Impact Factor ISSN: 1837-9664
J Cancer 2015; 6(2):160-168. doi:10.7150/jca.10873 This issue Cite
Research Paper
The Analysis of the Relationship between Multiple Myeloma Cells and Their Microenvironment
1. Department of Hematology, Jagiellonian University Medical College, Krakow, Poland
2. Department of Transplantation, Jagiellonian University Medical College, Krakow, Poland
3. Department of Infectious Diseases, Jagiellonian University Medical College, Krakow, Poland
4. Department of Radioligands, Faculty of Pharmacy, Jagiellonian University Medical College, Krakow, Poland
5. Department of Medicine, John Dempsey Hospital, University of Connecticut, Farmington, CT, USA
Received 2014-10-21; Accepted 2014-12-13; Published 2015-1-5
Abstract
The bone marrow microenvironment plays a key role in the stimulation of growth and survival of multiple myeloma (MM) cells. We investigated whether membrane microfragments (MFBs) exert a stimulatory effect on mesenchymal stem cell (MSC) gene expression or differentiation. MSCs from patients with multiple myeloma (MMBM-MSCs) proliferated at a slower rate than MSCs from healthy volunteers (BM-MSCs), and fewer MMBM-MSCs adhered to the substrate as compared to BM-MSCs. Phenotypic analysis revealed that MMBM-MSCs and BM-MSCs differed significantly in terms of their CD166 and CXCR4 expressions. In conclusion, our comparative analysis of mesenchymal cells from MM patients and healthy volunteers revealed differences in the genetic and phenotypic profiles of these two populations, their potential for osteodifferentiation, and expression of surface antigens. Moreover, we showed that membrane MFBs may alter the genetic profile of MSCs, leading to disorders of their osteodifferentiation, and interact with the WNT pathway via presentation of the DKK-1 protein.
Keywords: bone marrow, genotype, membrane microfragments, microenvironment, multiple myeloma, osteodifferentiation
Introduction
Multiple myeloma (MM) is an incurable hematologic malignancy of plasma cells that accumulate and proliferate in the bone marrow. Most MMs arise from bone marrow, which suggests that the bone marrow microenvironment plays a key role in the stimulation of growth and survival of MM cells [1-2]. However, the microenvironmental signals and their relevant intracellular messengers, which may serve as a potential target for therapeutic intervention, have not been fully investigated thus far. Bone marrow constitutes a source of mesenchymal stem cells (MSCs). MSCs, representing approximately 0.001% of all nucleated cells in bone marrow, are a heterogeneous population of adult stem cells with a high proliferative potential and the ability to differentiate. Friedenstein et al. were the first to isolate and characterize mesenchymal stem cells. They showed that MSCs have the potential to differentiate into such mesenchymal cell lines as osteocytes, adipocytes and chondrocytes [3]. Myeloma cells interact with bone marrow microenvironment consisting of endothelial cells, stromal cells, mesenchymal cells, osteoclasts, osteoblasts, immune cells, adipocytes and extracellular matrix. These interactions are responsible for homing of specific cells in the bone marrow niche, their proliferation, survival, and, in the case of MM cells, resistance to chemotherapy [4]. Direct contact of myeloma cells with the stroma via integrin receptors may be an important factor influencing their proliferation and survival [2]. The growth of myeloma cells is also modulated by cytokines or stromal bone growth factors, such as interleukin 6 (IL-6), hepatocyte growth factor (HGF), vascular endothelial growth factor (VEGF), insulin-like growth factor 1 (IGF-1), transforming growth factor-beta (TGF-β) and other factors that interfere with bone structure. Uncontrolled proliferation of myeloma cells, synthesizing excessive amounts of monoclonal immunoglobulins, as well as the interaction of these cells with the microenvironment, cause impairment of the immune system and lead to abnormal haematopoiesis, anaemia, renal dysfunction, and development of osteolytic bone lesions [5-7].
Both normal and neoplastic cells exfoliate their cellular membranes and secrete circular membrane fragments, which are referred to as microfragments (MFBs) [8]. MFBs extracted from intact cells are usually smaller than those from apoptotic cells. The amount of secreted MFBs increases during activation of cells, hypoxia and radiation damage. The damage results from the generation of reactive oxygen species and exposure to proteins derived from the activated complement cascade [9]. MFBs play an array of functions: they may stimulate other cells by expressing “complex signalling” ligands on their surface, deliver these ligands to target cells, deliver proteins, mRNA and bioactive lipids into target cells, thus transmitting infectious particles (e.g. HIV, prions), or perhaps even move entire organelles (e.g. mitochondria) to other cells [10]. Flow cytometric analysis of the normal and neoplastic cell lines showed the presence of small structures, which were considered to be fragments of cells. The amount of these fragments turned out to be specific for a given cell line [11]. Studies using electron microscopy demonstrated that these small fragments contain many membrane MFBs. Similar processes occur both in vitro and in vivo; MFBs released by tumour cells are present both in the microenvironment of the tumour and in the peripheral blood [12-16]. MFBs were demonstrated to play a significant role in lung cancer, particularly by facilitating its growth and metastatic potential [17].
The aim of this study was to evaluate the interaction between the bone marrow microenvironment and myeloma cells, and analyse the role of MFBs in the pathogenesis of MM.
Materials and Methods
Ethics
The study was conducted in accordance with the Declaration of Helsinki and approved by the local ethics committee.
