Impact Factor ISSN: 1837-9664
J Cancer 2023; 14(6):881-894. doi:10.7150/jca.82916 This issue Cite
Research Paper
MicroRNA 133A Regulates Cell Proliferation, Cell Migration, and Apoptosis in Colorectal Cancer by Suppressing CDH3 Expression
1. Department of Pathology, School of Medicine, Wonkwang University, Iksan, Chonbuk 54538, Republic of Korea
2. Digestive Disease Research Institute, Wonkwang University, Iksan, Chonbuk 54538, Republic of Korea
Received 2023-1-25; Accepted 2023-3-20; Published 2023-4-9
Abstract
MicroRNAs are endogenous, non-coding RNA that play an essential role in colorectal carcinoma (CRC) pathogenesis by targeting specific genes. This research aimed to determine and validate the target genes of the MIR133A associated with CRC. We verified that cadherin 3 (CDH3) is the direct target gene of MIR133A using a luciferase reporter assay, quantitative RT-PCR, and western blot analyses. CDH3 mRNA and protein expression were reduced significantly in CRC cells after transfection with MIR133A or siCDH3. We also verified that MIR133A regulated CDH3-mediated catenin, matrix metalloproteinase, apoptosis, and the epithelial-mesenchymal transition (EMT) pathway. Knockdown of CDH3 in CRC cell lines by siCDH3 produced similar results. Compared with adjacent non-tumor tissues, CDH3 protein expression was upregulated in CRC tissues, which is further confirmed by immunohistochemistry. Additionally, molecular and functional studies revealed that cell viability, migration, and colony formation were significantly reduced, and apoptosis was increased in CRC cell lines transfected with MIR133A or siCDH3. Our results suggest that MIR133A regulates CDH3 expression in human CRC.
Keywords: MIR133A, CDH3, Cell viability, Colorectal Cancer, EMT
Introduction
Colorectal carcinoma (CRC) is a common cancer of the digestive tract with a high death rate [1]. In 2020, about 19.3 million novel cases and 10 million cancer-related mortalities were recorded worldwide. CRC comprises about 10% of new cases and 9.4% of cancer-related deaths, which are increasing globally [2]. Several studies have shown a global prevalence of CRC of about 4-5%. CRC risk factor comprises age, inflammatory bowel disease (IBD) history, sedentary lifestyle, obesity, diet, smoking, and alcohol [3]. In many developing countries, high incidence and poor detection of CRC have increased the need for early diagnosis and treatment. Hence, different prognostic biomarkers, including treatments for CRC, should be explored.
Cadherins are cell adhesion molecules that help cells to adhere to each other and help cells migrate via epithelial-mesenchymal transition (EMT) [4,5]. They control calcium-dependent cellular adhesion and bind to various cell types and the extracellular matrix (ECM), maintaining cellular structure. Cadherin 3 (CDH3, also known as PCAD) has a molecular weight of 118kDa; comprises extracellular, transmembrane, and cytoplasmic domains; and promotes homotypic interactions. In normal cells, CDH3 regulates cell growth and migration, but CDH3 may have opposing effects in tumors depending on type: CDH3 promotes tumorigenesis in pancreatic, breast, and gastric cancer but has the opposite effect in non-small cell lung carcinoma, liver cancer, and melanoma [6,7]. A previous study reported that CDH3 is upregulated in colon adenocarcinoma, and patients with high CDH3 have a good prognosis for cancer [8,9]. The aberrant expression of CDH3 in the colonic mucosa of CRC is the effect of abnormal gene upregulation through CDH3 gene promoter hypomethylation [10,11].
MicroRNAs (miRNAs) are single-stranded, non-coding RNA that are expressed endogenously and attach to the target mRNA's 3′ untranslated regions (3′ UTR) and regulate gene expression by mRNA degradation or translational inhibition. Normal expression of miRNAs is tissue-specific and regulates various pathways involved in growth, cell differentiation, migration, and apoptosis, whereas aberrant expression of miRNAs leads to the development of human cancer [12]. While miRNAs play a crucial role in cancer biology, the molecular mechanism underlying miRNA expression in cancer is not well understood. Many studies have demonstrated associations of miRNAs, including microRNA 133A (MIR133A, also known as has-miR-133A), with colon cancer pathogenesis [13,14,15]. In our earlier research, we reported that MIR133A expression was reduced in CRC, which correlates with poor diagnosis in CRC patients [16,17]. The CDH3 gene was identified as one of the candidate targets of MIR133A in our previous study using MIR133A1 knock-in cells and was confirmed using TargetScan, miRanda, and miRWalk algorithms [16].
Here, we reported CDH3 as a direct target gene of MIR133A. We showed the association of MIR133A and CDH3 via human CRC cell lines and tissues. We have also used MTT, colony formation, migration assays, and flow cytometry to further verify the correlation. Our results suggest that MIR133A regulates CDH3 in human colorectal cancer and may be helpful for CRC.
Materials and Methods
Patient sample
This study used tissue samples from the Biobank of Wonkwang University Hospital. With Institutional Review Board permission and informed consent from participants (WKIRB-202006-BR-023), we received four CRC tissue and four adjacent normal samples from colon cancer patients [2 TNM stage 3 (1 female and 1 male) and 2 TNM stage 4 (1 female and 1 male)]. The mean ages of the colon cancer patients were 60. These samples were used to examine protein expression using western blotting and in situ CDH3 expression using immunohistochemistry.
Cell culture
Human CRC SW480, Caco2, and LoVo cell lines were acquired from Korean Cell Line Bank (KCLB, Seoul, Korea) or the American Type Culture Collection (ATCC). The cells were cultured in RPMI-1640 (Hyclone, UTAH, USA) or MEM Alpha (Hyclone, UTAH, USA) supplemented with 10% fetal bovine serum (FBS) in 5% CO2 at 37⁰C in a humidified atmosphere.
RNA extraction and quantitative real-time PCR
The RNA extraction and quantitative RT-PCR (qRT-PCR) were done as our previous study [18,19]. Each experiment was analyzed three times. The primers that were used are in Supplementary Table 1.
Transfection of oligonucleotides
The SW480 and Caco2 cells were cultured in RPMI and MEM Alpha medium. MIR133A mimic (hsa-MIR-133A, pre-miR miRNA precursor AM17100, product ID: PM12946) and negative control (Ambion, USA) were added at 50 nmol/mL for transfection, which was performed as previously described [20,21]. Lipofectamine RNAiMAX (Invitrogen) and siPORT NeoFX (Ambion) were used as transfecting agents. The CDH3 small interfering RNA (siRNA)(Sigma-Aldrich) and negative control siRNA (Ambion) were used according to the manufacturer's protocols. After transfection, cells were cultured for 72 hours for protein analyses.
Plasmid constructs and luciferase assay
Wild-type (WT) or mutant-type (MT) constructs of CDH3 3′ UTR were created using PCR primers as indicated in Supplementary Table 1. Then, constructs were ligated into the pmirGLO™ Dual-Luciferase miRNA target expression vector (Promega, USA). All constructs were confirmed by DNA sequencing. In this assay, 5 × 104 SW480 cells/well were seeded into 24-well plates and transfected with WT-CDH3 or MT-CDH3 constructs and with MIR133A or MIRNC (negative control) using lipofectamine (Invitrogen) and siPORTTM NeoFXTM transfecting agents (Ambion). The results were evaluated as previously described [22,23]. All experiments were repeated three times.
