast cancer cell lines (such as MDA-MB-231) express high levels of BTN3A3, while low-grade malignant cell lines (such as MCF-7) express lower levels of BTN3A3 [63]. BTN3A3 can also predict the sensitivity of patients with gastric cancer to fluorouracil chemotherapy [64]. It is reported that BTN3A3 may increase the sensitivity to chemotherapeutic drugs by inhibiting the biological process of EMT [64]. BTN3A3 is also associated with colon cancer and can be used as a potential cancer biomarker [65]. Compared with healthy controls, the expression of BTN3A3 in patients with ulcerative colitis is significantly up-regulated [66]. If ulcerative colitis is not treated as soon as possible, it is likely to develop into colon cancer. In patients with cervical cancer, compared with normal tissues, cervical cancer tissues showed a low level of BTN3A3 methylation, indicating that the immune system of women without cervical cancer may be activated by DNA demethylation [67]. According to research, BTN3A3 is associated with intestinal inflammation and colon cancer. There is a negative correlation between the expression of BTN3A3 and IFN-γ in colonic tissue of patients with ulcerative colitis [68].
4.2. BTN3A3 and its potential therapeutic strategy for breast cancer
LSECtin is a transmembrane protein highly expressed in tumor-associated macrophages (TAMs), while BTN3A3 is the receptor of LSECtin on breast cancer cells. It has been confirmed that the interaction between LSECtin and BTN3A3 can promote the stem cell characteristics of breast cancer. The extracellular IgC and IgV domains of BTN3A3 are necessary for its interaction with LSECtin, while the intracellular domains of BTN3A3 are necessary for promoting stem cell activity. In mice carrying human tumor xenografts, macrophage-specific LSECtin interference or BTN3A3 interference in breast cancer cells can slow tumor growth. Anti-BTN3A3 mAb destruction of LSECtin-BTN3A3 axis has therapeutic effect on breast cancer. Compared with monotherapy, the combination of anti-BTN3A3 (5E08) mAb and paclitaxel significantly reduced tumor growth. Thus, it can be seen that the LSECtin-BTN3A3 axis may be a unique target in the treatment of breast cancer, which has important clinical significance in the treatment of breast cancer in the future [63].
5. Conclusion and prospect
The BTN3A family, also called CD277, consists of three members: BTN3A1, BTN3A2 and BTN3A3. At present, the research on the BTN3A family is mainly focused on the activation of Vγ9Vδ2 T cells. The latest research found that the overexpression of NLR family CARD domain containing 5 (NLRC5) not only increased the levels of BTN3A1, BTN3A2 and BTN3A3, but also promoted the activation and killing of Vγ9Vδ2 T cells functionally. In contrast, CRISPR/Cas9-mediated knockout of BTN3A molecules almost eliminated the enhanced lethality induced by NLRC5 overexpressing cells. This new discovery about the relationship between BTN3A family and NLRC5 further discusses the mechanism of BTN3A family in the activation of Vγ9Vδ2 T cells, and brings more implications for the study of immunotherapy [69].
As the first member of the BTN3A family, the essential role of BTN3A1 in the activation of Vγ9Vδ2 T cells has been verified many times. The latest research shows that CD277 specific antibodies can transform BTN3A1 from immunosuppressive molecules to immunostimulatory molecules, thus dynamically stimulating anti-tumor immunity driven by αβ and γδ T cells to block the progression of established ovarian tumors [53]. Therefore, targeting BTN3A1 can exert the synergistic killing effect of αβ and γδ T cells on tumors, and may provide a therapeutic strategy for resisting the existing immunotherapy. Although there are few studies on BTN3A2 and BTN3A3 in tumors, more and more evidence shows that BTN3A2 and BTN3A3 also play an important role in tumors. For BTN3A2, it can mediate immune infiltration by regulating T cell receptor interaction and NF-κB signal pathway [60]. For BTN3A3, it can promote the stemness of breast cancer through its interaction with LSECtin [63].
Current studies have shown that members of the BTN3A family are expressed in a variety of tumors, and the expression levels are quite different in different types of tumors. In addition, the primary phenotypic mechanisms of BTN3A family in different tumors are also different (Table 1). This suggests that the BTN3A family has a high complexity in tumorigenesis and development. In a word, BTN3A family is closely related to tumorigenesis and development. With the deepening of its research, BTN3A family is expected to become a new tumor prediction and treatment target, and will have a broad application prospect in the field of tumor.