Participants
The study included 20 multiple myeloma patients (12 men and 8 women) treated at the Haematology Department, University Hospital in Krakow. The patients were aged between 45 and 78 years (median age 64.1 years) and showed bone marrow plasmacytosis (15-65%, mean 35.6%). The diagnosis of multiple myeloma was based on histological evidence of increased fraction of plasma cells in bone marrow biopsy specimens ( > 10% plasma cells), and the presence of monoclonal protein in blood or urine and presence of CRAB criteria [18]. Control group was comprised of 10 volunteers (5 men and 5 women) aged between 48 and 68 years (median age 64.3 years). Bone marrow aspirates from the posterior iliac crest were collected to sterile anticoagulant-coated (EDTA or heparin) Vacutainer tubes (Becton Dickinson, New Jersey, USA).
Isolation and culture of normal cells
We isolated bone marrow MSCs from healthy donors (BM-MSCs) and from multiple myeloma patients (MMBM-MSCs). MSCs were isolated according to the protocol described by Friedenstein et al. [3] Bone marrow mononuclear cells were isolated by centrifugation of the total bone marrow on ficoll gradient (1000 x g, 30 min) and subsequent selection via adhesion to a plastic surface. 4 x 105/cm2 mononuclear cells were seeded into plastic bottles (Sarstedt AG & Co., Numbrecht, Germany). Cultures of BM-MSCs and MMBM-MSCs were grown on Dulbecco's Modified Eagle's Medium (DMEM, Sigma-Aldrich, Missouri, USA), supplemented with 10% fetal bovine serum (FBS) for incubation of MSCs (Stem Cell Technologies, Vancouver, Canada). The mixture was supplemented with antibiotics (PAA Laboratories GmbH, Pasching, Austria) and for MMBM-MSCs with IL-6 (2 ng/ml). The cultures were incubated at 37°C in an atmosphere of 5% CO2 with 95% humidity. After reaching 90% density, the cells were passaged using 0.05% trypsin (trypsin-EDTA, PAA Laboratories GmbH) and seeded into new culture flasks at 1 x 104/cm2.
Cell viability during culturing was assessed using the Trypan Blue Exclusion Test. The MTS test (Cell Titer 96 Aqueous One Solution Cell Proliferation Assay) was performed to assess cells proliferation as per manufacturer instructions (Promega, Promega, Madison, WI, USA). Absorbance was measured using the EL800 plate reader (BIO-TEK Instruments, USA) at 490nm.
Induction and evaluation of MSC differentiation into osteoblasts
The role of the bone marrow microenvironment was studied on mesenchymal cell populations that had the potential to differentiate into osteogenic cells capable of forming the normal structure of bone tissue. BM-MSCs differentiated into osteoblasts after the third passage. In preparation for osteoinduction, 3-5 x 104 cells/well were seeded on 6-well plates and cultured in differentiation medium (DMEM, 10% FBS, 0.1 mM dexamethasone, 50 g/ml ascorbic acid, and 10 M glycerophosphate) for 21 days. The medium was changed twice a week. To visualize calcium phosphate deposited by osteoblasts, cultures were fixed in 4% paraformaldehyde (2 min), stained with alizarin Red S (Sigma, Missouri, USA) for 2 min, and washed with 70% ethanol. Stained calcium phosphate deposits were visualized using an Olympus microscope and photographed using an Olympus camera. Analogous experiments were carried out in the presence of MM membrane microfragments (MM-MFBs, 20ng/ml) in a differentiating medium.
Flow cytometric analysis of MSC phenotype
After the third passage, BM-MSCs and MMBM-MSCs were subjected to cytofluorometric analysis for the presence of surface antigens common to MSCs (CD73, CD166 and CD271) and surface antigens common to hematopoietic lines (CD45, CD34 and CXCR4). 1 x 105 cells were suspended in 100 ml PBS with 2% FBS and stained with the appropriate monoclonal antibody (Becton Dickinson) for 30 min at 4°C in the dark. Subsequently, the cells were harvested using a FACS Canto flow cytometer (Becton Dickinson) and analysed using FACS Diva software (Becton Dickinson).
Evaluation of gene expression using real-time PCR
After the third passage, MSCs were subjected to genotypic analysis with real-time PCR. We determined expressions of genes for Runx2, PPAR-γ, osteocalcin, collagen-α, matrix metalloproteinase 2 (MMP-2), matrix metalloproteinase 9 (MMP-9), interleukin 1 (IL-1), interleukin 8 (IL-8), interleukin 11 (IL-11), TGF-β, HGF, VEGF and β fibroblast growth factor (β-FGF). The results were expressed relative to the endogenous GADPH control and illustrated in arbitrary units. We compared the genotypes of MSCs from healthy donors and multiple myeloma patients, and analysed expressions of genes in cells exposed to membrane MFBs. RNA was isolated using a RNeasy Mini Kit (Qiagen, Valencia, CA, USA). The concentration and purity of the obtained RNA was evaluated by measuring absorbance at a 260 nm and 280 nm wavelength with a DU 640B spectrophotometer (Beckman Coulter, Fullerton, CA, USA). RNA was transcribed into cDNA using MMLV reverse transcriptase (Promega, Fitchburg, USA) and random primers (Promega). The levels of gene expressions were determined by quantitative real-time PCR (qRT-PCR) based on the specific TaqMan probe (Applied Biosystems, Foster City, CA, USA) using a ABI PRISM 7300 Sequence Detection System (Applied Biosystems). The components of the reaction mixture used in the qRT-PCR reaction were: TaqMan PCR Master Mix 25 μl, cDNA 100 ng, 20x probe 2.5 μl, water added until final volume was 50 μl. Probes used for qRT PCR were manufactured by Applied Biosystems accordingly: TaqMan RUNX2 (Hs00231692_ml), PPARγ (Hs01115513_ml), HGF (Hs00300159_m1), αCOL1 (Hs01076780_ml), OSTEOCALC (Hs01587814_ml), PTHR (Hs00896835_ml), MMP-2 (Hs00234579_m1), MMP-9 (Hs00234579_m1), TGFβ (Hs00998133_m1), IL-8 (Hs00174103_m1), IL-11 (Hs00899845_ml), IL-1 (Hs01055413_ml), VEGF (Hs00173626_m1), FGF2/β-FGF (Hs00266645_m1).