Western blot analyses
The SW480, Caco2, and LoVo cells (2 × 105 cells/well) were cultured in cell culture plates for 72 hours after transfection, followed by lysis with RIPA buffer, and lysates were centrifuged at 12000 RPM (4⁰C for 15 min). Protein concentration was calculated using BSA reagents (Thermo Fisher, USA). An equal amount of cell protein was boiled in Laemmli buffer and subjected to SDS-PAGE, followed by transfer to a PVDF membrane (Millipore Corporation, USA). The PVDF membranes were incubated with 3% BSA (BOVOGEN Biologicals, Australia) in 1× PBS with 1% Tween® 20 (iNtRON Biotechnology) for one hour at RT. Membranes were incubated overnight at 4⁰C with primary anti-CDH1 (3195S, 1:1000), -VIM (5741S, 1:1000), -CTNNB1 (610153, 1:2000), -GSK3B (NBP1-47470, 1:1000), -MMP1 (sc-373908, 1:500), -MMP2 (sc-13595, 1:500), -BCL2 (sc-7382, 1:500), -GAPDH (sc-47724, 1:2500), -CDH3 (sc74545, 1:500), -BIRC5 (sc-17779,1:500), -CTNND1 (sc-23873, 1:500), -CDH2 (sc-8424, 1:500), ‑CASP3(#9661, 1:1000), -CASP9 (#9502S, 1:1000), -CASP8 (#9746L, 1:1000), -RIPK3 (sc-374639, 1:500), and -MLKL (sc-293201, 1:500) antibodies and washed with PBST three times for 8 min/wash. After washing, membranes were incubated for one hour at RT with a secondary antibody (goat anti-rabbit, sc-2004, 1:5000, goat anti-mouse antibody 1:10000). Proteins were evaluated using ECL solution (Millipore) and FluorChem™-E System (Cell Biosciences, USA). For reblotting, a stripping buffer (Thermo Fisher, USA) was used. GAPDH was used as the control. Quantification was performed using ImageJ software. Each experiment was analyzed three times.
MTT assay of cell viability
The SW480 cell line (2 × 104) was used to assess cell viability. Transfection of SW480 cell line was done with 50 nM MIR133A or MIRNC in a 96-well plate. The MTT reagent (Sigma-Aldrich, USA) was dissolved at a concentration of 0.5 mg/ml in RPMI medium. After removing the RPMI media, MTT solution was added in every well. After covering in aluminum foil, cell plates were placed in 37⁰C incubator for three hours. After removing the media, DMSO (Duchefa Biochemie, Netherlands) was used to dissolve MTT crystals. The plates were then placed in a shaker for 30 min, and optical density was measured at 560 nm using a GloMax®-Multi+ Detection System (Promega). Each experiment was analyzed three times.
Scratch wound-healing assay
A monolayer of SW480 cells cultured in a 6-well plate was scratched to form a wound using a sterilized 200 µm pipette tip. Scratching was performed after 24 hours of transfection, and serum-free media was used to wash the wells to remove cell debris. The plates were then incubated for 72 hours. Representative photographs were obtained at 0, 24, 48, and 72 hours using a Canon DS-126431 camera system mounted in a phase-contrast Nikon TS100 microscope (10×). Each experiment was analyzed three times.
Colony-forming assay
The SW480 cells (1000 cells/well) were transfected with MIR133A (50nM) or MIRNC in a 6-well plate and cultured in an incubator with 5% CO2 at 37⁰ C for 15-20 days for the formation of colonies. Media was changed every 72 hours, and cells were washed with 1X PBS. Colonies were stained by using 0.5 ml of 6% glutaraldehyde and 0.5% crystal violet, followed by incubation at RT for 30 minutes and staining solution was discarded, and the cells were cleansed with distilled water and counted after drying in air. Each experiment was analyzed three times.
Immunohistochemistry (IHC)
Human colon sections were formalin-fixed, implanted in paraffin, and cut for IHC reaction. The expression of CDH3 was evaluated as described previously [21,24].
Apoptosis analysis (flow cytometry)
The SW480 (2 × 105 cells/well) were transfected with MIR133A (50nM) or MIRNC in 6-well plates for 72 hours. An Annexin V-FITC Apoptosis Detection Kit (Sigma-Aldrich, St. Louis, MO, USA) was used for flow cytometry as in our previous experiment [18,24].
Statistical analyses
Each experiment was analyzed three times. The results are reported as mean ± standard deviation. Microsoft Excel and GraphPad Prism 8 were used to complete all statistical analyses. Two-tailed Student's t-tests were used for comparison, and P values < 0.05 were considered significant.
Results
CDH3 is a direct target of MIR133A
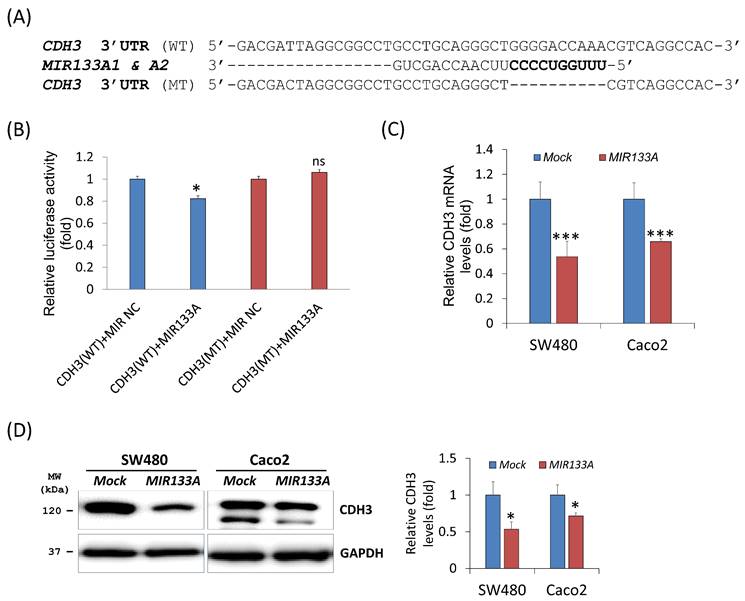
To evaluate the influence of MIR133A on the CDH3 3′ UTR region, we construct a wild-type (WT) CDH3 3′ UTR region (predicted to interact with MIR133A) into a luciferase vector (Fig. 1A). Compared with the negative control, luciferase activity was significantly reduced in the MIR133A-transfected cells (Fig. 1B). Additionally, a mutant-type (MT) CDH3 3′ UTR was constructed by deleting the seed sequence of MIR133A; however, MIR133A-mediated luciferase suppression was not present at the mutant-transfected cells (Fig. 1A, B). Furthermore, qRT-PCR was used to assess CDH3 mRNA expression in MIR133A-transfected cells. With respect to control cells, CDH3 mRNA expression was significantly reduced in MIR133A-transfected cells (Fig. 1C). Likewise, CDH3 protein expression was significantly decreased in the MIR133A-transfected group (Fig. 1D). These findings demonstrate that MIR133A directly targets CDH3.
CDH3 is a direct target of MIR133A (A) Sequence constructs of wild-type (WT) and mutant (MT) MIR133A target sites in the 3′ UTR of CDH3. A human CDH3 3′ UTR containing the WT and MT MIR133A binding constructs was cloned to a luciferase vector. (B) A luciferase reporter plasmid with WT or MT CDH3 3′ UTR was co-transfected into SW480 cells. Luciferase activity was assayed using a dual-luciferase system. Results are shown as the relative firefly luciferase activity normalized to that of renilla luciferase. (C) qRT-PCR analyses of CDH3 mRNA expression in SW480 and Caco2 cells after transfection with MIRNC or MIR133A. Following transfection, cells were incubated for 48 hour. Values are presented as fold change with respect to mock-transfected controls. (D) Western blot analyses of cellular CDH3 protein in SW480 and Caco2 cells after transfection with MIRNC or MIR133A. Following transfection, cells were incubated for 72 hours, and proteins were collected. Data were obtained from three independent experiments, and p values were calculated using t test (ns = not significant, * P < 0.05, *** P < 0.001).