BTN3A family and their primary mechanism of phenotype in different tumor types.
| Name | Cancer type | In vitro/in vivo | Primary mechanism of the phenotype | Reference |
|---|---|---|---|---|
| BTN3A1 | Ovarian cancer | In vitro | BTN3A1 inhibits the TCR-mediated proliferation of activated human T cells. | [2] |
| Colorectal cancer | In vitro | Zoledronate activates colorectal cancer cells in the presence of BTN3A1 to trigger Vδ2 T cells. | [22] | |
| Renal cancer | In vivo | BTN3A1 can predict response to nivolumab in metastatic clear cell renal carcinoma. | [29] | |
| BTN3A2 | Gastric cancer | In vitro | BTN3A2 promotes gastric cancer cell proliferation and invasion. | [56] |
| Pancreatic cancer | In vitro | BTN3A2 participates in Vγ9Vδ2 T cells anti-tumor functions towards pancreatic ductal adenocarcinoma. | [58] | |
| Ovarian cancer | In vitro, in vivo | Epithelial expression of BTN3A2 may modulate the infiltration of immune cells with the tumor. | [59] | |
| BTN3A3 | Breast cancer | In vitro | BTN3A3 enhances the stemness of breast cancer by interacting with LSECtin. | [63] |
| Colon cancer | In vivo | The increase of BTN3A3 expression is related to the decrease of IFN-γ level. | [68] |
Abbreviations
BTN: Butyrophilin; BTN3A: Butyrophilin 3A; BTN3A1: Butyrophilin Subfamily 3 Member A1; BTN3A2: Butyrophilin Subfamily 3 Member A2; BTN3A3: Butyrophilin Subfamily 3 Member A3; Ig: immunoglobulin; NK: natural killer; His: histidine; Arg: arginine; Lys: lysine; Tyr: tyrosine; TM: transmembrane domain; pAg: phosphate antigen; IPP: isopentenyl pyrophosphate; HMBPP: (E)-4-hydroxy-3-methyl-but-2-enyl pyrophosphate; BrHPP: phosphorylated bromohydrin; NMR: nuclear magnetic resonance; CSP: chemical shift perturbation; RhoB: Ras Homolog Family Member B; TCR: T-cell receptor; cHDMAPP: 1-hydroxy-2-methylpent-2-enyl-pyrophosphonate; WT: wild type; HR-: hormone receptor negative; HER2+: human epidermal growth factor receptor 2 positive; DMFS: distant metastasis-free survival; SNP: single nucleotide polymorphism; TAMs: tumor-associated macrophages; NLRC5: NLR family CARD domain containing 5.
Acknowledgements
This project was supported by the Xuzhou Municipal Science and Technology Project (KC20106); “Six-one” Project for High-level Health Talents in Jiangsu Province (LGY2019057); 2020 “333 Project” Award of Jiangsu Province (BRA2020252), Postgraduate Research & Practice Innovation Program of Jiangsu Province (KYCX20_2301 and KYCX20_2295).
Competing Interests
The authors have declared that no competing interest exists.