We analysed the relative expression of genes expressed as ΔΔCt, being the difference between the empiric value and the Ct of the reference gene.
Isolation of membrane MFBs
For MFBs isolation, human plasmacytoma lymphocyte B cells (U266B1, ATCC TIB-196) were cultured in RPMI Medium 1640 (Gibco, BRL, Grand Island NY, USA) with 10% FBS, 2 ml L-gluthamine and the following antibiotics: penicillin (100 IU/ml) and streptomycin (100 µg/ml). Cells were maintained at 37oC in a humidified atmosphere which contained 5% CO2 and were seeded into a 6-well plate (Sarsted, Germany) at a density of 5x105 cells/well in 2mL of medium.
MFBs released from cellular membranes from U266 cells were isolated according to method proposed by Baj-Krzyworzeka et al. 2006 [19]. Supernatants from MMBM-MSC cells were collected after centrifugation at 2.000g for 20 min. To remove cell debris samples were centrifuged again at 50.000g (RC28S, Sorvall, Newton, CT, USA) for 1 h at 4oC. Pellets were washed several times to remove FBS and finally resuspended in serum-free medium. Isolated microfragments were thus prepared for further analysis.
The effect of stimulation with membrane MFBs on MSC gene expression
To test the effect of MM-MFBs on mesenchymal cells, three independent experiments were performed. Mesenchymal cells from the third passage were stimulated with MM-MFBs at a concentration of 20 µg/ml, and genotypic analysis with real-time PCR was conducted 8, 24 and 48 h after the stimulation.
Differentiation of MSCs in the presence or absence of membrane MFBs
To compare the potential of MSCs to differentiate into osteoblasts, the cells from the third passage were subjected to osteoinduction. 3-5 x 104 cells/well were plated on 6-well plates and cultured for 7, 14 and 21 days in a differentiation medium containing dexamethasone, β-glycerophosphate and ascorbic acid. The medium was changed twice a week. After completing the culture, calcium phosphate deposited by the cells was stained with Alizarin Red S. The same procedures were carried out in the presence of MM-MFBs as well.
Statistical analysis
Statistical analysis was conducted with Microsoft Excel and GraphPad Prism 4.02 software. The experiments were carried out in at least three iterations. Statistical characteristics of quantitative variables are presented as arithmetic means with standard deviations or standard errors expressed as error bars on the graphs. The information on the number of experiments overall and the number of tests conducted during each specific experiment is given below each chart. Statistical significance of intergroup difference was verified with either the Student-t-test or one-way analysis of variance. The threshold of statistical significance was set at p<0.05.
Results
Characteristics of MSCs
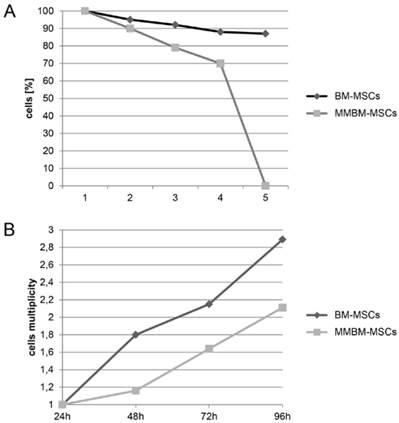
We observed that the number of MMBM-MSCs that adhered to the substrate during culture was lower than the respective number of BM-MSCs. Moreover, MMBM-MSCs grew at a slower rate than the control BM-MSCs (Figure 1 A and B).
A. The rate of proliferation of BM-MSCs and MMBM-MSCs. 1-4 denotes number of cell passages. B. The growth rate of BM-MSCs and MMBM-MSCs after 24, 48, 72 and 96 h of cells culturing.
Phenotypic analysis of MSCs
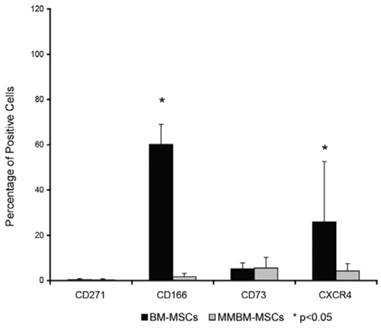
Both types of MSCs showed similar phenotypes, expressing surface antigens CD271 and CD73. However, MMBM-MSCs and BM-MSCs differed significantly in terms of their CD166 and CXCR4 expressions (Figure 2).
Comparison of surface antigens expressed by MMBM-MSCs and BM-MSCs. Cells were stained using the appropriate antigens and analysed using the FACS method. Statistically significant differences were documented for CD166 and CXCR4 antigens (n=5, p<0.05).
