Impact Factor ISSN: 1837-9664
J Cancer 2021; 12(20):6165-6169. doi:10.7150/jca.62154 This issue Cite
Research Paper
The contribution of YTHDF2 gene rs3738067 A>G to the Wilms tumor susceptibility
1. Department of Pathology, The First Affiliated Hospital of Kunming Medical University, Kunming 650031, Yunnan, China.
2. Department of Pediatric Surgery, Guangzhou Institute of Pediatrics, Guangdong Provincial Key Laboratory of Research in Structural Birth Defect Disease, Guangzhou Women and Children's Medical Center, Guangzhou Medical University, Guangzhou 510623, Guangdong, China.
3. Department of Clinical Laboratory, Yunnan Key Laboratory of Laboratory Medicine, the First Affiliated Hospital of Kunming Medical University, Kunming 650032, Yunnan, China.
4. Kunming Key Laboratory of Children Infection and Immunity, Yunnan Key Laboratory of Children's Major Disease Research, Yunnan Institute of Pediatrics Research, Yunnan Medical Center for Pediatric Diseases, Kunming Children's Hospital, Kunming 650228, Yunnan, China.
5. Department of Pediatric Surgery, the First Affiliated Hospital of Zhengzhou University, Zhengzhou 450052, Henan, China.
6. Department of Pediatric Surgery, the Second Affiliated Hospital of Xi'an Jiaotong University, Xi'an 710004, Shaanxi, China.
7. Department of Hematology, The Second Affiliated Hospital and Yuying Children's Hospital of Wenzhou Medical University, Wenzhou 325027, Zhejiang, China.
8. Department of Pathology, Children Hospital and Women Health Center of Shanxi, Taiyuan 030013, Shannxi, China.
9. Yunnan Key Laboratory of Stem Cell and Regenerative Medicine, Biomedical Engineering Research Center, Kunming Medical University, Kunming 650500, Yunnan, China.
#These authors contributed equally to this work.
Received 2021-4-29; Accepted 2021-8-9; Published 2021-8-26
Abstract
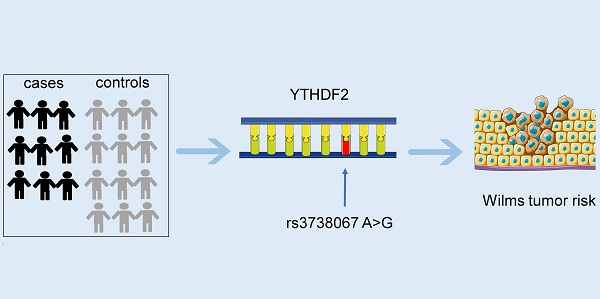
YTHDF2 is responsible for maintaining the dynamic N6-methyladenosine (m6A) modification balance and influences a variety of cancers. We tested whether YTHDF2 gene rs3738067 A>G polymorphism is related to Wilms tumor by genotyping samples of Chinese children (450 cases and 1317 controls). However, the rs3738067 A>G polymorphism showed no statistical significance with Wilms tumor susceptibility. Stratification analysis also revealed that there was no remarkable association of rs3738067 variant AG/GG genotype with Wilms tumor risk in every subgroup (age, gender, and clinical stages). In all, the results indicated YTHDF2 gene rs3738067 A>G polymorphism could not alter Wilms tumor risk significantly.
Keywords: Wilms tumor, susceptibility, m6A, YTHDF2, polymorphism
Introduction
Wilms tumor (nephroblastoma) is a typical common seen embryonal kidney cancer in childhood [1]. Its prevalence was about 1 in 10,000 children in Western populations [2], whereas 3.3 in one million children in China [3]. Wilms tumor usually represents epithelial, undifferentiated/blastemal, and stromal components in varying proportions [4, 5]. Moreover, Wilms tumor also displays heterologous elements such as cartilage, osteoid, and neural elements [6-8]. This heterogeneity suggests a complexity to the underlying causes of Wilms tumor [9].
