3.2
Impact Factor
ISSN: 1837-9664
J Cancer 2022; 13(1):184-201. doi:10.7150/jca.65225 This issue Cite
Research Paper
The diagnostic and prognostic significance of small nuclear ribonucleoprotein Sm D1 aberrantly high expression in hepatocellular carcinoma
1. The Fuzong Clinical Medical College of Fujian Medical University, Fuzhou, Fujian 350025, PR China.
2. Department of Hepatobiliary Surgery, 900 Hospital of the Joint Logistic Team, Fuzhou, Fujian 350025, PR China.
* Contributed equally.
Abstract
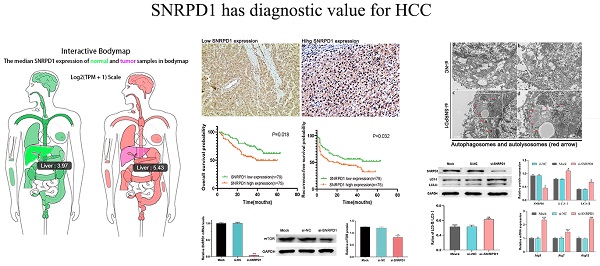
Small nuclear ribonucleoprotein Sm D1 (SNRPD1), one of the crucial genes encoding core spliceosome components, was abnormally highly expressed in multiple types of tumors. In this study, we investigated the diagnostic and prognostic significance of SNRPD1 in hepatocellular carcinoma (HCC). The investigation of datasets from GEO and TCGA databases revealed that SNRPD1 expression in HCC was significantly higher than adjacent normal liver tissues, which was validated by immunohistochemistry (IHC). Both GO, KEGG analysis showed that the SNRPD1 co-expressed genes mainly enriched in Cell division, Nuclear import, mRNA splicing via spliceosome, Ribosome, Cell cycle, etc. Survival analysis from the GSE14520 dataset and 154 HCC cohorts exhibited a significant association of high SNRPD1 expression with poor overall survival and recurrence-free survival. ROC analysis showed that the abnormally high SNRPD1 mRNA expression has diagnostic significance in distinguishing between HCC and normal liver tissue (AUC = 0.819). Gene set enrichment analysis (GSEA) demonstrated that the high expression of SNRPD1 might regulate HCC tumorigenesis and progression by affecting the cell cycle, mismatch repair, DNA replication, and RNA degradation, etc. The luciferase report assay revealed that SNRPD1 was the direct target gene of miR-100 manifested by decreased SNRPD1 expression and luciferase activity in the HCC cells upon miR-100 overexpression. Finally, SNRPD1 may as an oncogene affecting the progression of HCC through regulates the mTOR pathway and autophagy.
Keywords: SNRPD1, miR-100, Hepatocellular carcinoma, prognosis, mechanism.

 Global reach, higher impact
Global reach, higher impact