Genotypic analysis
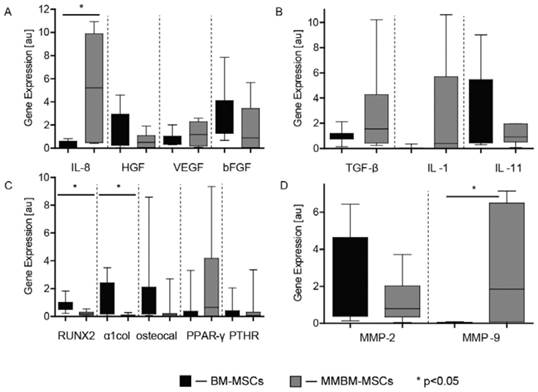
MMBM-MSCs showed significantly stronger expression of IL-8 and MMP-9 than BM-MSCs. The differences in the expressions of TGF-β, IL-1, VEGF and PPAR-γ turned out to be insignificant. Additionally, a decrease in the MMBM-MSC expression of RUNX2 and α1col was significantly when compared with control MSCs (Figure 3).
Genotyping of MMBM-MSCs and BM-MSCs from the third passage. Results were normalized, relative to the GAPDH control. The graphs show the level of mRNA for A) IL8, HGF, VEGF and β-FGF, B) TGF-β, IL-1 and IL-11, C) RUNX2, α1col, osteocalcin, PPAR-γ and PTHR, and D) MMP-2 and MMP-9 genes.
Characteristics of membrane MFBs
The expression of Dikkoph-1 (DKK-1), a protein involved in the WNT signalling pathway and bone osteolysis, was detected in the protein lysate. Also we observed expression of IL-8, VEGF and HGF in the MMBM-MSC membrane MFBs (data not shown).
Effect of stimulation with MFBs on the expression of MSC genes
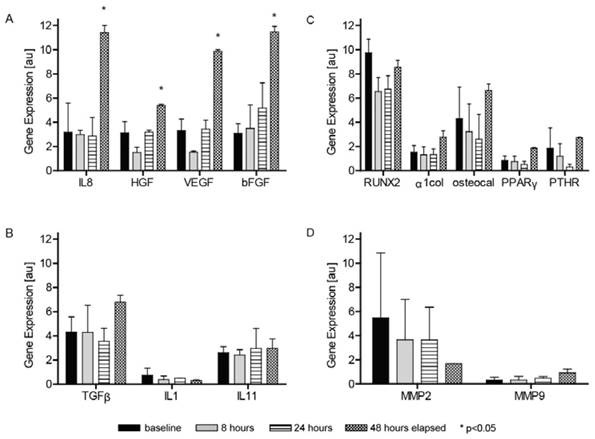
The levels of BM-MSC gene expressions baseline and after 8, 24 and 48 h of stimulation with MM-MFBs are presented on Figure 4. The increase in the gene expression turned out to be most pronounced after 48 h of stimulation when statistically significant changes in the expressions of genes encoding IL-8, HGF, VEGF and β-FGF were documented.
Expression of genes in MMBM-MSCs and BM-MSCs after 8-, 24- and 48-h stimulation with MM-MFBs. The cells were grown on 6-well plates in DMEM with 10% FBS for MSCs, 2 mM L-glutamine and antibiotics. Results were normalized, relative to the GAPDH control. The graphs show the level of mRNA for A) IL8, HGF, VEGF and β-FGF, B) TGF-β, IL-1 and IL-11, C) RUNX2, α1col, osteocalcin, PPAR-γ and PTHR, and D) MMP-2 and MMP-9 genes.
Differentiation of BM-MSCs in the presence or absence of membrane MFBs
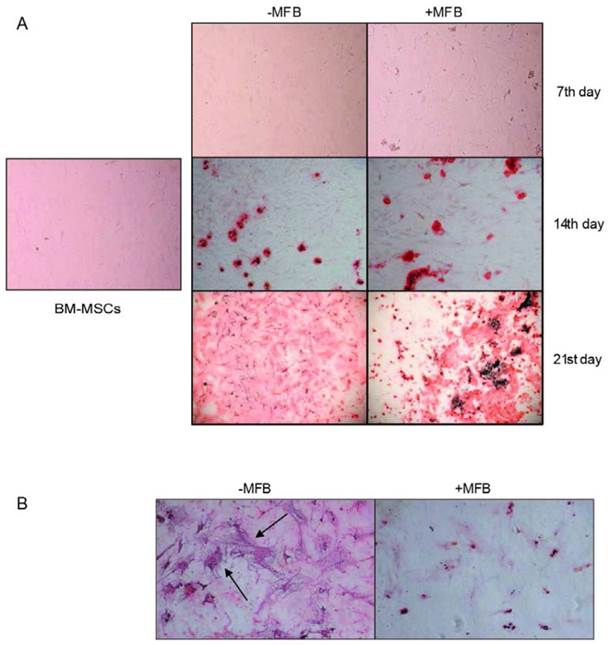
To determine their potential to differentiate, BM-MSCs from the third passage were subjected to osteoinduction. Neither presence of phosphate deposits nor differentiation was observed in the case of cells exposed to MM-MFBs, and all these cells eventually died (Figure 5).
Osteoinduction of BM-MSCs in the presence of MM-MFBs. The cells from the third passage were cultured for 7, 14 and 21 days in the differentiation medium (DMEM with 10% FBS, dexamethasone, β-glycerophosphate and ascorbic acid). Subsequently, the cells were fixed in 4% paraformaldehyde, and calcium phosphate deposits were stained with alizarin red S. Legend: + MFB - differentiation in presence of 20 µl/ ml myeloma membrane microfragments, -MFB - differentiation in absence of any additional factors, control - cells that were not subjected to the differentiation. Pictures were taken at a 100 x (A) and 200 x (B) magnification. Representative images are shown.