References
1. Cavaletto M, Giuffrida MG, Fortunato D, Gardano L, Dellavalle G, Napolitano L. et al. A proteomic approach to evaluate the butyrophilin gene family expression in human milk fat globule membrane. Proteomics. 2002;2:850-856
2. Cubillos-Ruiz JR, Martinez D, Scarlett UK, Rutkowski MR, Nesbeth YC, Camposeco-Jacobs AL. et al. CD277 is a negative co-stimulatory molecule universally expressed by ovarian cancer microenvironmental cells. Oncotarget. 2010;1:329-338
3. Afrache H, Gouret P, Ainouche S, Pontarotti P, Olive D. The butyrophilin (BTN) gene family: from milk fat to the regulation of the immune response. Immunogenetics. 2012;64:781-794
4. Arnett HA, Viney JL. Immune modulation by butyrophilins. Nat Rev Immunol. 2014;14:559-569
5. Franke WW, Heid HW, Grund C, Winter S, Freudenstein C, Schmid E. et al. Antibodies to the major insoluble milk fat globule membrane-associated protein: specific location in apical regions of lactating epithelial cells. J Cell Biol. 1981;89:485-494
6. Messal N, Mamessier E, Sylvain A, Celis-Gutierrez J, Thibult ML, Chetaille B. et al. Differential role for CD277 as a co-regulator of the immune signal in T and NK cells. Eur J Immunol. 2011;41:3443-3454
7. Henry J, Ribouchon M, Depetris D, Matteï M, Offer C, Tazi-Ahnini R. et al. Cloning, structural analysis, and mapping of the B30 and B7 multigenic families to the major histocompatibility complex (MHC) and other chromosomal regions. Immunogenetics. 1997;46:383-395
8. Afrache H, Pontarotti P, Abi-Rached L, Olive D. Evolutionary and polymorphism analyses reveal the central role of BTN3A2 in the concerted evolution of the BTN3 gene family. Immunogenetics. 2017;69:379-390
9. Harly C, Peigné CM, Scotet E. Molecules and mechanisms implicated in the peculiar antigenic activation process of human Vγ9Vδ2 T cells. Front Immunol. 2015;5:657
10. Yamashiro H, Yoshizaki S, Tadaki T, Egawa K, Seo N. Stimulation of human butyrophilin 3 molecules results in negative regulation of cellular immunity. J Leukocyte Biol. 2010;88:757-767
11. Compte E, Pontarotti P, Collette Y, Lopez M, Olive D. Frontline: Characterization of BT3 molecules belonging to the B7 family expressed on immune cells. Eur J Immunol. 2004;34:2089-2099
12. Rhodes DA, Chen HC, Price AJ, Keeble AH, Davey MS, James LC. et al. Activation of human γδ T cells by cytosolic interactions of BTN3A1 with soluble phosphoantigens and the cytoskeletal adaptor periplakin. J Immunol. 2015;194:2390-2398
13. Palakodeti A, Sandstrom A, Sundaresan L, Harly C, Nedellec S, Olive D. et al. The molecular basis for modulation of human Vγ9Vδ2 T cell responses by CD277/Butyrophilin-3 (BTN3A)-specific antibodies. J Biol Chem. 2012;287:32780-32790
14. Starick L, Riano F, Karunakaran MM, Kunzmann V, Li J, Kreiss M. et al. Butyrophilin 3A (BTN3A, CD277)-specific antibody 20.1 differentially activates Vγ9Vδ2 TCR clonotypes and interferes with phosphoantigen activation. Eur J Immunol. 2017;47:982-992
15. Blazquez JL, Benyamine A, Pasero C, Olive D. New insights into the regulation of γδ T cells by BTN3A and other BTN/BTNL in tumor immunity. Front Immunol. 2018;9:1601
16. Rhodes DA, Stammers M, Malcherek G, Beck S, Trowsdale J. The cluster of BTN genes in the extended major histocompatibility complex. Genomics. 2001;71:351-362
17. Adams EJ, Gu S, Luoma AM. Human gamma delta T cells: evolution and ligand recognition. Cell Immunol. 2015;296:31-40
18. Vavassori S, Kumar A, Wan GS, Ramanjaneyulu GS, Cavallari M, El Daker S. et al. Butyrophilin 3A1 binds phosphorylated antigens and stimulates human γδ T cells. Nat Immunol. 2013;14:908-916