Knowledge of the genetic underpinnings of Wilms tumor is growing. In 1990, the WT1 gene was first cloned as a Wilms tumor suppressor gene [10]. Subsequently, mutations in WTX and CTNNB1, loss of imprinting (LOI), or loss of heterozygosity (LOH) at 11p15 were found to contribute to Wilms tumor development [11-14]. Moreover, multiple genetic variants have been identified as Wilms tumor risk loci in genetic association studies [15-18]. However, all the identified gene mutations or single nucleotide polymorphisms (SNPs) could only explain a small set of the etiology of Wilms tumor [19, 20]. Thus, identification and characterization of more variants are indispensable in better unraveling the full genetic spectrum of Wilms tumor.
N6-methyladenosine (m6A) is a prevalent internal modification of mRNAs, taking up >80% of all RNA base methylation [21]. It regulates the processing, localization, translation, and eventual decay of RNA [22]. m6A modification is deposited by the methyltransferase complex (writer) composed of METTL3, METTL14, and WTAP [23]. Such modification could be reversed by m6A demethylases (erasers) including FTO and ALKBH5. Meanwhile, m6A associated RNA binding proteins (readers), including YTHDF1-3 and YTHDC1, also function in m6A modification by modulating mRNA fate [24, 25]. m6A modification is highly involved in the carcinogenesis and progression of multiple cancers [26-33]. YTHDF2, an m6A reader, usually recognizes m6A in the 3'-UTR of mRNA, leading to mRNA degradation [34]. The contribution of YTHDF2 gene to oncogenesis has been partly clarified, whereas correlations between YTHDF2 gene SNPs and Wilms tumor risk have not been analyzed. The current study addresses the association between YTHDF2 gene SNPs and Wilms tumor risk among children of Chinese ancestry.
Materials and Methods
Study subjects
We successfully enrolled 450 cases and 1317 controls to participate in this project. The cases were newly diagnosed and histopathologically confirmed to be Wilms tumor. Control subjects were healthy volunteers with no underlying medical disorder. Controls and cases were frequency-matched by geographically ethnicity, age, and gender. All subjects were Han Chinese children to lessen the genetic background differences. All subjects' guardians provided written consent before accepting any study-related activity. Our research was approved by the Institutional Review Board of the participating hospitals and conducted in accordance with the principles of the Declaration of Helsinki.
Polymorphism selection and genotyping
The selection of YTHDF2 gene rs3738067 A>G was based on previously described criteria [35-37]. Selection criteria were briefly depicted below: (1) the minor allele frequency (MAF) reported in HapMap was > 5% for Chinese Han subjects; (2) putative functional potentials SNPs located in the 5'- flanking region, exon, 5'- untranslated region (5' UTR), and 3' UTR, which might affect transcription activity or binding capacity of the microRNA binding site; (3) SNPs in low linkage disequilibrium with each other (R2<0.8). YTHDF2 gene rs3738067 A>G is located in transcription factor binding site (TFBS). DNA was extracted from blood using QIAamp DNA Blood mini kit (QIAGEN Inc., Valencia, CA). Genotyping was carried out using TaqMan technology (Applied Biosystems, Foster City, CA) [36]. The conditions of reactions were set as follow: pre-read stage at 60 °C for 30 seconds, holding stage at 95 °C 10 minutes, repeated 45 cycles each of denaturation at 95 °C for 15 seconds, annealing and extension at 60 °C for 1 minute. For quality control purposes, 10% of the samples genotyped were randomly duplicated blindly. Quality control analysis showed a concordance rate of 100%.
Statistical analysis
Chi-square test (for categorical variables) and Student t-test (for continuous variables) were employed to evaluate clinical variables differences in the case and control groups. A goodness-of-fit χ2 test served to know whether SNP rs3738067 A>G in the controls were agreed with Hardy-Weinberg equilibrium (HWE). The association between rs3738067 A>G and Wilms tumor risk was estimated by odds ratios (ORs) with 95% confidence intervals (CIs) calculated by logistic regression analyses. Statistical analyses were performed with SAS v10.0 (SAS Institute Inc., Cary, NC) and statistical significance was considered when P < 0.05.