Discussion
In this study, we compared the MSC from healthy donors and MM patients in order to understand the mechanisms that are responsible for the changes in the bone marrow microenvironment during MM progression. We compared the rate of proliferation of MMBM-MSCs and BM-MSCs. The growth rate of BM-MSCs turned out to be significantly higher than that of MMBM-MSCs. Furthermore, MSCs from healthy donors could be maintained in culture for a longer period of time and displayed the ability to deposit calcium phosphate. Conversely, MMBM-MSCs stopped proliferating and died after only four passages.
The phenotype of MSCs isolated from MM is different than in the case of their normal counterparts. Our analysis of the antigenic profile of cells from the third passage revealed that both MMBM-MSCs and BM-MSCs show relatively weak expression of CD271 and CD73 markers; this finding is consistent with previously published data [20-23]. CD166 turned out to be significantly decreased on the surface of MMBM-MSCs. Furthermore, MSCs showed strong expression of CXCR4 receptor. Cytokines play a key role in cell homing and migration in MM, among them one of the most recognized is stromal cell-derived factor 1α (SDF-1α, also known as CXCL12) and its receptor CXCR4 [24]. SDF-1α is produced by many cells of bone marrow including stromal cells, osteblasts and osteoclasts [25]. MM cells express SDF-1α and CXCR4 [26], overexpression of CXCR4 was confirmed in a number of cancers, including MM [27]. It is not fully understood what leads to CXCR4 overexpression in tumors, however it is postulated that the pro-inflammatory cytokines TNFα [28] and VEGF [29], hypoxia [30] or genetic factors [31] may all play a role. CXCR4 expression in MM cells has been reported to be inversely correlated with disease activity [26]. Wright et al. demonstrated that SDF-1α influences osteoclast precursors thus regulating bone resorption [32]. Inhibition of the SDF-1α/CXCR4 axis inhibits osteolysis in animal models [33]. The SDF-1α/CXCR4 axis is another potential target for therapy of MM. Compounds which are specific inhibitors of CXCR4 tend to hinder the process of MM homing [26,34]. Oliveira et al. reported that thalidomide down-regulated SDF-1α and CXCR4 expression in MM patients [35]. Niesvizky et al. showed that plasma levels of SDF-1α were significantly lower in patients receiving bortezomib, versus a non-bortezomib-based therapy [36]. Pandey et al. reported suppression of CXCR4 by gambogic acid (GA) led to down-regulation of invasion induced by the ligand SDF-1α. They found that in MM cells, GA suppressed the induction of differentiation of macrophages to osteoclasts in an IL-6 dependent mechanism [27]. Bao et al. reported that CXCR4 is a good prognostic indicator of survival in multiple myeloma patients [37].
The differences between MSCs derived from the BM of newly diagnosed myeloma patients and those from healthy donors can give insight into their role in the pathophysiology of MM and its consequences. Although the potential of MSCs from both origins is similar, the functionality and genetic features of MSC from MM can differ from their healthy counterparts [38]. Garcia-Gomez et al. analyzed the transcriptomic profile from both healthy donors and MM patients which were co-cultured with myeloma cell line MM.1S. Their study confirmed that there was up regulation in the genetic expression of CXCL1, CXCL5, and CXCL6 in both groups, while there was up regulation of the genetic expression of Neuregulin 3 and Norrie disease protein exclusively in the MM group [39]. CXCL1, CXCL5, and CXCL6 are all important in the pathophysiology of MM; several members of the CXC family of chemokines act as promoters of angiogenesis [40]. Neuregulin 3 is implicated in normal plasma cell differentiation and also in myeloma biology [41]. Norrie disease protein is involved in canonical Wntβ/-catenin signaling pathway [42].
In this study we analysed the expression of genes involved in differentiation of osteocytes, including genes active at the early stages of the process, such as Runx2 and PPAR-γ, and the genes that are specific for the later stages, such as genes for osteocalcin, collagen-1α, MMP-9 and MMP-2, which degrade denatured collagen and cell membrane components. We also analysed the expression of gene encoding IL-11, a cytokine synthesized by osteoblasts, eosinophils and stromal cells of bone marrow. Finally, we studied the expressions of genes for IL-1, pro-inflammatory cytokines, factors associated with angiogenesis (IL-8, HGF, VEGF, β-FGF), and TGF-β which is involved in the proliferation and differentiation of osteogenic cells. We showed significant differences between the gene expression profiles of MSCs from bone marrow of MM patients and healthy volunteers. MMBM-MSCs were characterized by significantly higher transcript levels of IL-8 and MMP-9, but significantly lower levels of Runx2 and collagen-1α. These findings are consistent with the results published by Todoerti et al., who also showed that the the microenvironment of MM patients differs from that of healthy donors [43]. Metalloproteinases are involved in the degradation of extracellular matrix proteins, and MMP-2 and MMP-9 preferentially digest components of the basement membrane. Moreover, the pro-inflammatory cytokines of MSCs, such as TGFβ1, IL-1β and TNFα, were shown to play a vital role in the stimulation and expression of MMPs and TIMPs [44]. Along with other enzymes, MMP-9 plays a role in the metabolism of type IV and V collagen, elastin and denatured collagen in normal tissue. During inflammation, stimulation by pro-inflammatory cytokines can result in a massive release of MMPs, including MMP-9, from inflammatory cells. IL-8 is a molecule associated with angiogenesis. Increased plasma levels of IL-8 were demonstrated by Kurihara et al., who studied the role of various factors (among them IL-6 and VEGF) in hematologic malignancies, and their correlation with the development of microcirculation [45]. They demonstrated that the expression of genes involved in osteogenesis, including genes active at the early stages of differentiation (i.e., Runx2), was significantly weaker in mesenchymal cells originating from MM patients. Moreover, they observed decreased expression of gene encoding collagen-α, which is normally activated during the late stages of collagen differentiation. In our study, control MSCs showed stronger expressions of Runx2, osteocalcin, PPAR-γ and collagen genes, which is consistent with the results published by [46], who analysed genes involved in bone formation.