MIR133A modulates the CDH3-mediated EMT pathway
The EMT represents a complex program in which epithelial cells lose their cell identity and attain a mesenchymal phenotype. EMT is also active in cancer cells, leading to apoptosis resistance, tumor metastasis, and the acquisition of cancer stem cell characteristics [25]. As CDH3 has been reported to affect cell migration and invasion [26], we aimed to verify the association between reduction in CDH3 expression mediated by MIR133A and induction of the EMT. Downstream proteins of the EMT pathway in SW480, Caco2, and LoVo cell lines were measured. Western blotting results showed that cadherin 2 (CDH2, also known as N-cadherin) and vimentin (VIM) expression was significantly reduced after MIR133A transfection in CRC cell lines (Fig. 2A). Additionally, CDH3 gene silencing revealed that expression of CDH2 and VIM proteins was significantly reduced (Fig. 2B). The expression of CDH1 (also known as E-cadherin) was not significantly changed in both MIR133A and siCDH3 overexpressed cell lines (Fig. 2A, B).
MIR133A modulates the CDH3-mediated EMT pathway Western blot analyses of CDH1, CDH2, and VIM protein expression in SW480 and Caco2 cell lines after transfection with MIR133A (A) or siCDH3 (B). (C) Western blot analyses of CDH3, CDH1, CDH2, and VIM protein expression in LoVo cell lines after transfection with MIR133A or siCDH3. Data were obtained from three independent experiments, and p values were calculated using t test (ns = not significant, * P < 0.05).
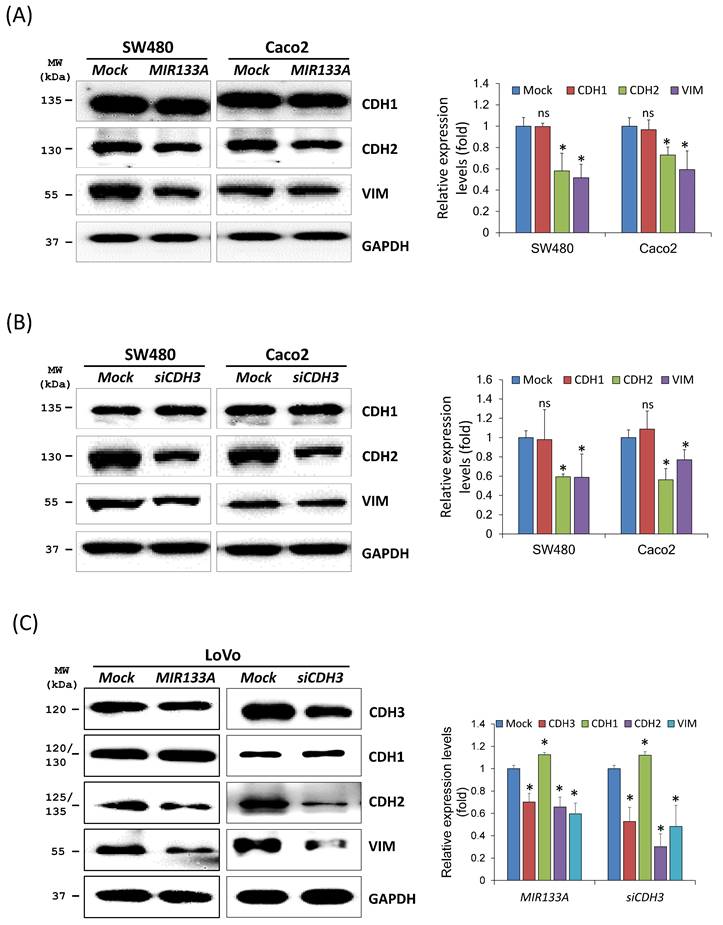
The ability to metastasize was more significant in the LoVo cell line, so we measured changes in CDH1, CDH2, and VIM expression using this line (Fig. 2C). Expression of both CDH2 and VIM was significantly downregulated by MIR133A and siCDH3 transfection, whereas CDH1 expression was upregulated by MIR133A and siCDH3 transfection (Fig. 2C). These data indicate that MIR133A regulates a CDH3-mediated EMT signaling pathway in CRC cells.
MIR133A modulates the catenin pathway in CRC cell lines
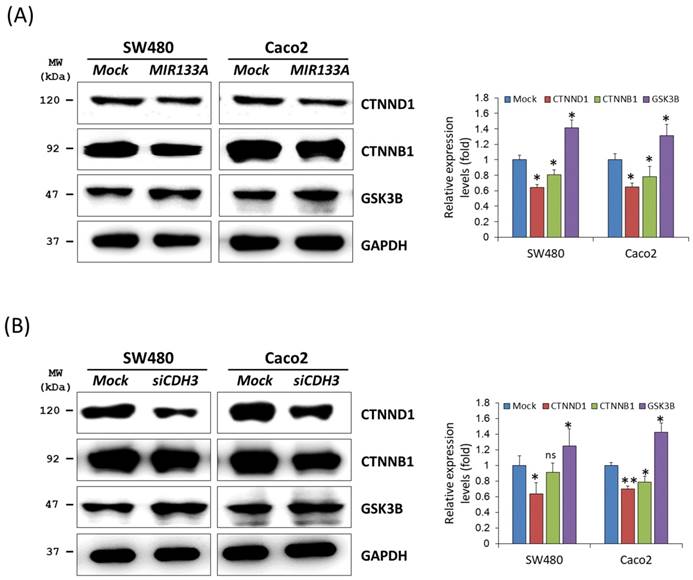
Cadherin mediates cellular adhesion and tissue morphology in association with catenin beta 1 (CTNNB1) [8]. Also, CDH3 is strongly associated with cytoplasmic accumulation of catenin delta 1 (CTNND1, also known as CAS and p120) [27]. To identify functional interactions of MIR133A within the catenin pathway, we carried out a western blot analysis using total cell isolates from SW480 and Caco2 cells. Our results showed that transfection of MIR133A in CRC cell lines significantly reduced CTNND1 and CTNNB1 protein expression in comparison to the mock (Fig. 3A). Additionally, siCDH3 transfection of SW480 and Caco2 cell lines was used to study the interaction of siCDH3 in the catenin pathway and indicated that protein expression of CTNND1and CTNNB1 was downregulated by CDH3 gene silencing (Fig. 3B). In comparison, glycogen synthase kinase 3 beta (GSK3B) was increased after both transfections (Fig. 3A, B).
MIR133A modulates the catenin pathway in CRC cell lines (A) Western blot analyses of CTNND1, CTNNB1, and GSK3B protein expression in SW480 or Caco2 cell lines after transfection with MIR133A. (B) Western blot analyses of CTNND1, CTNNB1, and GSK3B protein expression in SW480 or Caco2 cell lines after transfection with siCDH3. Data were obtained from three independent experiments, and p values were calculated using t test (ns = not significant, * P < 0.05, ** P < 0.01).
