Impact Factor
ISSN: 1837-9664
J Cancer 2022; 13(5):1540-1552. doi:10.7150/jca.60124 This issue Cite
Research Paper
circ_0000567/miR-421/TMEM100 Axis Promotes the Migration and Invasion of Lung Adenocarcinoma and Is Associated with Prognosis
Department of Thoracic Surgery II, Key Laboratory of Carcinogenesis and Translational Research (Ministry of Education), Peking University Cancer Hospital and Institute, Beijing, People's Republic of China
Abstract
Purpose: Due to the high metastatic ability and poor prognosis of lung adenocarcinoma (LUAD), we identified novel non-coding RNAs, which constitute approximately 60% of human transcripts, as prognostic biomarkers and potential therapeutic targets for LUAD.
Methods: In this study, we downloaded and analyzed microRNA (miRNA) datasets from The Cancer Genome Atlas (TCGA) to identify dysregulated miRNAs correlating with the overall survival (OS) of LUAD patients. miR-421, circ_0000567, and TMEM100 expression levels were examined by quantitative real-time polymerase chain reaction (qRT-PCR) in NSCLC tissues from 73 patients and adjacent normal tissues. Cell migration and invasion were assayed using wound healing and transwell assays. miR-421 target predictions were conducted using starBase, CircInteractome, circBank, TargetScan, miRanda, MirDB, miRpath, and Gene Expression Omnibus (GEO) databases. The circular structure and stability of circ_0000567 were verified by RNase R digestion and qRT-PCR using oligo(dT) and random primers. A luciferase reporter assay was used to evaluate the relationship between miR-421, circ_0000567, and TMEM100.
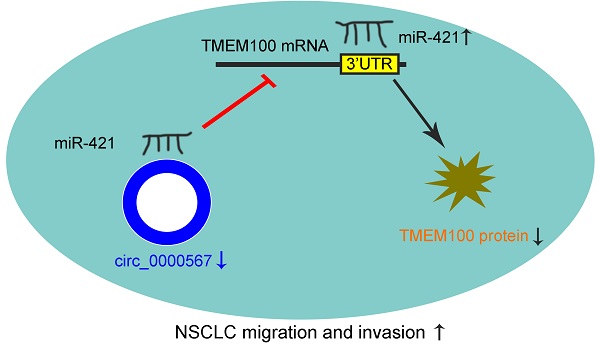
Results: The miRNA panel associated with OS in patients with LUAD was screened according to the hazard ratio (HR) of miRNAs from high to low. Based on the correlation between these miRNAs and OS, as well as miRNA expression levels, miR-421 was selected for further outcome analysis. High miR-421 expression was an independent risk factor for shorter OS in 73 patients collected from our department. Bioinformatic analyses, luciferase reporter assays, and functional assays showed that circ_0000567 could act as a sponge for miR-421 and prevent it from directly targeting the 3'-untranslated region of TMEM100 mRNA and further degrading it in LUAD. miR-421 expression was significantly upregulated, while circ_0000567 and TMEM100 were downregulated in tumor tissues of LUAD, compared to their counterparts in normal tissues. Gain- and loss-of-function assays showed that miR-421 promoted LUAD cell migration and invasion. Overexpression of circ_0000567 inhibited migration and invasion, whereas co-transfection of circ_0000567 and miR-421 mimics partly counteracted this effect. TMEM100 was upregulated by enhanced circ_0000567 in LUAD cells, and the expression of TMEM100 was inversely proportional to miR-421, whereas it was directly proportional to circ_0000567 in 73 LUAD specimens, which confirmed the competitive endogenous RNA (ceRNA) network.
Conclusion: Our findings suggest that miR-421 promotes the migration and invasion of lung adenocarcinoma via circ_0000567/miR-421/TMEM100 signaling and could be a prognostic biomarker for LUAD.
Keywords: LUAD, Hsa-miR-421, Hsa_circ_0000567, TMEM100, prognosis

 Global reach, higher impact
Global reach, higher impact