MFBs are important molecules involved in transduction of signal between cells, and thereby modulate functions of the latter. They activate intracellular signal transduction cascades via MAPK p42/44, AKT, and STAT proteins, which results in cellular proliferation and inhibition of apoptosis [10]. Furthermore, secretion of MFBs by tumour cells leads to activation of stromal factors stimulating angiogenesis and endothelial cell proliferation. Moreover, MFBs are involved in transfer of receptors between cells, e.g. in transfer of cell adhesion molecules from platelets to hematopoietic cells, which is reflected by enhanced adhesion of the latter to fibrinogen and endothelium [47,48]. The transfer of MFBs from activated platelets to lung cancer cells, results in increase adhesion of the latter to endothelium and their ability to metastasize in vivo, [17]. Tumour cells coated with MFBs that were transplanted into mice demonstrated greater metastatic capability than the non-encapsulated cells. Higher levels of MFBs in the peripheral blood of gastric cancer patients correlated with increased metastatic of the tumour and worse prognosis [14]. MFBs produced by tumour cells stimulate fibroblasts to secretion of various pro-angiogenic factors which act as chemo-attractants and enhance proliferation of endothelial cells, thus promoting angiogenesis [17].
Our analysis of membrane MFBs revealed the presence DKK-1 in microfragment protein lysates isolated from the supernatant of the U266 MM cell line. In the case of MM patients, elevated levels of DKK-1 are associated with osteolytic changes [49]. Tian et al. demonstrated involvement of DKK-1 in the inhibition of osteoblast activity in MM [50]. DKK-1 is an important component of the WNT signalling pathway and plays an important role in osteodifferentiation. DKK-1 secreted by plasma cells at a concentration above 12 ng/ml inhibited osteodifferentiation of murine mesenchymal cells in vitro. Expression of the gene transcript for DKK-1 correlated strongly with the severity of osteolytic lesions in myeloma patients [50]. However, other authors did not document the presence of DKK-1 in myeloma cells, which suggests that these cells do not universally express this protein [51,52]. We demonstrated that myeloma cells secrete MFBs containing inhibitors of the WNT pathway, the activation of which is required for proper development and modelling of skeletal tissue. Previous studies revealed that stimulation with MFBs alters the cells' potential to osteodifferentiate. We confirmed this observation, staining calcium phosphate deposits at 7th, 14th and 21st day of differentiation. In presence of MFBs, the cells did not proliferate and completely died after 3 weeks. Staining calcium phosphate deposits with von Kossa method confirmed that control mesenchymal cells are capable of osteodifferentiation. This suggests that myeloma MFBs exert negative effect on the bone, via transfer of protein inhibitors of the WNT pathway, and points to a previously unknown role of membrane microfragments.
We revealed that stimulation with MM-MFB alters expression of genes involved in angiogenesis, which points to an important role of microfragments in cell-to-cell signal transmission. Similar to previous studies, we observed a significant, time-dependent increase in the expression of IL-8, HGF, VEGF and β-FGF after 48 hours of incubation with myeloma MFBs. Although the differences in the expression of other genes turned out to be statistically insignificant, we observed a similar trend as in the comparative analysis of MMBM-MSCs and MB-MSCs. Longer exposure of MSCs to membrane MFBs would likely reveal the effect of microfragments on the expression of genes involved in osteogenesis.
In conclusion, our comparative analysis of mesenchymal cells from MM patients and healthy volunteers revealed differences in the genetic and phenotypic profiles of these two populations, their potential for osteodifferentiation, and expression of surface antigens. Moreover, we showed that membrane MFBs may alter the genetic profile of MSCs, leading to disorders of their osteodifferentiation, and interact with the WNT pathway via presentation of the DKK-1 protein.
Abbreviations
BM-MSCs: bone marrow mesenchymal stem cells; DMEM: Dulbecco's Modified Eagle's Medium; ESCs: embryonic stem cells; FBS: fetal bovine serum; HGF: hepatocyte growth factor; IGF-1: insulin-like growth factor 1; IL-1: interleukin 1; IL-6: interleukin 6; IL-8: interleukin 8; IL-11: interleukin 11; MFBs: microfragments; MM: multiple myeloma; MMBM-MSCs: bone marrow mesenchymal stem cells from multiple myeloma patients; MM-MFBs: membrane microfragments from myeloma patients; MMP-2: matrix metalloproteinase 2; MMP-9: matrix metalloproteinase 9; MSCs: mesenchymal stem cells; TGF-β: transforming growth factor β; VEGF: vascular endothelial growth factor; β-FGF: β fibroblast growth factor; SDF-1α: stromal cell-derived factor 1α.