MIR133A modulates the apoptosis pathway in CRC cell lines
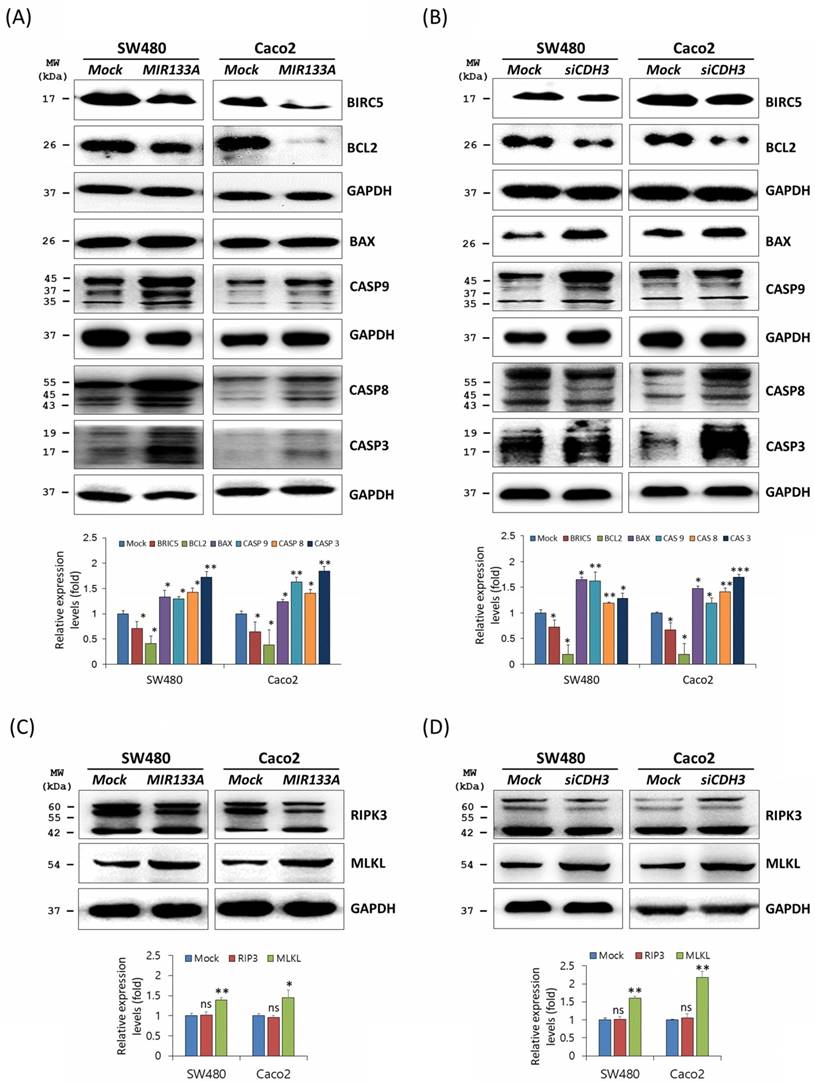
Apoptosis is programmed cell death during normal cell development and is a homeostatic mechanism to maintain cells [28]. To determine the functional interaction of MIR133A in the apoptosis pathway, western blot analyses was performed using SW480 and Caco2 cells. Expression of the BCL2 apoptosis regulator (BCL2) and the baculoviral IAP repeat-containing 5 (BIRC5, as known as API4 and survivin) was significantly reduced, whereas expression of the BCL2-associated X (BAX) apoptosis regulator, caspase 9 (CASP9), CASP8, and CASP3 was upregulated in MIR133A overexpressed SW480 and Caco2 cell lines (Fig. 4A). The data showed that BCL2 and BIRC5 were significantly downregulated and BAX, CASP9, CASP8, and CASP3 were increased by CDH3 gene silencing (Fig. 4B). We also assessed changes in expression of the necroptosis marker receptor-interacting serine/threonine kinase 3 (RIP3) and mixed lineage kinase domain-like pseudokinase (MLKL). The expression of MLKL was upregulated by MIR133A and siCDH3 transfection, whereas there was no change in RIP3 (Fig. 4C, D).
MIR133A modulates the apoptosis pathway in CRC cell lines (A) Western blot analyses of BIRC5, BCL2, BAX, CASP9, CASP8, and CASP3 protein expression in SW480 or Caco2 cell lines after transfection with MIR133A. (B) Western blot analyses of BIRC5, BCL2, BAX, CASP9, CASP8, and CASP3 protein expression in SW480 or Caco2 cell lines after transfection with siCDH3. (C, D) Western blot analyses of RIPK3 and MLKL protein expression after transfection with MIR133A or siCDH3. Data were obtained from three independent experiments, and p values were calculated using t test (ns = not significant, * P < 0.05, ** P < 0.01, *** P < 0.001).
MIR133A modulates the CDH3-mediated MMP1/MMP2 pathway of invasion and metastasis
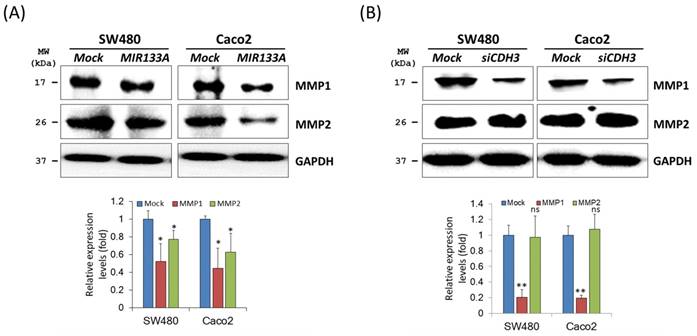
Matrix metalloproteinases (MMPs) are endopeptidases that can degrade ECM components [29]. In breast cancer, overexpression of CDH3 will increase cell invasion and migration and increase MMP1 and MMP2 expression [30]. To investigate the regulation of MIR133A in MMPs of CRC cells, western blot analyses of total cell isolates from SW480 and Caco2 cells were carried out. Expression of MMP1 and MMP2 was significantly reduced in MIR133A overexpressed SW480 and Caco2 cell lines (Fig. 5A). Additionally, siCDH3 transfection revealed that MMP1 was reduced in CRC cell lines, whereas MMP2 expression was unchanged (Fig. 5B). These results indicate that CDH3 regulates only MMP1 expression in colorectal cancer.
MIR133A modulates the CDH3-mediated MMP1/MMP2 pathway of invasion and metastasis (A) Western blot analyses of MMP1 and MMP2 protein expression in SW480 or Caco2 cell lines after transfection with MIR133A or mock transfection. (B) Western blot analyses of MMP1 and MMP2 protein expression in SW480 or Caco2 cell lines after transfection with siCDH3. Data were obtained from three independent experiments, and p values were calculated by t test (ns = not significant, * P < 0.05, ** P < 0.01).
MIR133A inhibits CRC cell growth and cell cycle progression
The functional analysis of MIR133A in SW480 and Caco2 cell lines was investigated using MTT, colony formation, cell migration, and flow cytometry assays. The MTT data confirmed that transfection of a MIR133A or CDH3 siRNA in SW480 and Caco2 cell lines significantly inhibited cell proliferation with respect to mock (Fig. 6A). The colony formation assay in the SW480 cell line indicated that transfection of MIR133A and siCDH3 significantly reduced the number of colonies (Fig. 6B). These data indicate that MIR133A targets CDH3 to suppress cell growth and colony formation in CRC cell lines.
