Impact Factor
ISSN: 1837-9664
J Cancer 2023; 14(10):1707-1724. doi:10.7150/jca.84215 This issue Cite
Research Paper
High Expression of Microtubule-associated Protein TBCB Predicts Adverse Outcome and Immunosuppression in Acute Myeloid Leukemia
1. State Key Laboratory of Experimental Hematology, National Clinical Research Center for Blood Diseases, Haihe Laboratory of Cell Ecosystem, Institute of Hematology & Blood Diseases Hospital, Chinese Academy of Medical Sciences & Peking Union Medical College, Tianjin, 300020, China.
2. Tianjin Institutes of Health Science, Tianjin 301600, China.
3. Regenerative Medicine Clinic, Institute of Hematology & Blood Diseases Hospital, Chinese Academy of Medical Sciences & Peking Union Medical College, Tianjin, 300020, China.
Abstract
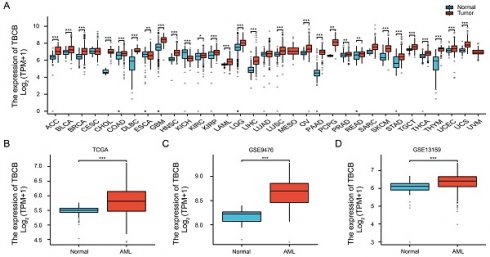
Acute myeloid leukemia (AML) is a devastating blood cancer with high heterogeneity and ill-fated outcome. Despite numerous advances in AML treatment, the prognosis remains poor for a significant proportion of patients. Consequently, it is necessary to accurately and comprehensively identify biomarkers as soon as possible to enhance the efficacy of diagnosis, prognosis and treatment of AML. In this study, we aimed to identify prognostic markers of AML by analyzing the cohorts from TCGA-LAML database and GEO microarray datasets. Interestingly, the transcriptional level of microtubule-associated protein TBCB in AML patients was noticeably increased when compared with normal individuals, and this was verified in two independent cohorts (GSE9476 and GSE13159) and with our AML patients. Furthermore, univariate and multivariate regression analysis revealed that high TBCB expression was an independent poor prognostic factor for AML. GO and GSEA enrichment analysis hinted that immune-related signaling pathways were enriched in up-regulated DEGs between two populations separated by the median expression level of TBCB. By constructing a protein-protein interaction network, we obtained six hub genes, all of which are immune-related molecules, and their expression levels were positively linked to that of TBCB. In addition, the high expression of three hub genes was significantly associated with a poor prognosis in AML. Moreover, we found that the tumor microenvironment in AML with high TBCB expression tended to be infiltrated by NK cells, especially CD56bright NK cells. The transcriptional levels of NK cell inhibitory receptors and their ligands were positively related to that of TBCB, and their high expression levels also predicted poor prognosis in AML. Notably, we found that the down-regulation of TBCB suppressed cell proliferation in AML cell lines by enhancing the apoptosis and cell cycle arrest. Finally, drug sensitivity prediction illustrated that cells with high TBCB expression were more responsive to ATRA and midostaurin but resistant to cytarabine, dasatinib, and imatinib. In conclusion, our findings shed light on the feasibility of TBCB as a potential predictor of poor outcome and to be an alternative target of treatment in AML.
Keywords: TBCB, prognosis, acute myeloid leukemia, bioinformatics, immune evasion, drug sensitivity

 Global reach, higher impact
Global reach, higher impact