Results
Population characteristics
Baseline characteristics of Wilms tumor cases and controls are shown in Table S1. 450 cases and 1317 controls were well matched in terms of age (P=0.668) and gender (P=0.157). Among the cases, 137 cases (30.44%) were classified into clinical stage I, 122 (27.11%) into clinical stage II, 119 (26.44%) into clinical stage III, 54 (12.00%) into clinical stage IV, and 18 (4.00%) could not be classified.
Association between YTHDF2 rs3738067 A>G polymorphism and Wilms tumor risk
A total of 441 cases and 1316 controls were successfully genotyped. The genotype of rs3738067 A>G and its association with Wilms tumor risk is shown in Table 1. The P value of HWE in control population of rs3738067 A>G was 0.860, meaning no violation of HWE. No significant difference in frequencies of genotype at rs3738067 A>G was observed between cases and controls (AG vs. AA: adjusted OR=0.91, 95% CI=0.73-1.15, P=0.444; GG vs. AA: adjusted OR=1.00, 95% CI=0.64-1.54, P=0.981; additive: adjusted OR=0.96, 95% CI=0.80-1.14, P=0.619; dominant: adjusted OR=0.93, 95% CI=0.75-1.15, P=0.491; recessive: adjusted OR=1.03, 95% CI=0.67-1. 85, P=0.892).
Association between YTHDF2 rs3738067 A>G polymorphism and Wilms tumor risk
| Genotype | Cases (N=441) | Controls (N=1316) | P a | Crude OR (95% CI) | P | Adjusted OR (95% CI) b | P b |
|---|---|---|---|---|---|---|---|
| rs3738067 (HWE=0.860) | |||||||
| AA | 253 (57.37) | 731 (55.55) | 1.00 | 1.00 | |||
| AG | 158 (35.83) | 498 (37.84) | 0.92 (0.73-1.15) | 0.457 | 0.91 (0.73-1.15) | 0.444 | |
| GG | 30 (6.80) | 87 (6.61) | 1.00 (0.64-1.55) | 0.987 | 1.00 (0.64-1.54) | 0.981 | |
| Additive | 0.632 | 0.96 (0.80-1.14) | 0.632 | 0.96 (0.80-1.14) | 0.619 | ||
| Dominant | 188 (42.63) | 585 (44.45) | 0.505 | 0.93 (0.75-1.15) | 0.505 | 0.93 (0.75-1.15) | 0.491 |
| Recessive | 411 (93.20) | 1229 (93.39) | 0.889 | 1.03 (0.67-1.59) | 0.888 | 1.03 (0.67-1.58) | 0.892 |
HWE, Hardy-Weinberg equilibrium; OR, odds ratio; CI, confidence interval.
a χ2 test for genotype distributions between Wilms tumor patients and cancer-free controls.
b Adjusted for age and gender.
Stratification analysis
We further performed stratification analysis based on age, gender, and clinical stages (Table 2). Similarly, we did not observe any association between the rs3738067 A>G polymorphism and Wilms tumor risk in all subgroups.