MIR133A inhibits CRC cell growth and cell cycle progression (A) MTT assay showed the reduction of cell viability of CRC cell lines after transfection with MIR133A or siCDH3. (B) Colony formation in SW480 cells was determined using a colony formation assay after transfection with MIR133A and siCDH3. (C) A scratch-wound-healing assay was performed in SW480 cells after transfection with MIR133A and siCDH3. Migration distance was measured at 0, 1, 2, and 3 days after wounding. (D) Flow cytometric analyses of apoptosis or necrosis in SW480 cells transfected with MIR133A and siCDH3. The number of each box indicates the percentage of annexin V-positive and PI-positive cells. Representative data from at least three independent experiments are shown. Each bar represents mean fold change above or below the control (± S.D). Differences were considered statistically significant compared with control (ns = not significant, * P < 0.05, ** P < 0.01, *** P < 0.001).
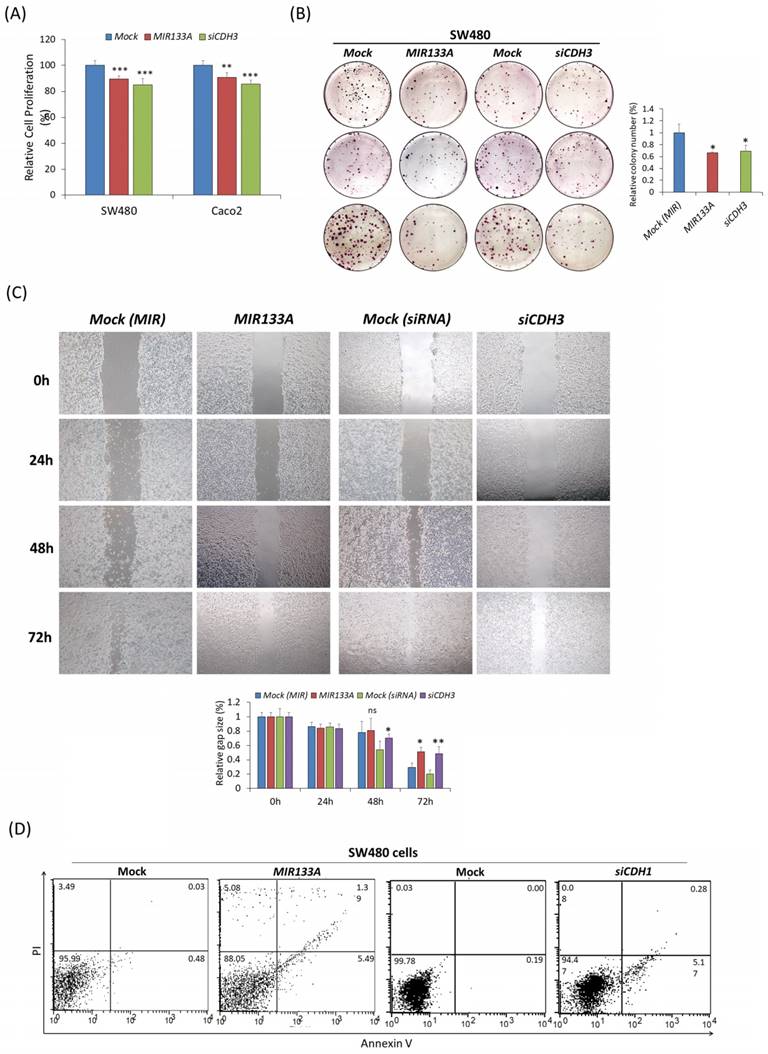
Next, to study the effect of MIR133A on migration activity, SW480 cells were transfected with a MIR133A and siCDH3. The result indicated that migration of SW480 cells was reduced in MIR133A-transfected groups and was decreased significantly 72 hours post-transfection. Cells transfected with CDH3 siRNA showed similar inhibition of migration (Fig. 6C). These findings imply MIR133A regulates CDH3 to limit migration.
Also, apoptosis data in figure 6D revealed that apoptosis and necrosis in SW480 cells were enhanced after MIR133A and siCDH3 transfection compared with the mock (Fig. 6D).
CDH3 expression in human CRC tissues
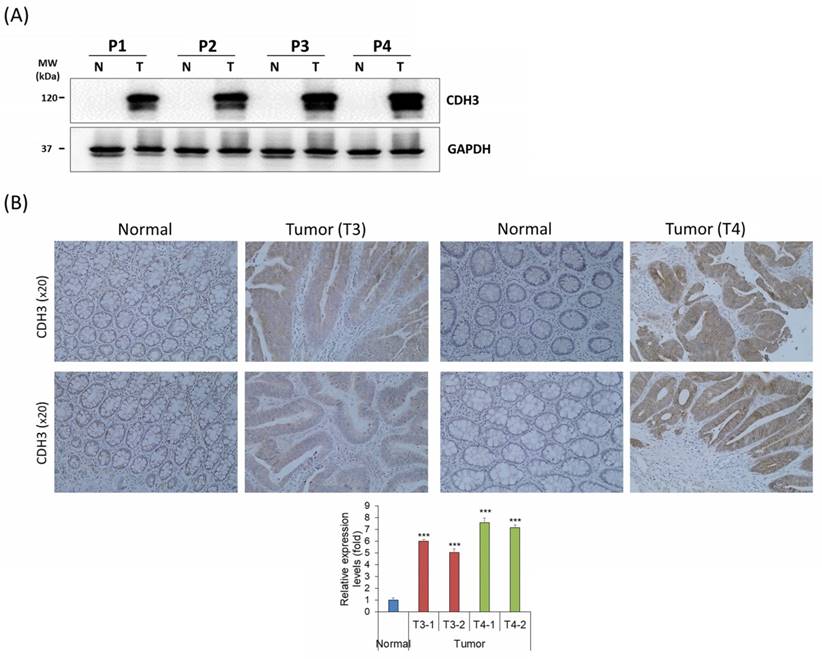
CDH3 expression was analyzed in four human CRC tissue samples and adjacent colon tissues using western blot analysis and IHC. The CDH3 protein expression was significantly upregulated in all CRC tissues compared with the healthy normal colon tissues (Fig. 7A, B). The result displayed that CDH3 expression increased with tumor progression (Fig. 7A, B).
Endogenous CDH3 expression in human CRC tissues (A) CDH3 protein expression in paired human CRC and adjacent normal colon tissue samples from four patients (P#). P1 and P2 were tumor (T) stage 3 (T3), and P3 and P4 were T stage 4 (T4). (B) CDH3 IHC expression in paired colon cancer and adjacent normal tissue samples (N) (20× magnification) from four patients. Differences were considered statistically significant compared with control (*** P < 0.001).
Discussion
Colon cancer is the prominent cause of global cancer-associated mortality. Millions of new cases and thousands of cancer-related deaths are recorded each year. Activation of oncogenes and suppression of tumor suppressor genes are thought to lead to the initiation of CRC. An improved understanding of our knowledge of the oncogenes, mechanism of development, invasion, and reoccurrence of CRC, including validation of new biological markers of colon cancer, would aid in the early identification and diagnosis of colon cancer [31]. Many studies have shown that miRNAs may have therapeutic significance as biomarkers as they regulate post-transcriptional gene expression in multicellular organisms. Our previous study showed that MIR133A is downregulated in CRC, which has also been reported in other studies [13,15,16, 32]. However, the association of the MIR133A and CDH3 genes in CRC is unexplored.
We identified CDH3 as the target gene of MIR133A (Fig. 1A, B) in CRC cells, and CDH3 mRNA and protein expression was significantly reduced in MIR133A over-expressed cell lines (Fig. 1C, D). These data revealed that CDH3 expression is negatively linked with MIR133A in CRC tissues. Additionally, CDH3 protein expression was dramatically increased in CRC tissue in comparison with mock (Fig. 7A, B). Our data signified that the downregulation of MIR133A in CRC progression upregulates CDH3 expression in human CRC tissues.
