Impact Factor
ISSN: 1837-9664
J Cancer 2022; 13(5):1588-1602. doi:10.7150/jca.66949 This issue Cite
Research Paper
Prognostic Values and Underlying Regulatory Network of Cohesin Subunits in Esophageal Carcinoma
State Key Laboratory of Bioactive Substances and Function of Natural Medicine, Institute of Materia Medica, Chinese Academy of Medical Sciences and Peking Union Medical College, Beijing 100050, China.
Abstract
Background: Cohesin is a highly conserved and ubiquitously expressed protein complex. While increasing evidence suggests that cohesin dysregulation is vital in the carcinogenesis of numerous malignancies, little is known about the prognostic values and potential mechanisms of cohesin subunits and direct regulators in esophageal carcinoma (ESCA).
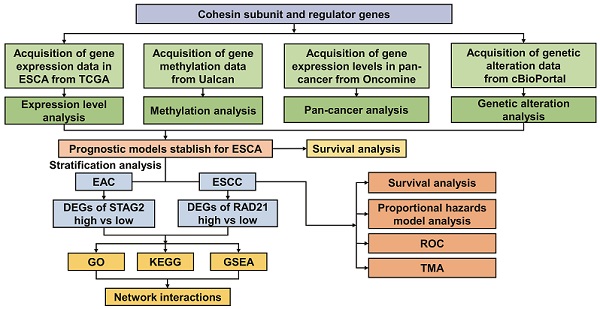
Methods: RNA-sequencing data from The Cancer Genome Atlas (TCGA) and Genome Tissue Expression (GTEx) were used. The subunits and regulators of cohesin affecting the prognosis of ESCA were screened by Kaplan-Meier survival analysis; univariate and multivariate Cox regression analyses were performed; and the receiver-operating characteristic (ROC) curve was determined. The ESCA hazard model and nomogram map were constructed by integrating the clinical data. We used functional analysis and protein-protein interaction (PPI) networks to explore underlying pathways. Finally, immunohistochemistry was performed to examine the expression levels of cohesin subunits in tissue microarray (TMA).
Results: Transcriptome data from multiple ESCA patient datasets showed cohesin subunits SMC1A, SMC1B, SMC3, STAG1, STAG2, RAD21, and cohesin regulators including ESCO2, NIPBL, MAU2, WAPL, PDS5A and PDS5B were all upregulated in ESCA tissues compared to normal tissues. Survival analysis demonstrated that high STAG2 expression was significantly associated with poorer overall survival (OS) and progression-free survival (PFS) in esophageal adenocarcinoma (EAC). In contrast, high RAD21 expression was significantly correlated with better OS in esophageal squamous cell carcinoma (ESCC). Moreover, STAG2 and RAD21 were identified as independent prognostic factors and predictive biomarkers in EAC and ESCC, respectively. Functional enrichment analysis further revealed that STAG2 and RAD21 were mainly involved in the mitotic nuclear division, DNA repair, angiogenesis, epithelial-mesenchymal transition (EMT), and oncogenic signaling pathways. PPI analysis illustrated that STAG2 and RAD21 could cross-talk through cancer-associated modules and performed the core roles of the whole PPI network. Using TMA, STAG2 protein expression positively correlated with lymph node metastasis and advanced clinical stage of EAC patients, whereas there was a negative correlation between RAD21 protein expression and the malignant clinicopathological parameters in ESCC.
Conclusion: These findings suggest that STAG2 and RAD21 can be used as predictive biomarkers for risk assessment and prognostic stratification in ESCA, which provide potential novel insights into molecular targets of ESCA.
Keywords: STAG2, RAD21, esophageal carcinoma (ESCA), biomarker, prognosis

 Global reach, higher impact
Global reach, higher impact