Impact Factor
ISSN: 1837-9664
J Cancer 2021; 12(19):5807-5816. doi:10.7150/jca.47557 This issue Cite
Research Paper
Identification of a prognostic 4-mRNA signature in laryngeal squamous cell carcinoma
1. Department of Otorhinolaryngology-head and neck surgery, Shanghai General Hospital, Shanghai Jiao Tong University School of Medicine, Shanghai, China.
2. Department of Otorhinolaryngology-head and neck surgery, Shanghai General Hospital, Shanghai, China.
Abstract
Background: Laryngeal squamous cell carcinoma (LSCC) is one of the most common malignancy in the respiratory tract and could reduce the quality of life seriously like dyspnea, dysphonia and dysphagia. Moreover, 5-year survival rate has decreased over the past 40 years. This study was designed to identify mRNAs that related to prognosis in LSCC to enable early detection and outcome improvement.
Methods: Gene expression profiles from Gene Expression Omnibus (GEO) (GSE59102, GSE84957) and The Cancer Genome Atlas (TCGA) were analyzed to identify differentially expressed genes (DEGs) with the help of bioinformatics tools. Functional enrichment analyses including Gene Ontology (GO) and pathway analysis were carried out to investigate the role of those genes and underlying molecular mechanisms in LSCC. Cox's regression analyses (univariate, LASSO and multivariate in order) were utilized to identify DEGs related with patients' overall survival and a 4-mRNA-based prognostic risk score model was established. Univariate and multivariate Cox's regression analyses were then performed on LSCC data (90 patients left) to identify independent predictors of OS, including the signature and clinicopathologic variables. The prognostic value of the gene signature was further validated and the genes were analyzed by GEPIA to get pan-cancer expression profiles.
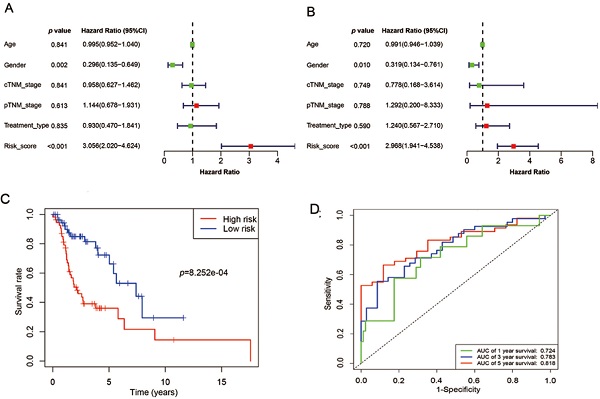
Results: 444 differentially expressed mRNAs (250 up-regulated, 194 down-regulated) were identified based on the threshold of fold change > 2 and adjusted p value < 0.05. Univariate Cox's regression analysis showed that high risk score (HR: 3.056, 95% confidence interval [CI]: 0.135-0.649, p<0.001) and female (HR: 0.296, 95% CI: 2.020-4.624, p=0.002) were associated with relatively poor prognosis. Further multivariate Cox's regression analysis indicated that risk score and gender were independent prognostic factors (p<0.05). The risk score model could stratify patients into high- and low‑risk groups, which presents significantly differential overall survival (p= 8.252e-04). The AUCs of 1-, 3- and 5-year OS were 0.724, 0.783 and 0.818, respectively.
Conclusions: Our study provides evidence that the four-mRNA signature could serve as a biomarker to predict prognosis in LSCC, especially in long-term.
Keywords: Laryngeal squamous cell carcinoma, prognosis, risk model, GEO, TCGA

 Global reach, higher impact
Global reach, higher impact