Stratify analysis of YTHDF2 rs3738067 A>G polymorphism with Wilms tumor risk
| Variables | rs3738067 (case/control) | Crude OR | P | Adjusted OR a | P a | |
|---|---|---|---|---|---|---|
| AA | AG/GG | (95% CI) | (95% CI) | |||
| Age, month | ||||||
| ≤18 | 84/272 | 59/228 | 0.84 (0.58-1.22) | 0.357 | 0.86 (0.59-1.25) | 0.431 |
| >18 | 169/459 | 129/357 | 0.98 (0.75-1.28) | 0.891 | 1.00 (0.76-1.31) | 0.990 |
| Gender | ||||||
| Females | 111/320 | 96/248 | 1.12 (0.81-1.54) | 0.501 | 1.13 (0.82-1.55) | 0.467 |
| Males | 142/411 | 92/337 | 0.79 (0.59-1.07) | 0.123 | 0.79 (0.59-1.07) | 0.129 |
| Clinical stages | ||||||
| I | 83/731 | 54/585 | 0.81 (0.57-1.17) | 0.259 | 0.82 (0.57-1.17) | 0.274 |
| II | 70/731 | 49/585 | 0.88 (0.60-1.28) | 0.491 | 0.88 (0.60-1.28) | 0.500 |
| III | 63/731 | 54/585 | 1.07 (0.73-1.57) | 0.723 | 1.05 (0.72-1.54) | 0.809 |
| IV | 29/731 | 23/585 | 0.99 (0.57-1.73) | 0.975 | 0.98 (0.60-1.71) | 0.937 |
| I+II | 153/731 | 103/585 | 0.84 (0.64-1.11) | 0.214 | 0.84 (0.64-1.11) | 0.221 |
| III+IV | 92/731 | 77/585 | 1.05 (0.76-1.44) | 0.785 | 1.02 (0.74-1.41) | 0.901 |
OR, odds ratio; CI, confidence interval.
a Adjusted for age and gender, omitting the corresponding variable.
Discussion
The current knowledge of genetic predisposition to Wilms tumor is incomplete. SNPs in m6A-related genes are highly implicated in the risk of cancer. We hypothesized YTHDF2 gene SNPs may also influence the risk of Wilms tumor. This pilot study provides the first indication that YTHDF2 gene rs3738067 A>G could not impact Wilms tumor risk in Chinese children.
The final consequences of m6A modification on mRNA fate are executed by “reader” proteins. These proteins mainly included the YTH family (YTHDC1-2 and YTHDF1-3), HNRNPA2B1, and eIF3. YTHDF2 recognizes m6A mRNA within the GACU/A consensus to induce degradation of methylated transcripts [38]. Cytoplasmic YTHDF1 and YTHDF3 could bind to m6A to initiate the translation of m6A-containing transcripts [39], while IGF2BP protein could enhance the stability of target mRNA [40]. Growing evidence has been added to support the critical role of YTHDF2 in the regulation of cancer cell proliferation and migration. Jasmin Paris et al. [41] found that YTHDF2 is highly expressed across multiple human acute myeloid leukemia (AML) and is required for initiation and propagation in AML. YTHDF2 shortens the half-life of various m6A transcripts that contribute to the overall integrity of self-renewing leukemic stem cells (LSCs) function. Therefore, YTHDF2 could be treated as a unique therapeutic target for AML therapy. In hepatocellular carcinoma (HCC), YTHDF2 was found to be reversely associated with the survival of patients. Knockdown of YTHDF2 resulted in impaired stemness in liver cancer cells. Mechanistically, YTHDF2 could regulate m6A methylation of OCT4 mRNA and thus promote the liver cancer stem cell phenotype and HCC metastasis [42]. Xie et al. [43] found that the METTL3/YTHDF2 m6A axis contributed to bladder cancer progression by directly degrading the mRNAs of the tumor suppressors KLF4 and SETD7. YTHDF2 was also found to be upregulated in lung cancer tissues and promotes lung cancer cell growth. Mechanistically, YTHDF2 acts as a lung cancer promoter to facilitate 6-phosphogluconate dehydrogenase (6PGD) mRNA translation through binding to the m6A modification site of 6PGD [44].