Acknowledgements
Conception and design: A. Jurczyszyn, A. Zebzda, M. Majka; acquisition of data: A. Jurczyszyn, A. Zebzda, M. Majka; analysis and interpretation of data: A. Jurczyszyn, A. Zebzda, J. Czepiel, J. Gdula-Argasińska, W. Perucki, M. Majka; writing, review, and/or revision of the manuscript: A. Jurczyszyn, A. Zebzda, J. Czepiel, J. Gdula-Argasińska, W. Perucki, M. Majka.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Derksen PW, de Gorter DJ, Meijer HP. et al. The hepatocyte growth factor/Met pathway controls proliferation and apoptosis in multiple myeloma. Leukemia. 2003;17:764-74
2. Gahrton G. New therapeutic targets in multiple myeloma. Lancet. 2004;364:1648-9
3. Friedenstein AJ, Gorskaja JF, Kulagina NN. Fibroblast precursors in normal and irradiated mouse hematopoietic organs. Exp Hematol. 1976;4:267-74
4. De Raeve HR, Vanderkerken K. The role of the bone marrow microenvironment in multiple myeloma. Histol Histopathol. 2005;20:1227-50
5. Podar K, Tai YT, Lin BK. et al. Vascular endothelial growth factor-induced migration of multiple myeloma cells is associated with beta 1 integrin- and phosphatidylinositol 3-kinase-dependent PKC alpha activation. J Biol Chem. 2002;277:7875-81
6. Jurczyszyn A, Czepiel J, Biesiada G. et al. HGF, sIL-6R and TGF-β1 play a significant role in the progression of multiple myeloma. J Cancer. 2014;5:518-24
7. Lesko E, Majka M. The biological role of HGF-MET axis in tumor growth and development of metastasis. Front Biosci. 2008;13:1271-80
8. Fevrier B, Raposo G. Exosomes: endosomal-derived vesicles shipping extracellular messages. Curr Opin Cell Biol. 2004;16:415-21
9. Hugel B, Martinez MC, Kunzelmann C. et al. Membrane microparticles: two sides of the coin. Physiology. 2005;20:22-7
10. Ratajczak J, Wysoczynski M, Hayek F. et al. Membrane-derived microvesicles: important and underappreciated mediators of cell-to-cell communication. Leukemia. 2006;20:1487-95
11. Beaudoin AR, Grondin G. Shedding of vesicular material from the cell surface of eukaryotic cells: different cellular phenomena. Biochim Biophys Acta. 1991;1071:203-19
12. Ginestra A, Miceli D, Dolo V. et al. Membrane vesicles in ovarian cancer fluids: a new potential marker. Anticancer Res. 1999;19:3439-45
13. Kim CW, Lee HM, Lee TH. et al. Extracellular membrane vesicles from tumor cells promote angiogenesis via sphingomyelin. Cancer Res. 2002;62:6312-7
14. Kim HK, Song KS, Park YS. et al. Elevated levels of circulating platelet microparticles, VEGF, IL-6 and RANTES in patients with gastric cancer: possible role of a metastasis predictor. Eur J Cancer. 2003;39:184-91
15. McCarty OJ, Mousa SA, Bray PF. et al. Immobilized platelets support human colon carcinoma cell tethering, rolling, and firm adhesion under dynamic flow conditions. Blood. 2000;96:1789-97
16. Taucher S, Salat A, Gnant M. et al. Impact of pretreatment thrombocytosis on survival in primary breast cancer. Thromb Haemost. 2003;89:1098-106
17. Janowska-Wieczorek A, Wysoczynski M, Kijowski J. et al. Microvesicles derived from activated platelets induce metastasis and angiogenesis in lung cancer. Int J Cancer. 2005;113:752-60
18. Durie BG, Harousseau JL, Miguel JS. et al. International uniform response criteria for multiple myeloma. Leukemia. 2006;20:1467-1473
19. Baj-Krzyworzeka M, Szatanek R, Weglarczyk K. et al. Tumour-derived microvesicles carry several surface determinants and mRNA of tumour cells and transfer some of these determinants to monocytes. Cancer Immunol Immunother. 2006;55:808-18
20. Katz AJ, Tholpady A, Tholpady SS. et al. Cell surface and transcriptional characterization of human adipose-derived adherent stromal (hADAS) cells. Stem Cells. 2005;23:412-23
21. Lee RH, Kim B, Choi I. et al. Characterization and expression analysis of mesenchymal stem cells from human bone marrow and adipose tissue. Cell Physiol Biochem. 2004;14:311-24
22. Rebelatto CK, Aguiar AM, Moretao MP. et al. Dissimilar differentiation of mesenchymal stem cells from bone marrow, umbilical cord blood, and adipose tissue. Exp Biol Med. 2008;233:901-13
23. Wagner W, Wein F, Seckinger A. et al. Comparative characteristics of mesenchymal stem cells from human bone marrow, adipose tissue, and umbilical cord blood. Exp Hematol. 2005;33:1402-16
24. Raman D, Baugher PJ, Thu YM. et al. Role of chemokines in tumor growth. Cancer Lett. 2007;256:137-65
25. Le Y, Zhou Y, Iribarren P. et al. Chemokines and chemokine receptors: their manifold roles in homeostasis and disease. Cell Mol Immunol. 2004;1:95-104
26. Alsayed Y, Ngo H, Runnels J. et al. Mechanisms of regulation of CXCR4/SDF-1 (CXCL12)-dependent migration and homing in multiple myeloma. Blood. 2007;109:2708-17
27. Pandey MK, Kale VP, Song C. et al. Gambogic acid inhibits multiple myeloma mediated osteoclastogenesis through suppression of chemokine receptor CXCR4 signaling pathways. Exp Hematol. 2014;42:883-96