19. Gu S, Sachleben JR, Boughter CT, Nawrocka WI, Borowska MT, Tarrasch JT. et al. Phosphoantigen-induced conformational change of butyrophilin 3A1 (BTN3A1) and its implication on Vγ9Vδ2 T cell activation. Proc Natl Acad Sci U S A. 2017;114:E7311-E7320
20. Nerdal PT, Peters C, Oberg HH, Zlatev H, Lettau M, Quabius ES. et al. Butyrophilin 3A/CD277-dependent activation of human γδ T cells: accessory cell capacity of distinct leukocyte populations. J Immunol. 2016;197:3059-3068
21. Sandstrom A, Peigné CM, Léger A, Crooks JE, Konczak F, Gesnel MC. et al. The intracellular B30.2 domain of butyrophilin 3A1 binds phosphoantigens to mediate activation of human Vγ9Vδ2 T cells. Immunity. 2014;40:490-500
22. Zocchi MR, Costa D, Venè R, Tosetti F, Ferrari N, Minghelli S. et al. Zoledronate can induce colorectal cancer microenvironment expressing BTN3A1 to stimulate effector γδ T cells with anti-tumor activity. Oncoimmunology. 2017;6:e1278099
23. Poggi A, Zocchi MR. γδ T lymphocytes as a first line of immune defense: old and new ways of antigen recognition and implications for cancer immunotherapy. Front Immunol. 2014;5:575
24. Benyamine A, Le Roy A, Mamessier E, Gertner-Dardenne J, Castanier C, Orlanducci F. et al. BTN3A molecules considerably improve Vγ9Vδ2T cells-based immunotherapy in acute myeloid leukemia. Oncoimmunology. 2016;5:e1146843
25. Tazi-Ahnini R, Henry J, Offer C, Bouissou-Bouchouata C, Mather IH, Pontarotti P. Cloning, localization, and structure of new members of the butyrophilin gene family in the juxta-telomeric region of the major histocompatibility complex. Immunogenetics. 1997;47:55-63
26. Fang J, Chen F, Liu D, Gu F, Chen Z, Wang Y. Prognostic value of immune checkpoint molecules in breast cancer. Bioscience Rep. 2020;40:BSR20201054
27. Dobosz P, Stempor PA, Roszik J, Herman A, Layani A, Berger R. et al. Checkpoint genes at the cancer side of the immunological synapse in bladder cancer. Transl Oncol. 2020;13:193-200
28. Bian B, Fanale D, Dusetti N, Roque J, Pastor S, Chretien AS. et al. Prognostic significance of circulating PD-1, PD-L1, pan-BTN3As, BTN3A1 and BTLA in patients with pancreatic adenocarcinoma. Oncoimmunology. 2019;8:e1561120
29. Incorvaia L, Fanale D, Badalamenti G, Porta C, Olive D, Luca ID. et al. Baseline plasma levels of soluble PD-1, PD-L1, and BTN3A1 predict response to nivolumab treatment in patients with metastatic renal cell carcinoma: a step toward a biomarker for therapeutic decisions. Oncoimmunology. 2020;9:e1832348
30. Gu S, Borowska MT, Boughter CT, Adams EJ. Butyrophilin3A proteins and Vγ9Vδ2 T cell activation. Semin Cell Dev Biol. 2018;84:65-74
31. Sebestyen Z, Scheper W, Vyborova A, Gu S, Rychnavska Z, Schiffler M. et al. RhoB mediates phosphoantigen recognition by Vγ9Vδ2 T cell receptor. Cell Rep. 2016;15:1973-1985
32. Moulin M, Alguacil J, Gu S, Mehtougui A, Adams EJ, Peyrottes S. et al. Vγ9Vδ2 T cell activation by strongly agonistic nucleotidic phosphoantigens. Cell Mol Life Sci. 2017;74:4353-4367
33. Salim M, Knowles TJ, Baker AT, Davey MS, Jeeves M, Sridhar P. et al. BTN3A1 discriminates γδ T cell phosphoantigens from non-antigenic small molecules via a conformational sensor in its B30.2 domain. ACS Chem Biol. 2017;12:2631-2643
34. Wang H, Henry O, Distefano MD, Wang YC, Räikkönen J, Mönkkönen J. et al. Butyrophilin 3A1 plays an essential role in prenyl pyrophosphate stimulation of human Vγ2Vδ2 T cells. J Immunol. 2013;191:1029-1042
35. Wang H, Morita CT. Sensor function for butyrophilin 3A1 in prenyl pyrophosphate stimulation of human Vγ2Vδ2 T cells. J Immunol. 2015;195:4583-4594