Contributions of the m6A gene to cancer are highly acknowledged, yet research on m6A critical gene SNPs on cancer risk is still at the primary stage. Multiple SNPs located in m6A critical genes have been found to impact the risk of cancer. Our previous studies also revealed the involvement of m6A gene SNPs in Wilms tumor [45-47]. For YTHDF2 gene SNPs and cancer risk, only one study has been conducted. In 2020, Meng et al. [48] performed the first genetic association study regarding m6A modification critical gene SNPs and cancer risk. They genotyped 240 SNPs within 20 m6A modification-related genes in samples of 2082 colorectal cancer cases and 2308 healthy controls. One SNP, rs4654320, in the YTHDF2 gene was included in the analysis. However, all the SNPs including YTHDF2 rs4654320 could not predispose to colorectal cancer, except for one SNP rs118049207 in the SND1 gene. In 2012, a genome-wide association study was carried out on Wilms tumor. The authors used cases recruited through oncology clinics in North America to identify genetic variants that confer susceptibility to Wilms tumor. They selected SNPs that demonstrated an association of a significance level of P<5×10-5 for the replication phase. They failed to detect YTHDF2 gene SNPs that were associated with Wilms tumor risk [14]. Until now, no available reports have been carried out to explore the role of YTHDF2 gene SNPs on Wilms tumor risk. Thus, here we set as a pioneer to determine the role of YTHDF2 gene SNPs on Wilms tumor risk. The current clinical analysis indicated YTHDF2 rs3738067 A>G could not impact Wilms tumor risk in Chinese children. We then analyzed the role of rs3738067 A>G in Wilms tumor risk using stratification analysis but still obtained negative results. Several potential reasons may help to interpret these null relationships: 1) the weak impact of SNP rs3738067 A>G; 2) the insufficient statistical power caused by moderate sample size; 3) influence of other potential pertinent factors, including modifications of environmental factors (parental exposures to pesticides, paternal occupation) [49, 50] and genetic-environmental factors.
Among the weaknesses are that the relatively small sample size of the study is underpowered to detect the weak impact of SNPs. In addition, the outcome of variant rs3738067 A>G on Wilms tumor risk was only assessed by genetic analysis. Environmental factors that greatly modified the risk of Wilms tumor remained unaccounted in the current study. Moreover, the risk variant identification here was only conducted in Chinese descendants, whether the effect of YTHDF2 gene rs3738067 A>G can be generalized to other ethnicities needs to be confirmed. Last, the relationship was only determined in the genetic model. The relationship between YTHDF2 and Wilms tumor from the protein level is warranted to be determined.
Taken together, our results suggested that the YTHDF2 rs3738067 A>G polymorphism did not show a significant association with the risk of Wilms tumor in a population of Chinese children. Investigations are warranted to verify this assessment and to further evaluate the underlying role of YTHDF2 rs3738067 A>G on the risk for Wilms tumor.
Abbreviations
LOI: loss of imprinting; LOH: loss of heterozygosity; SNP: single nucleotide polymorphism; m6A: N6-methyladenosine; MAF: minor allele frequency; UTR: untranslated region; TFBS: transcription factor binding site; HWE: Hardy-Weinberg equilibrium; OR: odds ratio; CI: confidence interval; AML: acute myeloid leukemia; LSC: leukemic stem cell; HCC: hepatocellular carcinoma; 6PGD: 6-phosphogluconate dehydrogenase.
Supplementary Material
Supplementary table.
Acknowledgements
This study was funded by grants from the National Natural Science Foundation of China (No: 82003523, 81960294), Natural Science Foundation of Guangdong Province (No: 2019A1515010360), and Guangdong Provincial Key Laboratory of Research in Structural Birth Defect Disease (No: 2019B030301004).
Competing Interests
The authors have declared that no competing interest exists.
