Impact Factor ISSN: 1837-9664
J Cancer 2022; 13(1):21-33. doi:10.7150/jca.64817 This issue Cite
Research Paper
Identification and Validation of a Novel 2-LncRNAs Signature Associated with m6A Regulation in Colorectal Cancer
1. Department of Organ Transplantation and Hepatobiliary, The First Affiliated Hospital of China Medical University, No.155 Nanjing North Street, Heping District, Shenyang 110001, Liaoning Province, China.
2. Department of Ultrasound, Xiang'an Hospital of Xiamen University, No.2000 Xiang'an East Road, Xiang'an District, Xiamen 361101, Fujian Province, China.
3. Department of Colorectal Surgery, Cancer Hospital of China Medical University, Liaoning Cancer Hospital and Institute. No.44 Xiaoheyan Road, Dadong District, Shenyang 110042, Liaoning Province, China.
# These authors contributed equally to this work.
Abstract
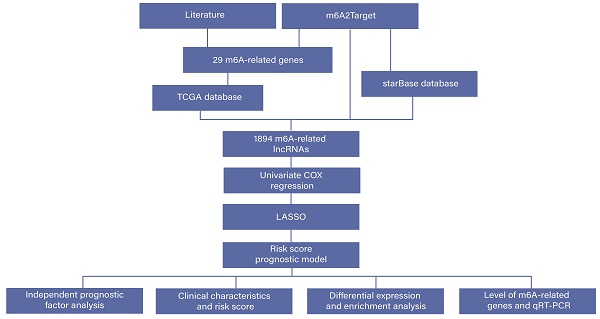
Colorectal cancer (CRC) is one of the most common tumors in the digestive system, and it is urgent to identify a new biomarker for the diagnosis and treatment of CRC. N6-methyladenosine (m6A) is an abundant mRNA modification and is almost involved in every aspect of physiological processes. In this study, we constructed a novel m6A-related 2-lncRNAs signature that can predict the prognosis of CRC. We obtained m6A-related lncRNAs and identified prognostic lncRNAs through univariate Cox regression analysis and least absolute shrinkage and selection operator (LASSO) analysis, then constructed a prognostic model based on the risk score, and we also verified the stability of the model. In addition, differential expression analysis between the high- and low-risk subgroups was performed. A total of 1,894 m6A-related lncRNAs were screened from various sources. Using univariate Cox regression analysis and survival analysis, two lncRNAs (AL135999.1 and AL049840.4) were identified (P < 0.05), and the coefficients of lncRNAs were calculated by LASSO. The high-risk group had worse clinical outcomes and overall survival (OS) than the low-risk group, and the risk score can serve as an independent prognostic factor in CRC. In addition, different stages of CRC also showed a different level of risk score. Finally, we found that two lncRNAs were differentially expressed (P < 0.01) in CRC patients, and AL135999.1 may be relevant to m6A modification mediated by methyltransferase-like 3 (METTL3) in CRC. In summary, we constructed a reliable 2-lncRNAs signature based on the risk score, and we identified two m6A-related prognostic lncRNAs, AL135999.1 and AL049840.4. The novel 2-lncRNAs signature plays an essential role in predicting the prognosis of CRC.
Keywords: colorectal cancer, N6-methyladenosine, biomarker, lncRNA signature, prognostic model.

 Global reach, higher impact
Global reach, higher impact