After selecting the target gene and verifying mRNA and protein levels, we assessed the regulation of MIR133A in various signaling pathways involved in CRC tumorigenesis. The EMT converts static epithelial cells into an invasive mesenchymal phenotype. Downregulation of the CDH1 gene and upregulation of CDH3, CDH2, and VIM in EMT is a hallmark of cancer metastasis [33]. High expression of CDH3 disrupts epithelial adhesion and fosters the development of a more mesenchymal and progenitor-like phenotype, ultimately resulting in cancer [30]. In this study, CDH3, VIM, and CDH2 were inhibited in MIR133A and siCDH3 overexpressed cell lines (Fig. 2A, B, C), whereas there was no change in CDH1 expression (Fig. 2A, B). The extent of metastasis and degree of invasiveness of cancer cells are related to cancer prognosis. The ability to metastasize and MMP activities were found to be greater in the LoVo cell line than in SW480 cells [34,35]. Hence, we assessed the regulation of the EMT pathway by MIR133A and siCDH3 transfection in the LoVo cell line. The expression of CDH2 and VIM was downregulated, and CDH1 was upregulated by MIR133A and siCDH3 transfection (Fig. 2C). These results suggest that MIR133A regulates the EMT pathway of colon cancer by regulating CDH3-dependent CDH2 and VIM expression. Similarly, MIR133A regulates the CRC progression and metastasis by targeting LIM and SH3 protein 1 and inhibiting the MAPK pathway [36].
Increased CDH3 expression also increases cell motility and migration in cancer by interfering with CTNND1 and CDH1. CDH3 binds to the juxtamembrane domain of CTNND1 and prevents the binding of CDH1, which stabilizes CTNND1 [37]. In this study, we investigated CTNND1 protein expression in a MIR133A overexpressed cell line. We found that CTNND1 expression is reduced after transfection with MIR133A (Fig. 3A), and siCDH3 overexpression produced similar results (Fig. 3B). The cadherin-catenin junction helps to maintain the cell structure and cell integrity and CTNNB1 helps in cell adhesion. A previous study reported that CTNNB1 is highly expressed in colorectal cancer [38]. AKT phosphorylates CTNNB1 at Ser552, dissociating it from the WNT complex and leading to its accumulation in the cytosol and nucleus. MIR133A deactivates the PI3K/AKT pathway in CRC cell lines, and PI3K/AKT inhibits GSK3B and promotes cyclin D1 to increase its transcriptional activity, ultimately increasing GSK3B and decreasing CTNNB1 [16,39]. In this study, overexpression of MIR133A regulated the CDH3-mediated GSK3B and CTNNB1 signaling pathways (Fig. 3A, B).
The MMP family comprises enzymes that degrade the extracellular matrix. The MMPs play an essential role in morphogenesis and wound healing [40] and are involved in initiating metastasis, invasion, and carcinogenesis from colorectal adenomas [41]. Both MMP1 mRNA and protein are highly expressed in various cancers and play a vital role in initiating tumorigenesis by disrupting the stroma and releasing growth factors. The degradation of the extracellular matrix (ECM) by MMP1 increases cell-ECM interaction, resulting in cell separation from ECM and leading to invasion and metastasis [42]. In addition, MMP2 plays a vital role in disrupting the ECM during metastasis [43]. Tumor cells can produce MMP2 or influence host cells to secrete it, as active MMP2 is routinely detected in malignant tumors [44]. Overexpression of CDH3 increases MMP activity, which is accountable for cleavage and shedding [45]. Our results suggest that MMP1 expression is downregulated by both MIR133A and siCDH3 transfection in CRC cells (Fig. 5A). However, MMP2 expression was only downregulated by MIR133A transfection and not by siCDH3 transfection (Fig. 5B). These data indicate that MIR133A regulates CDH3-dependent MMP1 expression, while MMP2 expression is not CDH3-dependent in CRC cells.
Likewise, tumor formation is influenced by the balance between cell growth and apoptosis. If apoptosis is reduced, cell proliferation may become uncontrollable, which leads to cancer [46]. Hence, reducing cell proliferation and inducing apoptosis may be an excellent approach for cancer treatment. This study linked cell apoptosis with various genes, like BCL2, BAX, BIRC5, CASP family, RIP3, and MLKL. For example, BCL2 is abnormally expressed in CRC and linked to tumor growth and development, while BAX is a BCL2 family protein and regulates cell development and apoptosis. The expression of BAX is reduced in colon cancer, which is associated with the advancement of cancer [47, 48]. Similarly, increased BIRC5 expression prevents apoptosis through inhibition of CASP3 and CASP7 and is also involved in chemoresistance and angiogenesis in CRC. BIRC5 is also directly correlated with anti-apoptotic proteins TACE and MCL1 and inversely correlated with the pro-apoptotic gene BAX [49]. In our study, increased BAX and decreased BCL2 and BIRC5 expression were detected in both MIR133A-and siCDH3-transfected CRC cells (Fig. 4A, B). Associations between CASP3 and CASP8 expression and tumorigenesis and poor prognosis of CRC have been reported [50]. Here, our results showed that CASP9, CASP8, and CASP3 are increased by both MIR133A and siCDH3 transfection in CRC cells (Fig. 4A, B). We also assessed the expression of the necroptosis markers RIP3 and MLKL and found that MIR133A and siCDH3 transfection upregulated the expression of MLKL, whereas there was no change in RIP3 (Fig. 4C, D). To further validate the effects on apoptosis, FACS analysis was done, which indicated that transfection of MIR133A and siCDH3 increased apoptosis in CRC cell lines (Fig. 6D). Here, our results indicate that CASP9, CASP8, and CASP3 expression is increased by MIR133A transfection in CRC cells (Fig. 4A, B), leading to increased apoptosis (Fig. 6D). Likewise, a study showed that MIR133A is associated with G0/G1 phase arrest and regulates the upregulation of the G1 phase regulator [51].
In addition, colony formation, migration, and MTT assays showed decreased colony formation, reduced migration rate, and diminished cell viability of CRC cell line after transfection with MIR133A and siCDH3 (Fig. 6A, B, C). These results suggest that the reduction of cell viability and impaired migration of CRC cell lines were due to increased apoptosis following MIR133A transfection, which was also seen in western blot analyses, as the expression of apoptotic proteins like BAX, CASP9, CASP8, and CASP3 was increased, while that of BCL2 and BIRC5 was reduced (Fig 4). Overall, these data indicate that MIR133A regulates apoptosis in the human CRC cell line. Similarly, the regulation of MIR133A in cell migration, colony formation, and invasion is reported by various other studies. MIR133A suppresses cell proliferation viability, migration, and invasion by regulating AQP1 and eIF4A1 and inhibits cell proliferation by targeting SENP1 [52,53,54].