References
1. Aldrink JH, Heaton TE, Dasgupta R, Lautz TB, Malek MM, Abdessalam SF. et al. Update on Wilms tumor. J Pediatr Surg. 2019;54:390-7
2. Breslow N, Olshan A, Beckwith JB, Green DM. Epidemiology of Wilms tumor. Med Pediatr Oncol. 1993;21:172-81
3. Bao PP, Li K, Wu CX, Huang ZZ, Wang CF, Xiang YM. et al. [Recent incidences and trends of childhood malignant solid tumors in Shanghai, 2002-2010]. Zhonghua Er Ke Za Zhi. 2013;51:288-94
4. Phelps HM, Kaviany S, Borinstein SC, Lovvorn HN 3rd. Biological Drivers of Wilms Tumor Prognosis and Treatment. Children (Basel). 2018;5:145
5. Rivera MN, Haber DA. Wilms' tumour: connecting tumorigenesis and organ development in the kidney. Nat Rev Cancer. 2005;5:699-712
6. Zhuang Z, Merino MJ, Vortmeyer AO, Bryant B, Lash AE, Wang C. et al. Identical genetic changes in different histologic components of Wilms' tumors. J Natl Cancer Inst. 1997;89:1148-52
7. Hussong JW, Perkins SL, Huff V, McDonald JM, Pysher TJ, Beckwith JB. et al. Familial Wilms' tumor with neural elements: characterization by histology, immunohistochemistry, and genetic analysis. Pediatr Dev Pathol. 2000;3:561-7
8. Cecchetto G, Alaggio R, Scarzello G, Dall'Igna P, Martino A, Bisogno G. et al. Teratoid Wilms' tumor: report of a unilateral case. J Pediatr Surg. 2003;38:259-61
9. Hohenstein P, Pritchard-Jones K, Charlton J. The yin and yang of kidney development and Wilms' tumors. Genes Dev. 2015;29:467-82
10. Haber DA, Buckler AJ, Glaser T, Call KM, Pelletier J, Sohn RL. et al. An internal deletion within an 11p13 zinc finger gene contributes to the development of Wilms' tumor. Cell. 1990;61:1257-69
11. Pelletier J, Bruening W, Li FP, Haber DA, Glaser T, Housman DE. WT1 mutations contribute to abnormal genital system development and hereditary Wilms' tumour. Nature. 1991;353:431-4
12. Treger TD, Chowdhury T, Pritchard-Jones K, Behjati S. The genetic changes of Wilms tumour. Nat Rev Nephrol. 2019;15:240-51
13. Ruteshouser EC, Robinson SM, Huff V. Wilms tumor genetics: mutations in WT1, WTX, and CTNNB1 account for only about one-third of tumors. Genes Chromosomes Cancer. 2008;47:461-70
14. Turnbull C, Perdeaux ER, Pernet D, Naranjo A, Renwick A, Seal S. et al. A genome-wide association study identifies susceptibility loci for Wilms tumor. Nat Genet. 2012;44:681-4
15. Fu W, Zhuo Z, Hua RX, Fu K, Jia W, Zhu J. et al. Association of KRAS and NRAS gene polymorphisms with Wilms tumor risk: a four-center case-control study. Aging (Albany NY). 2019;11:1551-63
16. Liu P, Zhuo Z, Li W, Cheng J, Zhou H, He J. et al. TP53 rs1042522 C>G polymorphism and Wilms tumor susceptibility in Chinese children: a four-center case-control study. Biosci Rep. 2019;39:BSR20181891