36. Harly C, Guillaume Y, Nedellec S, Peigné CM, Mönkkönen H, Mönkkönen J. et al. Key implication of CD277/butyrophilin-3 (BTN3A) in cellular stress sensing by a major human γδ T-cell subset. Blood. 2012;120:2269-2279
37. Hsiao CHC, Lin X, Barney RJ, Shippy RR, Li J, Vinogradova O. et al. Synthesis of a phosphoantigen prodrug that potently activates Vγ9Vδ2 T-lymphocytes. Chem Biol. 2014;21:945-954
38. Wang H, Nada MH, Tanaka Y, Sakuraba S, Morita CT. Critical roles for coiled-coil dimers of butyrophilin 3A1 in the sensing of prenyl pyrophosphates by human Vγ2Vδ2 T cells. J Immunol. 2019;203:607-626
39. Peigné CM, Léger A, Gesnel MC, Konczak F, Olive D, Bonneville M. et al. The juxtamembrane domain of butyrophilin BTN3A1 controls phosphoantigen-mediated activation of human Vγ9Vδ2 T cells. J Immunol. 2017;198:4228-4234
40. Nguyen K, Li J, Puthenveetil R, Lin X, Poe MM, Hsiao CHC. et al. The butyrophilin 3A1 intracellular domain undergoes a conformational change involving the juxtamembrane region. FASEB J. 2017;31:4697-4706
41. Vantourout P, Laing A, Woodward MJ, Zlatareva I, Apolonia L, Jones AW. et al. Heteromeric interactions regulate butyrophilin (BTN) and BTN-like molecules governing γδ T cell biology. Proc Natl Acad Sci U S A. 2018;115:1039-1044
42. Gu S, Nawrocka W, Adams EJ. Sensing of pyrophosphate metabolites by Vγ9Vδ2 T cells. Front Immunol. 2015;5:688
43. Guo BL, Liu Z, Aldrich WA, Lopez RD. Innate anti-breast cancer immunity of apoptosis-resistant human γδ-T cells. Breast Cancer Res Treat. 2005;93:169-175
44. Kitayama J, Atomi Y, Nagawa H, Kuroda A, Mutoh T, Minami M. et al. Functional analysis of TCR gamma delta+ T cells in tumour-infiltrating lymphocytes (TIL) of human pancreatic cancer. Clin Exp Immunol. 1993;93:442-447
45. Viey E, Fromont G, Escudier B, Morel Y, Rocha SD, Chouaib S. et al. Phosphostim-activated γδ T cells kill autologous metastatic renal cell carcinoma. J Immunol. 2005;174:1338-1347
46. Viey E, Laplace C, Escudier B. Peripheral γδ T-lymphocytes as an innovative tool in immunotherapy for metastatic renal cell carcinoma. Expert Rev Anticancer Ther. 2005;5:973-986
47. Kobayashi H, Tanaka Y, Yagi J, Osaka Y, Nakazawa H, Uchiyama T. et al. Safety profile and anti-tumor effects of adoptive immunotherapy using gamma-delta T cells against advanced renal cell carcinoma: a pilot study. Cancer Immunol Immunother. 2007;56:469-476
48. Kabelitz D, Wesch D, Pitters E, Zöller M. Characterization of tumor reactivity of human Vγ9Vδ2 γδ T cells in vitro and in SCID mice in vivo. J Immunol. 2004;173:6767-6776
49. Otto M, Barfield RC, Martin WJ, Iyengar R, Leung W, Leimig T. et al. Combination immunotherapy with clinical-scale enriched human γδ T cells, hu14.18 antibody, and the immunocytokine Fc-IL7 in disseminated neuroblastoma. Clin Cancer Res. 2005;11:8486-8491
50. Domae E, Hirai Y, Ikeo T, Goda S, Tsuji K. Human Vγ9Vδ2 T cells show potent antitumor activity against zoledronate-sensitized OSCC cell lines. J BUON. 2018;23:132-138
51. Kunzmann V, Bauer E, Feurle J, Weissinger F, Tony HP, Wilhelm M. Stimulation of γδ T cells by aminobisphosphonates and induction of antiplasma cell activity in multiple myeloma. Blood. 2000;96:384-392
52. Mariani S, Muraro M, Pantaleoni F, Fiore F, Nuschak B, Peola S. et al. Effector γδ T cells and tumor cells as immune targets of zoledronic acid in multiple myeloma. Leukemia. 2005;19:664-670
53. Payne KK, Mine JA, Biswas S, Chaurio RA, Perales-Puchalt A, Anadon CM. et al. BTN3A1 governs antitumor responses by coordinating αβ and γδ T cells. Science. 2020;369:942-949