Conclusion
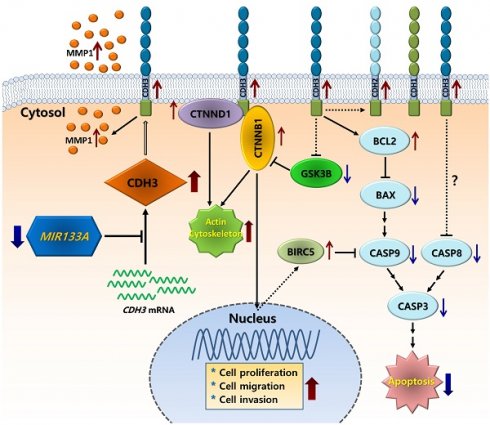
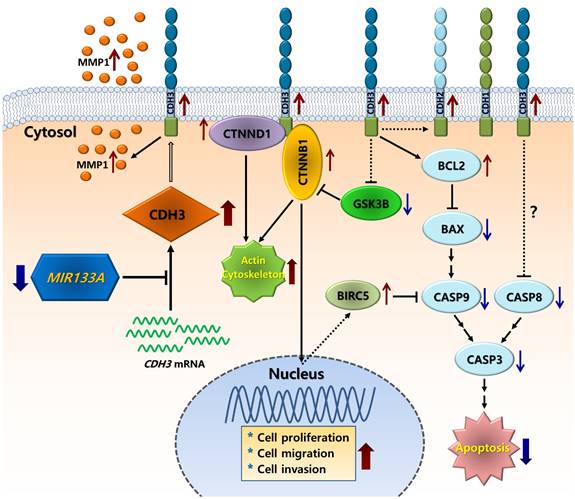
In conclusion, we showed that CDH3 is a direct target of MIR133A, and CDH3 is overexpressed in CRC tissues in comparison with their respective normal tissues. Expression of both CDH3 mRNA and protein was significantly downregulated in CRC cells upon transfection with MIR133A. The silencing of CDH3 in CRC cell lines by siCDH3 produced similar results. Furthermore, our molecular and functional studies suggest that MIR133A regulates CDH3 to induce apoptosis and reduce cell viability, migration, and colony formation in CRC cell lines. Our results suggest that the reduction of MIR133A during human CRC progression upregulates CDH3 expression. The upregulation of intracellular CDH3 may, in turn, affect downstream or associated signal pathways to upregulate cell proliferation, cell migration, and colony formation, as seen in our experiments, while apoptosis was downregulated in CRC cells (Fig. 8). Although we did not demonstrate these effects in vivo, MIR133A produces a potent antitumor effect in vitro. Our results suggest that MIR133A regulates CDH3 in human CRC; therefore, MIR133A may be a therapeutic target in human CRC, which needs to be examined further.
Schematic diagram of the putative mechanism of MIR133A in CRC Reduction of MIR133A expression in CRC cells or tissues leads to upregulation of cellular CDH3 expression. Upregulation of CDH3 directly or indirectly activates downstream or associated pathways, such as CTNNB1, CTNND1, GSK3B, MMP1, and apoptosis pathways, resulting in increased cell proliferation, cell migration, and cell invasion.
Abbreviations
3' UTR, three prime untranslated region; BAX, BCL2 associated X; BCL2, BCL2 apoptosis regulator; BIRC5, baculoviral IAP repeat-containing 5; CASP3, caspase 3; CASP8, caspase 8; CASP9, caspase 9; CDH1, epithelial cadherin; CDH2, neural cadherin; CDH3, cadherin 3; CRC, colorectal carcinoma; CTNNB1, catenin beta 1; CTNND1, catenin delta 1; EMT, epithelial-mesenchymal transition; GSK3β, glycogen synthase kinase 3 beta; IBD, inflammatory bowel disease; IHC, immunohistochemistry; MIR133A, microRNA 133A; MLKL, mixed lineage kinase domain-like pseudokinase; MMP1, matrix metalloproteinase-1; MMP2, matrix metalloproteinase-2; qRT-PCR, quantitative Real time polymerase chain reaction; RIP3, receptor-interacting serine/threonine kinase 3; siCDH3, small interfering cadherin-3; VIM, vimentin.
Supplementary Material
Supplementary figure and table.
Acknowledgements
This study used biospecimens from Wonkwang University Hospital's Biobank, a member of the National Biobank of Korea supported by the Ministry of Health and Welfare.
Funding
This project was funded by the National Research Foundation of Korea (NRF) grant funded by the Korea government (MSIT); 2020R1A2C2003882.
Author contributions
The authors stated: Conceived and designed experiments: Chae-Soo-Cheon. Conducted and analyzed the experiments: Grinsun Sharma and Soo-Cheon Chae. Contributed Reagents/materials and discussion: Ji-Su Mo and Santosh Lamichhane. Grinsun Sharma and Soo-Cheon Chae contributed to the draft of manuscript.
Ethics statement
Every process done in this study comprising humans were in accord with Helsinki declaration and approved by the Committee of Ethical Standards of Wonkwang University, Republic of Korea (WKIRB-202006-BR-023).
Data availability statement
This article and supplementary figures contain all studied data.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Javid G, Zargar SA, Rather S. et al. Incidence of colorectal cancer in Kashmir valley, India. Indian J Gastroenterol. 2011;30:7-11
2. Sung H, Ferlay J, Siegel RL. et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: Cancer J Clin. 2021;71:209-49
3. Mármol I, Sánchez-de-Diego C, Dieste AP. et al. Colorectal carcinoma: A general overview and future perspectives in colorectal cancer. Int J Mol Sci. 2017;18:197
4. Alimperti S, Andreadis ST. CDH2 and CDH11 act as regulators of stem cell fate decisions. Stem Cell Res. 2015;14:270-82
5. Taneyhill LA, Schiffmacher AT. Should I stay or should I go? Cadherin function and regulation in the neural crest. Genesis. 2017;55:e23028
6. Zhou Y, Chi Y, Bhandari A. et al. Downregulated CDH3 decreases proliferation, migration, and invasion in thyroid cancer. Am J Transl Res. 2020;12:3057-67
7. Paredes J, Correia AL, Ribeiro AS. et al. P-cadherin expression in breast cancer: A review. Breast Cancer Res. 2007;9:214
8. Kumara HS, Bellini GA, Caballero OL. et al. P-cadherin (CDH3) is overexpressed in colorectal tumors and has potential as a serum marker for colorectal cancer monitoring. Oncoscience. 2017;4:139-47
9. Xu Y, Zhao J, Dai X. et al. High expression of CDH3 predicts a good prognosis for colon adenocarcinoma patients. Exp Ther Med. 2019;18:841-47
10. Milicic A, Harrison LA, Goodlad RA. et al. Ectopic expression of P-cadherin correlates with promoter hypomethylation early in colorectal carcinogenesis and enhanced intestinal crypt fission in vivo. Cancer Res. 2008;68:7760-68
11. Paredes J, Albergaria A, Oliveira JT. et al. P-cadherin overexpression is an indicator of clinical outcome in invasive breast carcinomas and is associated with CDH3 promoter hypomethylation. Clin Cancer Res. 2005;11:5869-77
12. Bandres E, Agirre X, Bitarte N. et al. Epigenetic regulation of microRNA expression in colorectal cancer. Int J Cancer. 2009;125:2737-43
13. Dong Y, Zhao J, Wu CW. et al. Tumor suppressor functions of MiR-133a in colorectal cancer. Mol Cancer Res. 2013;11:1051-60
14. Hua YT, Xu WX, Li H. et al. Emerging roles of MiR-133a in human cancers. J Cancer. 2021;12:198-206
15. Li W, Chen A, Xiong L. et al. MiR-133a acts as a tumor suppressor in colorectal cancer by targeting eIF4A1. Tumor Biol. 2017;39:1010428317698389
16. Lamichhane S, Mo JS, Sharma G. et al. MIR133A regulates cell proliferation, migration, and apoptosis by targeting SOX9 in human colorectal cancer cells. Am J Cancer Res. 2022;12:3223-41
17. Wang LL, Du LT, Li J. et al. Decreased expression of MiR-133a correlates with poor prognosis in colorectal cancer patients. World J Gastroenterol. 2014;20:11340-46