17. Ferrara M, Capozzi L, Russo R. Impact of the MTHFR C677T polymorphism on risk of Wilms tumor: case-control study. J Pediatr Hematol Oncol. 2009;31:256-8
18. Zhuo Z, Fu W, Liu J, Cheng J, Zhou H, Zhang J. et al. LIN28A gene polymorphisms confer Wilms tumour susceptibility: A four-centre case-control study. J Cell Mol Med. 2019;23:7105-10
19. Maciaszek JL, Oak N, Nichols KE. Recent advances in Wilms tumor predisposition. Hum Mol Genet. 2020;29:R138-49
20. Cresswell GD, Apps JR, Chagtai T, Mifsud B, Bentley CC, Maschietto M. et al. Intra-Tumor Genetic Heterogeneity in Wilms Tumor: Clonal Evolution and Clinical Implications. EBioMedicine. 2016;9:120-9
21. Balacco DL, Soller M. The m(6)A Writer: Rise of a Machine for Growing Tasks. Biochemistry. 2019;58:363-78
22. Wei CM, Gershowitz A, Moss B. Methylated nucleotides block 5' terminus of HeLa cell messenger RNA. Cell. 1975;4:379-86
23. Liu L, Wang Y, Wu J, Liu J, Qin Z, Fan H. N(6)-Methyladenosine: A Potential Breakthrough for Human Cancer. Mol Ther Nucleic Acids. 2020;19:804-13
24. Lan Q, Liu PY, Haase J, Bell JL, Huttelmaier S, Liu T. The Critical Role of RNA m(6)A Methylation in Cancer. Cancer Res. 2019;79:1285-92
25. Chen XY, Zhang J, Zhu JS. The role of m(6)A RNA methylation in human cancer. Mol Cancer. 2019;18:103
26. Cai J, Yang F, Zhan H, Situ J, Li W, Mao Y. et al. RNA m(6)A Methyltransferase METTL3 Promotes The Growth Of Prostate Cancer By Regulating Hedgehog Pathway. Onco Targets Ther. 2019;12:9143-52
27. Chao Y, Shang J, Ji W. ALKBH5-m(6)A-FOXM1 signaling axis promotes proliferation and invasion of lung adenocarcinoma cells under intermittent hypoxia. Biochem Biophys Res Commun. 2020;521:499-506
28. Chen X, Xu M, Xu X, Zeng K, Liu X, Sun L. et al. METTL14 Suppresses CRC Progression via Regulating N6-Methyladenosine-Dependent Primary miR-375 Processing. Mol Ther. 2019;28:599-612
29. Cheng M, Sheng L, Gao Q, Xiong Q, Zhang H, Wu M. et al. The m(6)A methyltransferase METTL3 promotes bladder cancer progression via AFF4/NF-kappaB/MYC signaling network. Oncogene. 2019;38:3667-80
30. Dahal U, Le K, Gupta M. RNA m6A methyltransferase METTL3 regulates invasiveness of melanoma cells by matrix metallopeptidase 2. Melanoma Res. 2019;29:382-9
31. Hou J, Zhang H, Liu J, Zhao Z, Wang J, Lu Z. et al. YTHDF2 reduction fuels inflammation and vascular abnormalization in hepatocellular carcinoma. Mol Cancer. 2019;18:163
32. Li J, Han Y, Zhang H, Qian Z, Jia W, Gao Y. et al. The m6A demethylase FTO promotes the growth of lung cancer cells by regulating the m6A level of USP7 mRNA. Biochem Biophys Res Commun. 2019;512:479-85
33. Cui X, Wang Z, Li J, Zhu J, Ren Z, Zhang D. et al. Cross talk between RNA N6-methyladenosine methyltransferase-like 3 and miR-186 regulates hepatoblastoma progression through Wnt/beta-catenin signalling pathway. Cell Prolif. 2020: e12768.
