Impact Factor
ISSN: 1837-9664
J Cancer 2022; 13(1):62-75. doi:10.7150/jca.66016 This issue Cite
Research Paper
Ago-RIP Sequencing Identifies New MicroRNA-449a-5p Target Genes Increasing Sorafenib Efficacy in Hepatocellular Carcinoma
1. Department of Human Genetics, Hannover Medical School, Hannover, Germany.
2. Research Core Unit Genomics, Hannover Medical School, Hannover, Germany.
3. Hannover Unified Biobank (HUB), Hannover Medical School, Hannover, Germany.
Abstract
BACKGROUND: Patients with hepatocellular carcinoma (HCC) have very limited treatment options. For the last fourteen years, the multi-tyrosine kinase inhibitor sorafenib has been used as standard-of-care therapeutic agent in advanced HCC. Unfortunately, drug resistance develops in many cases. Therefore, we aimed to find a way to mitigate drug resistance and to improve the sorafenib efficacy in HCC cells. MicroRNAs play a significant role in targeting genes involved in tumor control suggesting microRNA/sorafenib combination therapy as a promising treatment option in advanced HCC.
METHODS: MiR-449a-5p target genes were identified by Ago-RIP sequencing and validated by luciferase reporter assays and expression analyses. Target gene expression and survival data were analyzed in public HCC datasets. Tumor-relevant functional effects of miR-449a-5p and its target genes as well as their impact on the effects of sorafenib were analyzed using in vitro assays. An indirect transwell co-culture system was used to survey anti-angiogenic effects of miR-449a-5p.
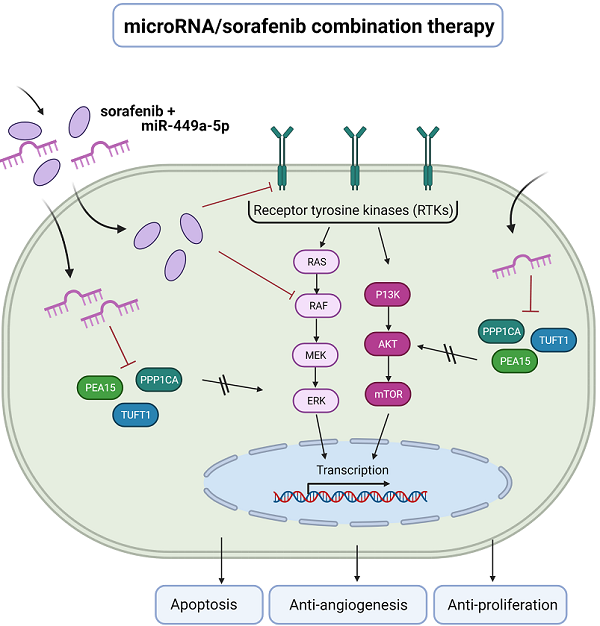
RESULTS: PEA15, PPP1CA and TUFT1 were identified as direct target genes of miR-449a-5p. Overexpression of these genes correlated with a poor outcome of HCC patients. Transfection with miR-449a-5p and repression of miR-449a-5p target genes inhibited cell proliferation and angiogenesis, induced apoptosis and reduced AKT and ERK signaling in HLE and Huh7 cells. Importantly, miR-449a-5p potentiated the efficacy of sorafenib in HCC cells via downregulation of PEA15, PPP1CA and TUFT1.
CONCLUSIONS: This study provides detailed insights into the targetome and regulatory network of miR-449a-5p. Our results demonstrate for the first time that targeting PEA15, PPP1CA and TUFT1 via miR-449a overexpression could have significant implications in counteracting sorafenib resistance suggesting miR-449a-5p as a promising candidate for a microRNA/sorafenib combination therapy.
Keywords: Liver cancer, microRNA combination therapy, drug resistance, microRNA target genes, Ago-RIP sequencing, multi-tyrosine kinase inhibitor

 Global reach, higher impact
Global reach, higher impact