28. Kulbe H, Hagemann T, Szlosarek PW. et al. The inflammatory cytokine tumor necrosis factor-alpha regulates chemokine receptor expression on ovarian cancer cells. Cancer Res. 2005;65:10355-62
29. Lipton A. Bisphosphonate therapy in the oncology setting. Expert Opin Emerg Drugs. 2003;8:469-88
30. Schioppa T, Uranchimeg B, Saccani A. et al. Regulation of the chemokine receptor CXCR4 by hypoxia. J Exp Med. 2003;198:1391-402
31. Bhandari D, Trejo J, Benovic JL. et al. Arrestin-2 interacts with the ubiquitin-protein isopeptide ligase atrophin-interacting protein 4 and mediates endosomal sorting of the chemokine receptor CXCR4. J Biol Chem. 2007;282:36971-9
32. Wright LM, Maloney W, Yu X. et al. Stromal cell-derived factor-1 binding to its chemokine receptor CXCR4 on precursor cells promotes the chemotactic recruitment, development and survival of human osteoclasts. Bone. 2005;36:840-53
33. Ooi LL, Dunstan CR. CXCL12/CXCR4 axis in tissue targeting and bone destructionin cancer and multiple myeloma. J Bone Miner Res. 2009;24:1147-9
34. Azab AK, Runnels JM, Pitsillides C. et al. CXCR4 inhibitor AMD3100 disrupts the interaction of multiple myeloma cells with the bone marrow microenvironment and enhances their sensitivity to therapy. Blood. 2009;113:4341-51
35. Oliveira AM, Maria DA, Metzger M. et al. Thalidomide treatment down-regulates SDF-1alpha and CXCR4 expression in multiple myeloma patients. Leuk Res. 2009;33:970-3
36. Niesvizky R, Mark TM, Ward M. et al. Over-coming the response plateau in multiple myeloma: a novel bortezomib-basedstrategy for secondary induction and high-yield CD34+ stem cell mobilization. Clin Cancer Res. 2013;19:1534-46
37. Bao L, Lai Y, Liu Y. et al. CXCR4 is a good survival prognostic indicator in multiple myeloma patients. Leuk Res. 2013;37:1083-8
38. Garcia-Gomez A, Sanchez-Guijo F, Consuelo del Cañizo M. et al. Multiple myeloma mesenchymal stromal cells: Contribution to myeloma bone disease and therapeutics. World J Stem Cells. 2014;6:322-343
39. Garcia-Gomez A, De Las Rias J, Ocio EM. et al. Transcriptomic profile induced in bone marrow mesenchymal stromal cells after interaction with multiple myeloma cells: implications in myeloma progression and myeloma bone disease. Oncotarget. 2014;5:8284-305
40. Strieter RM, Polverini PJ, Kunkel SL. et al. The functional role of the ELR motif in CXC chemokine-mediated angiogenesis. J Biol Chem. 1995;270:27348-27357
41. Mahtouk K, Cremer FW, Reme T. et al. Heparan sulphate proteoglycans are essential for the myeloma cell growth activity of EGF-family ligands in multiple myeloma. Oncogene. 2006;25:7180-7191
42. Braunger BM, Tamm ER. The different functions of Norrin. Adv Exp Med Biol. 2012;723:679-683
43. Todoerti K, Lisignoli G, Storti P. et al. Distinct transcriptional profiles characterize bone microenvironment mesenchymal cells rather than osteoblasts in relationship with multiple myeloma bone disease. Exp Hematol. 2010;38:141-53
44. Ries C, Egea V, Karow M. et al. MMP-2, MT1-MMP, and TIMP-2 are essential for the invasive capacity of human mesenchymal stem cells: differential regulation by inflammatory cytokines. Blood. 2007;109:4055-63
45. Kurihara N, Bertolini D, Suda T. et al. IL-6 stimulates osteoclast-like multinucleated cell formation in long term human marrow cultures by inducing IL-1 release. J Immunol. 1990;144:4226-30
46. Noel D, Caton D, Roche S. et al. Cell specific differences between human adipose-derived and mesenchymal-stromal cells despite similar differentiation potentials. Exp Cell Res. 2008;314:1575-84
47. Baj-Krzyworzeka M, Majka M, Pratico D. et al. Platelet-derived microparticles stimulate proliferation, survival, adhesion, and chemotaxis of hematopoietic cells. Exp Hematol. 2002;30:450-9
48. Janowska-Wieczorek A, Majka M, Kijowski J, Baj-Krzyworzeka M, Reca R, Turner AR. et al. Platelet-derived microparticles bind to hematopoietic stem/progenitor cells and enhance their engraftment. Blood. 2001;98:3143-9
49. Semenov MV, Zhang X, He X. DKK1 antagonizes Wnt signaling without promotion of LRP6 internalization and degradation. J Biol Chem. 2008;283:21427-32
50. Tian E, Zhan F, Walker R. et al. The role of the Wnt-signaling antagonist DKK1 in the development of osteolytic lesions in multiple myeloma. N Engl J Med. 2003;349:2483-94
51. Ehrlich LA, Chung HY, Ghobrial I. et al. IL-3 is a potential inhibitor of osteoblast differentiation in multiple myeloma. Blood. 2005;106:1407-14
52. Zangari M, Esseltine D, Lee CK. et al. Response to bortezomib is associated to osteoblastic activation in patients with multiple myeloma. Br J Haematol. 2005;131:71-3
Author contact
Corresponding author: Professor Marcin Majka, Department of Transplantation, Jagiellonian University Medical College, 265 Wielicka St., 30-663 Krakow, Poland. E-mail: mmajkakrakow.pl, tel/fax 48-12-658-20-11.

 Global reach, higher impact
Global reach, higher impact