34. Wang X, Lu Z, Gomez A, Hon GC, Yue Y, Han D. et al. N6-methyladenosine-dependent regulation of messenger RNA stability. Nature. 2014;505:117-20
35. He J, Wang F, Zhu J, Zhang R, Yang T, Zou Y. et al. Association of potentially functional variants in the XPG gene with neuroblastoma risk in a Chinese population. J Cell Mol Med. 2016;20:1481-90
36. Zhuo ZJ, Liu W, Zhang J, Zhu J, Zhang R, Tang J. et al. Functional Polymorphisms at ERCC1/XPF Genes Confer Neuroblastoma Risk in Chinese Children. EBioMedicine. 2018;30:113-9
37. Zhuo Z, Lu H, Zhu J, Hua RX, Li Y, Yang Z. et al. METTL14 Gene Polymorphisms Confer Neuroblastoma Susceptibility: An Eight-Center Case-Control Study. Mol Ther Nucleic Acids. 2020;22:17-26
38. Du H, Zhao Y, He J, Zhang Y, Xi H, Liu M. et al. YTHDF2 destabilizes m(6)A-containing RNA through direct recruitment of the CCR4-NOT deadenylase complex. Nat Commun. 2016;7:12626
39. Shi H, Wang X, Lu Z, Zhao BS, Ma H, Hsu PJ. et al. YTHDF3 facilitates translation and decay of N(6)-methyladenosine-modified RNA. Cell Res. 2017;27:315-28
40. Huang H, Weng H, Sun W, Qin X, Shi H, Wu H. et al. Recognition of RNA N(6)-methyladenosine by IGF2BP proteins enhances mRNA stability and translation. Nat Cell Biol. 2018;20:285-95
41. Paris J, Morgan M, Campos J, Spencer GJ, Shmakova A, Ivanova I. et al. Targeting the RNA m(6)A Reader YTHDF2 Selectively Compromises Cancer Stem Cells in Acute Myeloid Leukemia. Cell Stem Cell. 2019;25:137-48.e6
42. Zhang C, Huang S, Zhuang H, Ruan S, Zhou Z, Huang K. et al. YTHDF2 promotes the liver cancer stem cell phenotype and cancer metastasis by regulating OCT4 expression via m6A RNA methylation. Oncogene. 2020;39:4507-18
43. Xie H, Li J, Ying Y, Yan H, Jin K, Ma X. et al. METTL3/YTHDF2 m(6) A axis promotes tumorigenesis by degrading SETD7 and KLF4 mRNAs in bladder cancer. J Cell Mol Med. 2020;24:4092-104
44. Sheng H, Li Z, Su S, Sun W, Zhang X, Li L. et al. YTH domain family 2 promotes lung cancer cell growth by facilitating 6-phosphogluconate dehydrogenase mRNA translation. Carcinogenesis. 2020;41:541-50
45. Hua RX, Liu J, Fu W, Zhu J, Zhang J, Cheng J. et al. ALKBH5 gene polymorphisms and Wilms tumor risk in Chinese children: A five-center case-control study. J Clin Lab Anal. 2020;34:e23251
46. Ma L, Hua RX, Lin H, Zhu J, Fu W, Lin A. et al. The contribution of WTAP gene variants to Wilms tumor susceptibility. Gene. 2020;754:144839
47. Lin A, Zhou M, Hua RX, Zhang J, Zhou H, Li S. et al. METTL3 polymorphisms and Wilms tumor susceptibility in Chinese children: A five-center case-control study. J Gene Med. 2020;22:e3255
48. Meng Y, Li S, Gu D, Xu K, Du M, Zhu L. et al. Genetic variants in m6A modification genes are associated with colorectal cancer risk. Carcinogenesis. 2020;41:8-17
49. Sharpe CR, Franco EL, de Camargo B, Lopes LF, Barreto JH, Johnsson RR. et al. Parental exposures to pesticides and risk of Wilms' tumor in Brazil. Am J Epidemiol. 1995;141:210-7
50. Wilkins JR 3rd, Sinks TH Jr. Occupational exposures among fathers of children with Wilms' tumor. J Occup Med. 1984;26:427-35
Author contact
Corresponding authors: Shan Yan, Yunnan Key Laboratory of Stem Cell and Regenerative Medicine, Biomedical Engineering Research Center, Kunming Medical University, No. 1168 Chunrongxi Road, Kunming 650500, Yunnan, China, E-mail: yanshanedu.cn; or Jing He, Department of Pediatric Surgery, Guangzhou Institute of Pediatrics, Guangdong Provincial Key Laboratory of Research in Structural Birth Defect Disease, Guangzhou Women and Children's Medical Center, Guangzhou Medical University, 9 Jinsui Road, Guangzhou 510623, Guangdong, China, E-mail: hejing198374com.

 Global reach, higher impact
Global reach, higher impact