Impact Factor
ISSN: 1837-9664
J Cancer 2022; 13(8):2490-2503. doi:10.7150/jca.69274 This issue Cite
Research Paper
New Clues to Prognostic Biomarkers of Four Hematological Malignancies
1. Department of Biochemistry and Biotechnology, Kwame Nkrumah University of Science and Technology (KNUST), Kumasi, Ghana.
2. Kumasi Centre for Collaborative Research in Tropical Medicine (KCCR), Kumasi, Ghana.
Abstract
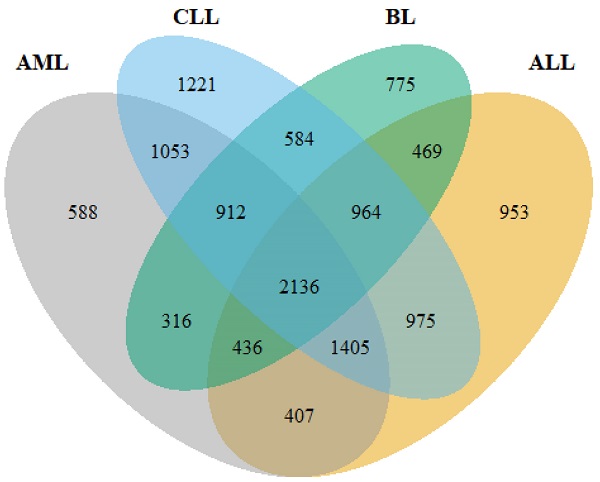
Globally, one out of every two reported cases of hematologic malignancies (HMs) results in death. Each year approximately 1.24 million cases of HMs are recorded, of which 58% become fatal. Early detection remains critical in the management and treatment of HMs. However, this is thwarted by the inadequate number of reliable biomarkers. In this study, we mined public databases for RNA-seq data on four common HMs intending to identify novel biomarkers that could serve as HM management and treatment targets. A standard RNA-seq analysis pipeline was strictly adhered to in identifying differentially expressed genes (DEGs) with DESeq2, limma+voom and edgeR. We further performed gene enrichment analysis, protein-protein interaction (PPI) network analysis, survival analysis and tumor immune infiltration level detection on the genes using G:Profiler, Cytoscape and STRING, GEPIA tool and TIMER, respectively. A total of 2,136 highly-ranked DEGs were identified in HM vs. non-HM samples. Gene ontology and pathway enrichment analyses revealed the DEGs to be mainly enriched in steroid biosynthesis (5.075×10-4), cholesterol biosynthesis (2.525×10-8), protein binding (3.308×10-18), catalytic activity (2.158×10-10) and biogenesis (5.929×10-8). The PPI network resulted in 60 hub genes which were verified with data from TCGA, MET500, CPTAC and GTEx projects. Survival analyses with clinical data from TCGA showed that high expression of SRSF1, SRSF6, UBE2Z and PCF11, and low expression of HECW2 were correlated with poor prognosis in HMs. In summary, our study unraveled essential genes that could serve as potential biomarkers for prognosis and may serve as drug targets for HM management.
Keywords: Hematologic malignancies, Hub genes, RNA-seq, Lymphoma, Biomarker

 Global reach, higher impact
Global reach, higher impact