54. Lu H, Dai W, Guo J, Wang D, Wen S, Yang L. et al. High abundance of intratumoral γδ T cells favors a better prognosis in head and neck squamous cell carcinoma: a bioinformatic analysis. Front Immunol. 2020;11:573920
55. Hoeres T, Smetak M, Pretscher D, Wilhelm M. Improving the efficiency of Vγ9Vδ2 T-cell immunotherapy in cancer. Front Immunol. 2018;9:800
56. Zhu M, Yan C, Ren C, Huang X, Zhu X, Gu H. et al. Exome array analysis identifies variants in SPOCD1 and BTN3A2 that affect risk for gastric cancer. Gastroenterology. 2017;152:2011-2021
57. Ni J, Deng B, Zhu M, Wang Y, Yan C, Wang T. et al. Integration of GWAS and eQTL analysis to identify risk loci and susceptibility genes for gastric cancer. Front Genet. 2020;11:679
58. Benyamine A, Loncle C, Foucher E, Blazquez JL, Castanier C, Chrétien AS. et al. BTN3A is a prognosis marker and a promising target for Vγ9Vδ2 T cells based-immunotherapy in pancreatic ductal adenocarcinoma (PDAC). Oncoimmunology. 2018;7:e1372080
59. Page CL, Marineau A, Bonza PK, Rahimi K, Cyr L, Labouba I. et al. BTN3A2 expression in epithelial ovarian cancer is associated with higher tumor infiltrating T cells and a better prognosis. PloS One. 2012;7:e38541
60. Cai P, Lu Z, Wu J, Qin X, Wang Z, Zhang Z. et al. BTN3A2 serves as a prognostic marker and favors immune infiltration in triple-negative breast cancer. J Cell Biochem. 2020;121:2643-2654
61. Han J, Choi YL, Kim H, Choi JY, Lee SK, Lee JE. et al. MMP11 and CD2 as novel prognostic factors in hormone receptor-negative, HER2-positive breast cancer. Breast Cancer Res Treat. 2017;164:41-56
62. Peedicayil A, Vierkant RA, Hartmann LC, Fridley BL, Fredericksen ZS, White KL. et al. Risk of ovarian cancer and inherited variants in relapse-associated genes. PloS One. 2010;5:e8884
63. Liu D, Lu Q, Wang X, Wang J, Lu N, Jiang Z. et al. LSECtin on tumor-associated macrophages enhances breast cancer stemness via interaction with its receptor BTN3A3. Cell Res. 2019;29:365-378
64. Pan J, Dai Q, Xiang Z, Liu B, Li C. Three biomarkers predict gastric cancer patients' susceptibility to fluorouracil-based chemotherapy. J Cancer. 2019;10:2953-2960
65. Kiyamova R, Garifulin O, Gryshkova V, Kostianets O, Shyian M, Gout I. et al. Preliminary study of thyroid and colon cancers-associated antigens and their cognate autoantibodies as potential cancer biomarkers. Biomarkers. 2012;17:362-371
66. Lebrero-Fernández C, Wenzel UA, Akeus P, Wang Y, Strid H, Simrén M. et al. Altered expression of Butyrophilin (BTN) and BTN-like (BTNL) genes in intestinal inflammation and colon cancer. Immun Inflamm Dis. 2016;4:191-200
67. Gašperov NM, Farkas SA, Nilsson TK, Grce M. Epigenetic activation of immune genes in cervical cancer. Immunol Lett. 2014;162:256-257
68. Lebrero-Fernández C, Wenzel UA, Akeus P, Wang Y, Strid H, Simrén M. et al. Altered expression of Butyrophilin (BTN) and BTN-like (BTNL) genes in intestinal inflammation and colon cancer. Immun Inflamm Dis. 2016;4(2):191-200
69. Dang AT, Strietz J, Zenobi A, Khameneh HJ, Brandl SM, Lozza L. et al. NLRC5 promotes transcription of BTN3A1-3 genes and Vγ9Vδ2 T cell-mediated killing. iScience. 2021;24(1):101900
Author contact
Corresponding authors: Shaohua Fan, School of Life Science, Jiangsu Normal University, Xuzhou, Jiangsu, 221116, China. Tel: +86 516 83403170, E-mail: fshflyedu.cn; Yanyan Wang, Department of Ultrasonic Medicine, Xuzhou First People's Hospital, Xuzhou, Jiangsu, 221000, China. Tel: +86 516 68383289, E-mail: lulizhan335com

 Global reach, higher impact
Global reach, higher impact