Impact Factor
ISSN: 1837-9664
J Cancer 2022; 13(5):1512-1522. doi:10.7150/jca.66241 This issue Cite
Research Paper
Multi-omics analysis identifies distinct subtypes with clinical relevance in lung adenocarcinoma harboring KEAP1/NFE2L2
1. Department of Thoracic Surgery, Shanghai Pulmonary Hospital, Tongji University, Shanghai, China.
2. Department of Thoracic Surgery, Zhongshan Hospital, Fudan University, Shanghai, China.
3. Department of General Surgery, Xingtai Third Hospital, Hebei Province, China.
4. Department of Cardiothoracic Surgery, Zhangqiu District People's Hospital, Shandong Province, China.
5. Department of Thoracic Surgery, Shanghai General Hospital, Shanghai, China.
#These authors contributed equally to this work.
Abstract
Backgrounds: Lung adenocarcinoma is one of the most common malignant tumors, in which KEAP1-NFE2L2 pathway is altered frequently. The biological features and intrinsic heterogeneities of KEAP1/NFE2L2-mutant lung adenocarcinoma remain unclear.
Methods: Multiplatform data from The Cancer Genome Atlas (TCGA) were acquired to identify two subtypes of lung adenocarcinoma harboring KEAP1/NFE2L2 mutations.
Bioinformatic analyses, including immune microenvironment, methylation level and mutational signature, were performed to characterize the intrinsic heterogeneities. Meanwhile, initial results were validated by using in silico assessment of common lung adenocarcinoma cell lines, which revealed consistent features of mutant subtypes. Furthermore, drug sensitivity screening was conducted based on public datasets.
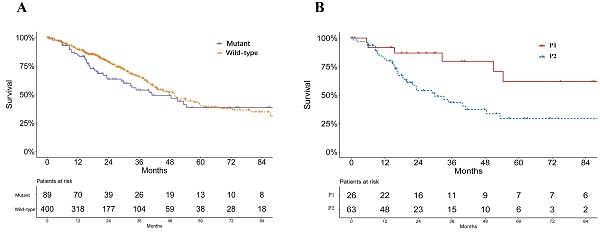
Results: Two mutant subtypes (P1 and P2) of 89 patients were identified in TCGA. P2 patients had significantly higher levels of smoking and worse survival compared with P1 patients. The P2 subset was characterized by active immune microenvironment and more smoking-induced genomic alterations with respect to methylation and somatic mutations. Validations of the corresponding features in 20 mutant cell lines were achieved. Several compounds which were sensitive to mutant subtypes of lung adenocarcinoma were identified, such as inhibitors of PI3K/Akt and IGF1R signaling pathways.
Conclusions: KEAP1/NFE2L2-mutant lung adenocarcinoma showed potential heterogeneities. The intrinsic heterogeneities of KEAP1/NFE2L2 were associated with immune microenvironment and smoking-related genomic aberrations.
Keywords: lung adenocarcinoma, KEAP1, mutation, NFE2L2

 Global reach, higher impact
Global reach, higher impact