Impact Factor ISSN: 1837-9664
J Cancer 2022; 13(7):2086-2104. doi:10.7150/jca.68673 This issue Cite
Research Paper
Comprehensive analysis of expression signature and immune microenvironment signature of biomarker Endothelin Receptor Type A in stomach adenocarcinoma
1. Department of Gastroenterology, The First Affiliated Hospital of China Medical University, Shenyang, Liaoning, China.
2. Department of Orthopedics, The First Affiliated Hospital of China Medical University, Shenyang, Liaoning, China.
3. Department of Organ transplantation and Hepatobiliary, The First Affiliated Hospital of China Medical University, Shenyang, Liaoning, China.
4. Department of Paediatrics, Wuhan Children's Hospital (Wuhan Maternal and Child Healthcare Hospital), Tongji Medical College, Huazhong University of Science and Technology, Wuhan, China.
Abstract
Background: EDNRA (Endothelin Receptor Type A) is closely associated with tumor progression in many tumor types. However, the functional mechanism of EDNRA in stomach adenocarcinoma (STAD) remains to be elucidated.
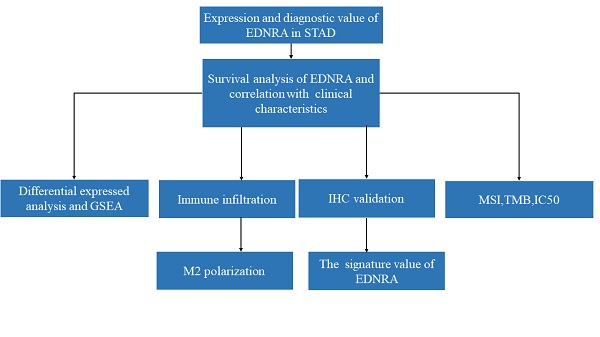
Methods: ENDRA expression levels in STAD were assessed. A Receiver Operating Characteristic (ROC) curve was constructed to measure the diagnostic value of EDNRA. The correlation between ENDRA expression levels and patient clinical-pathological characteristics was analyzed. The survival and prognostic significance were validated using Kaplan-Meier and Cox regression and confirmed by the immunohistochemistry cohorts. Differentially expressed genes of EDNRA in STAD were determined, and EDNRA related functional enrichment and biological pathways involved in STAD were obtained by Gene-Set Enrichment Analysis (GSEA). The correlation between EDNRA expression in STAD and immune cell infiltration was assessed using the CIBERSORT and Spearman correlation analysis, and the correlation between EDNRA and TMB, MSI, IC50, and immune checkpoints was examined.
Results: EDNRA expression was significantly higher in STAD than in normal tissues (P < 0.001) and associated with worse overall survival (OS). EDNRA expression was significantly associated with T stage, histological type, histologic grade, and TP53 status. Cox regression analysis revealed that primary therapy outcome, age, tumor status, and EDNRA were independent prognostic factors for OS. Multivariate analysis revealed that EDNRA expression, tumor status, age, and primary therapy outcome influenced patient prognosis. GSEA was significantly enriched in several pathways and biological processes, which include Immunoregulatory, Hedgehog, WNT, PI3K-AKT.NK cells, Tem, macrophages, and mast cells were substantially positively correlated with EDNRA expression in the STAD microenvironment. Notably, high EDNRA expression may promote M2 macrophages to block PD-1-mediated immunotherapy and induce immunosuppression. In addition, patients with high expression of EDNRA might be resistant to the treatment of several anti-tumor drugs.
Conclusion: Our results suggest that EDNRA was closely related to clinicopathologic characteristics, poor prognosis, and promoted macrophage differentiation and synergistic role in immunosuppression.
Keywords: EDNRA, Tumor biomarker, Immune checkpoint, M2 macrophage, Immunoregulatory

 Global reach, higher impact
Global reach, higher impact