18. Alam KJ, Mo JS, Han SH. et al. MicroRNA 375 regulates proliferation and migration of colon cancer cells by suppressing the CTGF-EGFR signaling pathway. Int J Cancer. 2017;141:1614-29
19. Mo JS, Chae SC. MicroRNA 452 regulates ASB8, NOL8, and CDR2 expression in colorectal cancer cells. Genes & Genomics. 2021;43:33-41
20. Mo JS, Alam KJ, Kim HS. et al. MicroRNA 429 regulates mucin gene expression and secretion in murine model of colitis. J Crohns Colitis. 2016;10:837-49
21. Lamichhane S, Mo JS, Sharma G. et al. MicroRNA 452 regulates IL20RA mediated JAK1/STAT3 pathway in inflammatory colitis and colorectal cancer. Inflamm Res. 2021;70:903-14
22. Mo JS, Park WC, Choi SC. et al. MicroRNA 452 regulates cell proliferation, cell migration, and angiogenesis in colorectal cancer by suppressing VEGFA expression. Cancers. 2019;11:1613
23. Han SH, Mo JS, Park WC. et al. Reduced microRNA 375 in colorectal cancer upregulates metadherin-mediated signaling. World J Gastroenterol. 2019;25:6495-507
24. Mo JS, Han SH, Yun KJ. et al. MicroRNA 429 regulates the expression of CHMP5 in the inflammatory colitis and colorectal cancer cells. Inflamm Res. 2018;67:985-96
25. Georgakopoulos-Soares I, Chartoumpekis DV, Kyriazopoulou V. et al. EMT factors and metabolic pathways in cancer. Front Oncol. 2020;10:499
26. Zhou Y, Chi Y, Bhandari A. et al. Downregulated CDH3 decreases proliferation, migration, and invasion in thyroid cancer. Am J Transl Res. 2020;12:3057-67
27. Taniuchi K, Nakagawa H, Hosokawa M. et al. Overexpressed P-cadherin/CDH3 promotes motility of pancreatic cancer cells by interacting with p120ctn and activating rho-family GTPases. Cancer Res. 2005;65:3092-99
28. Wang H. MicroRNAs and apoptosis in colorectal cancer. Int J Mol Sci. 2020;21:5353
29. Herszényi L, Hritz I, Lakatos G. et al. The behavior of matrix metalloproteinases and their inhibitors in colorectal cancer. Int J Mol Sci. 2012;13:13240-63
30. Vieira AF, Paredes J. P-cadherin and the journey to cancer metastasis. Mol Cancer. 2015;14:178
31. Zeng M, Zhu L, Li L. et al. MiR-378 suppresses the proliferation, migration and invasion of colon cancer cells by inhibiting SDAD1. Cell Mol Biol Lett. 2017;22:12
32. Wan TM, Lam CS, Ng L. et al. The clinicopathological significance of miR-133a in colorectal cancer. Dis. Markers. 2014:1-8
33. Huang Lx, Hu Cy, Jing L. et al. MicroRNA-219-5p inhibits epithelial-mesenchymal transition and metastasis of colorectal cancer by targeting lymphoid enhancer-binding factor 1. Cancer Sci. 2017;108:1985-95
34. Ribeiro AS, Paredes J. P-cadherin linking breast cancer stem cells and invasion: A promising marker to identify an intermediate/metastable EMT state. Front Oncol. 2015;4:371
35. Sul JY, Song IS, Bae CS. et al. Metastatic potential of human colon cancer cell lines, LoVo and SW480. Journal of the Korean Cancer Association. 1995;27:209-23
36. Wang H, An H, Wang B. et al. MiR-133a represses tumour growth and metastasis in colorectal cancer by targeting LIM and SH3 protein 1 and inhibiting the MAPK pathway. Eur J Cancer. 2013;49:3924-35
37. Li C, Ma H, Wang Y. et al. Excess PLAC8 promotes an unconventional ERK2-dependent EMT in colon cancer. J Clin Investig. 2014;124:2172-87
38. Wang B, Li X, Liu L. et al. β-catenin: Oncogenic role and therapeutic target in cervical cancer. Biol Res. 2020;53:33
39. Fang D, Hawke D, Zheng Y. et al. Phosphorylation of beta-catenin by AKT promotes beta-catenin transcriptional activity. J Biol Chem. 2007;282:11221-29
40. Nagase H, Visse R, Murphy G. Structure and function of matrix metalloproteinases and TIMPs. Cardiovasc Res. 2006;69:562-73
41. Herszényi L, Hritz I, Lakatos G. et al. The behavior of matrix metalloproteinases and their inhibitors in colorectal cancer. Int J Mol Sci. 2012;13:13240-63
42. Poola I, Dewitty RL, Marshalleck JJ. et al. Identification of MMP-1 as a putative breast cancer predictive marker by global gene expression analysis. Nat Med. 2005;11:481-83
43. Damodharan U, Ganesan R, Radhakrishnan UC. Expression of MMP2 and MMP9 (gelatinases A and B) in human colon cancer cells. Appl Biochem Biotechnol. 2011;165:1245-52
44. Mendes O, Kim HT, Lungu G. et al. MMP2 role in breast cancer brain metastasis development and its regulation by TIMP2 and ERK1/2. Clin Exp Metastasis. 2007;24:341-51
45. Ribeiro A, Albergaria A, Sousa B. et al. Extracellular cleavage and shedding of P-cadherin: A mechanism underlying the invasive behaviour of breast cancer cells. Oncogene. 2010;29:392-402
46. Wong RS. Apoptosis in cancer: From pathogenesis to treatment. J Exp Clin. Cancer Res. 2011;30:87
47. Poincloux L, Durando X, Seitz JF. et al. Loss of Bcl-2 expression in colon cancer: A prognostic factor for recurrence in stage II colon cancer. Surg Oncol. 2009;18:357-365
48. Pryczynicz A, Gryko M, Niewiarowska K. et al. Bax protein may influence the invasion of colorectal cancer. World J Gastroenterol. 2014;20:1305-10
49. Hernandez JM, Farma JM, Coppola D. et al. Expression of the antiapoptotic protein survivin in colon cancer. Clin Colorectal Cancer. 2011;10:188-93
50. Yao Q, Wang W, Jin J. et al. Synergistic role of caspase-8 and caspase-3 expressions: Prognostic and predictive biomarkers in colorectal cancer. Cancer Biomark. 2018;21:899-908
51. Dong Y, Zhao J, Wu CW. et al. Tumor Suppressor Functions of miR-133a in Colorectal Cancer. Mol Cancer Res. 2013;11:1051-60
52. Kong B, Zhao S, Kang X. et al. MicroRNA 133a 3p inhibits cell proliferation, migration and invasion in colorectal cancer by targeting AQP1. Oncol Lett. 2021;22:1-10
53. Li W, Chen A, Xiong L. et al. MiR-133a acts as a tumor suppressor in colorectal cancer by targeting eIF4A1. Tumor Biol. 2017;39:1-9
54. Zhou GQ, Han F, Shi ZL. et al. MiR-133a-3p targets SUMO-specific protease 1 to inhibit cell proliferation and cell cycle progress in colorectal cancer. Oncol Res. 2018;26:795-800
Author contact
Corresponding author: Soo-Cheon Chae Department of Pathology, School of Medicine, Wonkwang University, Iksan, Chonbuk 54538, Republic of Korea TEL. +82-63-8506954, Email: chaescac.kr ORCID No. 0000-0002-5427-714X

 Global reach, higher impact
Global reach, higher impact